Intelligent Meal Planning: Kefir’s Role in Nutrition
Intelligent Meal Planning: Algorithmic Inclusion of Kefir in Nutritional Recommendations
Abbas Maazallahi¹, Mohammad Amir Salari², Payam Norouzzadeh³, Eli Snir⁴, Maria Jose Romo-Palafox⁵, Bahareh Rahmani⁶*
- University of Tehran, Computer Science Department, Tehran, Iran
- Saint Louis University, Computer Science Department, Saint Louis, MO
- Saint Louis University, Professional Studies Department, Saint Louis, MO
- Washington University in Saint Louis, Business School, Saint Louis, MO
- Saint Louis University, College of Health Science, Saint Louis, MO
- Saint Louis University, Health & Clinical Outcome Research Department, Saint Louis, MO
OPEN ACCESS
PUBLISHED 31 October 2025
CITATION; Maazallahi, A., Salari, MA., et al., 2025. Intelligent Meal Planning: Algorithmic Inclusion of Kefir in Nutritional Recommendations. Medical Research Archives, [online] 13(10). https://doi.org/10.18103/mra.v13i10.6956
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI: https://doi.org/10.18103/mra.v13i10.6956
ISSN 2375-1924
ABSTRACT
This study employs advanced machine learning techniques to systematically analyze comprehensive nutritional data from the Food and Nutrient Database for Dietary Studies (FNDDS), focusing on dairy products. The primary goal is to distinguish the nutritional profiles of fermented dairy foods, such as kefir and yogurt, from non-fermented dairy counterparts, like milk and cream. Leveraging a robust dataset encompassing detailed nutrient information, this research aims to identify unique nutritional characteristics inherent to fermented dairy products that may contribute significantly to dietary interventions aimed at health enhancement and chronic disease prevention. Findings from this analysis offer practical insights for dietary planning, emphasizing evidence-based nutritional recommendations, and underscore the critical role of fermented dairy in personalized healthcare strategies.
Keywords: Diet, Disease, Machine Learning, Nutritional Interventions, Data Analysis.
1. Introduction
Nutritional science increasingly recognizes diet as a pivotal factor influencing human health, disease prevention, and overall wellness. As chronic diseases such as cardiovascular conditions, diabetes, and various forms of cancer impose substantial burdens globally, understanding how specific dietary components affect health outcomes has become imperative. Dairy products, long appreciated for their nutritional contributions, have garnered significant interest due to emerging evidence suggesting distinct health benefits associated with fermented dairy products, such as kefir, yogurt, and cultured cheeses, compared to their non-fermented counterparts like milk and cream. Fermented dairy products undergo microbiological processes that enhance their nutritional and probiotic profiles, potentially offering health advantages beyond basic nutrient provision. However, the differentiation between the nutritional profiles of fermented and non-fermented dairy products has not been thoroughly characterized through comprehensive data-driven approaches. This study addresses this gap by applying sophisticated machine learning techniques to detailed nutritional data extracted from the Food and Nutrient Database for Dietary Studies (FNDDS).
By systematically comparing fermented and non-fermented dairy products using machine learning models, we aim to identify and highlight distinct nutritional components that uniquely characterize fermented dairy. Such insights have the potential to refine dietary guidelines, support the development of targeted nutritional interventions, and enhance public health strategies. Ultimately, this research contributes to the broader integration of precision nutrition into healthcare practices, underscoring the importance of dietary choices informed by rigorous, evidence-based analysis.
In the contemporary medical discourse, “food as medicine” is not a mere aphorism but an integrative approach towards combating the burgeoning global epidemic of diet-related chronic diseases. The intricate relationship between nutrition and chronic disease management is undeniable, with diet being recognized as a modifiable risk factor for a host of chronic conditions existing in isolation or comorbidly. Non-communicable diseases (NCDs) such as cardiovascular disease, diabetes, and various forms of cancer pose a substantial burden on healthcare systems across the globe, affecting populations in both developed and developing nations. The significance of dietary patterns in the context of these diseases is profound, as the Global Burden of Disease Study illustrates, positioning diet as a leading contributor to morbidity and mortality worldwide. Moreover, nutritional genomics heralds a new frontier, suggesting that genome-based dietary guidelines could markedly improve public health outcomes. The “food is medicine” paradigm, which encompasses interventions like medically tailored meals and produce prescription programs, is increasingly being piloted with the intention of embedding these into the structure of healthcare systems.
However, the integration of such nutrition interventions into healthcare is encumbered by a multitude of challenges. There is a pressing need for rigorous research to address the knowledge gaps and test the efficacy of various approaches. Clinicians, at the vanguard of patient care, require enhanced education and training to adeptly navigate and utilize these interventions. Furthermore, the establishment of sustainable funding streams is critical to ensure equitable patient access to these potentially life-altering nutrition interventions.
A comprehensive review explicates how ML facilitates early detection of various diseases, which is pivotal for effective treatment. In the sphere of nutritional epidemiology, observational data is crucial for identifying associations between diet, nutritional results, and disease risk. However, traditional analytic methods have limitations, such as inadequate incorporation of correlations and nonlinear behaviors in dietary data. This gap presents an opportunity for machine learning to advance the field.
Furthermore, the application of ML in foodborne disease surveillance has been showcased through initiatives like China’s National Foodborne Disease Outbreak Surveillance System, which relies on a wealth of data from case reports and outbreak surveillance. The importance of accurately predicting foodborne disease pathogens is critical, not only for public health but also for the economic implications of outbreaks. Machine learning prediction models are becoming indispensable in identifying these pathogens and aiding in the prevention and treatment of foodborne diseases.
Lastly, the deployment of machine learning tools in healthcare decision-making is expanding, particularly in disease prediction and detection. These tools are instrumental in diagnosing diseases at early stages, thereby simplifying treatment and increasing the likelihood of patient recovery. RIP-M showcases machine learning’s impact on healthcare by enhancing gene network analysis, crucial for understanding disease pathways. It optimizes the clustering of co-expressed genes, facilitating targeted gene-based treatments. This method reflects the potential of machine learning to refine diagnostic precision and personalize patient care. Through its application, RIP-M underscores the transformative role of ML in advancing healthcare outcomes.
In the related work, the integration of machine learning (ML) in nutritional epidemiology is emerging as a transformative approach to understanding the complex associations between diet and disease risk. The current methodologies in nutritional data analysis face limitations due to their insufficient handling of nonlinear behaviors and interactions between dietary components. Machine learning offers robust solutions to these challenges, enhancing the precision and validity of dietary assessments and interventions.
Furthermore, advancements in ML have facilitated the development of systems capable of accurately classifying food images and estimating nutritional attributes. Such technologies are pivotal for maintaining a balanced diet and can potentially be used to tailor dietary interventions to prevent and treat obesity and other diet-related diseases.
Big data combined with machine learning presents a promising frontier in nutritional epidemiology. These tools can help address measurement errors, diet’s complexity as exposure, and residual confounding, thereby offering new methods of dietary measurement and modeling the intricate relations between diet and diseases. Machine learning methods have been widely applied in studies related to obesity, providing insights that can improve the efficiency of interventions targeting nutrient intake and dietary patterns.
The computational diet, involving the use of computational methods in nutrition research, has shown that the human microbiome may modulate health in ways related to diet. This insight is crucial for developing dietary interventions that consider the unique microbiological aspects of human health.
Lastly, a perspective on big data and machine learning in nutritional epidemiology highlights how these approaches could revolutionize dietary measurement and modeling, offering tools to address the complexity of diet and its relations with diseases. The application of these methods promises to enhance the development of precise and effective dietary interventions for disease management.
These references underscore the potential of machine learning to revolutionize our understanding of the intricate relationship between food, diet, and disease, paving the way for personalized dietary interventions informed by comprehensive data analysis.
This project endeavors to scrutinize an extensive array of foods and their nutritional components, leveraging a comprehensive dataset encompassing over 7000 foods to discern their impacts on various nutritional measures. Through systematic classification, we aim to lay a foundation for tailored nutritional packages, grounded in empirical data. This foundation can be for dietary prescriptions aligned with the principles of personalized medicine. Such a venture not only promises to enhance our understanding of the nexus between diet and disease but also paves the way for evidence-based dietary interventions that could revolutionize patient care and disease management.
2. Dataset Description
The Food and Nutrient Database for Dietary Studies (FNDDS) is a detailed nutritional database used to analyze dietary intake data, particularly collected from national surveys such as NHANES. Two key datasets from the FNDDS used in this study are:
- FNDDS Nutrient Values Dataset: This dataset contains detailed nutrient profiles for numerous food items, including information on macro- and micronutrients such as calories, carbohydrates, proteins, fats (lipids), dietary fibers, vitamins, and minerals. Each nutrient value is standardized per 100 grams, facilitating direct comparisons between food items.
- Foods and Beverages Dataset: This dataset provides detailed descriptions of foods and beverages, categorizing them into various groups such as dairy products, meats, vegetables, fruits, grains, beverages, and snacks. It includes descriptive attributes that identify the specific types of food items, their preparation methods, and any unique characteristics.
From these datasets, dairy foods can be specifically distinguished based on their categorization in the Foods and Beverages dataset. Dairy items include both fermented products like kefir, yogurt, and cheese, and non-fermented items like milk, cream, and butter. By linking the categorization from the Foods and Beverages dataset with nutrient values from the Nutrient Values dataset, we construct a comprehensive analytical dataset explicitly tailored to analyze and compare nutritional profiles between fermented and non-fermented dairy products.
By systematically comparing fermented and non-fermented dairy products using machine learning models, we aim to identify and highlight distinct nutritional components that uniquely characterize fermented dairy. Such insights have the potential to refine dietary guidelines, support the development of targeted nutritional interventions, and enhance public health strategies. Ultimately, this research contributes to the broader integration of precision nutrition into healthcare practices, underscoring the importance of dietary choices informed by rigorous, evidence-based analysis.
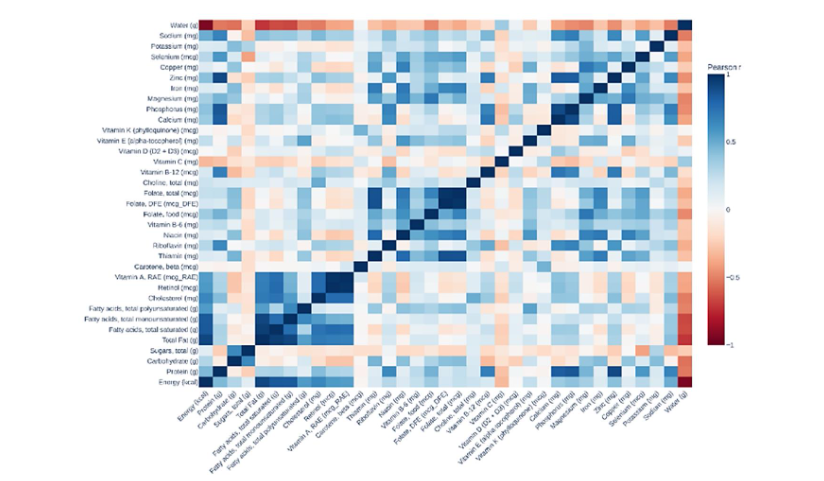
2.1 CORRELATION OF NUTRITIONAL FEATURES
In exploring nutritional profiles within dairy products, several key nutrients were analyzed, including macronutrients such as carbohydrates, proteins, and fats, alongside micronutrients including vitamins and minerals such as calcium, phosphorus, vitamin B-12, vitamin D, potassium, sodium, and cholesterol. Correlation analysis between these nutrients reveals meaningful associations, which are visualized through correlation heatmaps. Strong positive correlations are evident between energy (calories), total fat, and saturated fatty acids, indicating their interconnected dietary presence. Conversely, nutrients such as water content show strong negative correlations with macronutrients and energy density. Additionally, micronutrients like calcium and phosphorus exhibit significant correlations, reflecting their joint presence in dairy products. Understanding these nutrient relationships is crucial for developing dietary guidelines and effectively differentiating the nutritional profiles of fermented versus non-fermented dairy products. By systematically comparing fermented and non-fermented dairy products using machine learning models, we aim to identify and highlight distinct nutritional components that uniquely characterize fermented dairy. Such insights have the potential to refine dietary guidelines, support the development of targeted nutritional interventions, and enhance public health strategies. Ultimately, this research contributes to the broader integration of precision nutrition into healthcare practices, underscoring the importance of dietary choices informed by rigorous, evidence-based analysis.
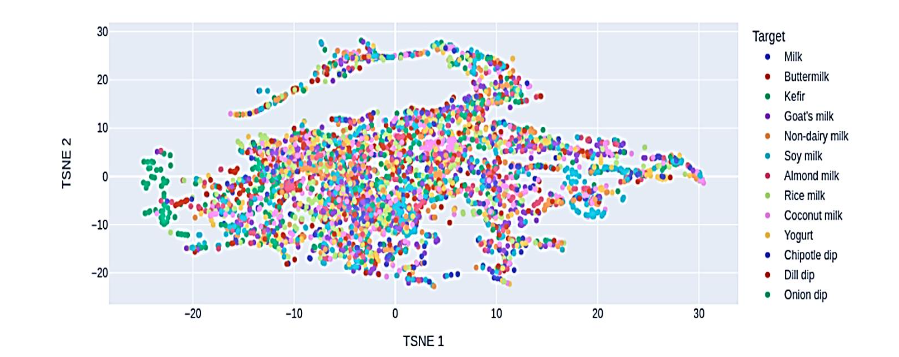
2.2 t-DISTRIBUTED STOCHASTIC NEIGHBOR EMBEDDING
In our exploratory analysis employing t-Distributed Stochastic Neighbor Embedding (t-SNE), a notable pattern emerged from the scatter plot, offering valuable insights into the structure of our food dataset. The plot, a result of t-SNE’s capability to project high-dimensional data into a lower-dimensional space, revealed interesting distinctions among various food categories. A striking observation was the distinct separation of certain categories, such as ‘kefir’, which appeared as a clearly isolated cluster. This distinctiveness of ‘kefir’ suggests unique nutritional characteristics or a specific combination of features that set it apart from other food items in the dataset. Conversely, the t-SNE scatter plot also highlighted a degree of overlap among several other food categories. Unlike ‘kefir’, these categories did not form distinct clusters, indicating a closer nutritional similarity or less pronounced differentiation in their feature space. This blending of categories in the t-SNE visualization can be attributed to t-SNE’s focus on preserving local similarities: foods sharing similar nutritional profiles appear closer in the reduced space, leading to these overlapping regions.
3. Methodology
3.1 DATA PREPARATION
The initial phase involved extracting dairy-specific data from the FNDDS Foods and Beverages dataset and merging it with corresponding nutrient values from the FNDDS Nutrient Values dataset. Foods were classified into two primary categories: fermented (e.g., yogurt, kefir, cheese) and non-fermented (e.g., milk, cream, butter). Standardization per 100 grams ensured comparability across different food items.
3.2 FEATURE ENGINEERING
Nutritional variables, including macro- and micronutrients, were selected based on their relevance to dietary interventions and health outcomes. Features included carbohydrates, proteins, fats (total lipids), saturated and unsaturated fatty acids, cholesterol, dietary fiber, water content, energy (calories), and key vitamins and minerals (calcium, phosphorus, potassium, sodium, vitamins B-12, D, and others).

Based on figure 3 the feature importance analysis revealed that Vitamin E (alpha-tocopherol), sodium, riboflavin, and Vitamin K (phylloquinone) were among the most influential features in differentiating fermented from non-fermented dairy products. Fatty acids, particularly polyunsaturated fats, retinol, and folate, were also identified as significant nutritional differentiators, indicating unique profiles associated with fermentation processes.
3.3 CORRELATION ANALYSIS
Correlation analysis using Pearson’s correlation coefficient assessed interrelationships among nutritional variables. Strong positive correlations were found between energy density, total fats, and saturated fats, indicating joint increases in these nutrients within dairy products. Negative correlations emerged between water content and dense nutrients, emphasizing the inverse relationship between water content and caloric density.
3.4 MODEL DEVELOPMENT AND TRAINING
Machine learning models, including Random Forest, Support Vector Machines (SVM), and Gradient Boosting (XGBoost), were trained to classify dairy products into fermented or non-fermented categories. Data was randomly partitioned into training (70%) and testing (30%) sets. Model hyperparameters were optimized through cross-validation.
3.5 MODEL EVALUATION AND VALIDATION
Models were evaluated using accuracy, precision, recall, F1-score, and ROC-AUC. Feature importance analysis identified key nutrients distinguishing fermented from non-fermented dairy foods. Validation ensured robustness and reliability of results, providing actionable insights into the nutritional distinctiveness of fermented dairy products.
The trained machine learning models demonstrated strong performance in distinguishing fermented from non-fermented dairy foods. The classification report (Table 1) illustrates excellent overall accuracy (97%), with precision for fermented products (label 1) at 92%, and recall at 75%, reflecting a high precision but moderate sensitivity due to class imbalance. Non-fermented products (label 0) showed near-perfect precision and recall, underscoring the distinctiveness of their nutritional profiles. These findings affirm the potential for nutritional profiling to effectively categorize dairy products based on fermentation status, providing valuable guidance for targeted dietary recommendations.
| precision | recall | f1-score | support | |
|---|---|---|---|---|
| Non-fermented | 0.98 | 0.99 | 0.99 | 172 |
| fermented | 0.92 | 0.75 | 0.83 | 16 |
| accuracy | 0.97 | 188 | ||
| macro avg | 0.95 | 0.87 | 0.91 | 188 |
| weighted avg | 0.97 | 0.97 | 0.97 | 188 |
Discussion
The findings from this study provide clear evidence of distinct nutritional profiles between fermented and non-fermented dairy products. The application of machine learning models successfully differentiated these two categories based on their nutrient compositions, with specific micronutrients like Vitamin E, sodium, riboflavin, and Vitamin K significantly contributing to this differentiation. The nutritional distinctiveness of fermented dairy products, particularly their enhanced levels of vitamins and beneficial fatty acids, supports the hypothesis that fermentation processes enrich dairy with additional health-promoting properties.
Furthermore, the nutritional markers identified in this study align closely with current nutritional science literature emphasizing the health benefits associated with fermented dairy, such as improved gut health, enhanced immune function, and potential roles in disease prevention. The lower recall observed for fermented products due to class imbalance highlights a limitation of the current analysis and indicates an area for further exploration through expanded datasets or alternative analytical approaches.
These results not only validate the utility of machine learning approaches in nutritional epidemiology but also provide actionable insights that can refine dietary guidelines and personalized nutritional interventions. Integrating these findings into public health initiatives could significantly contribute to chronic disease prevention and management strategies through dietary modification.
The comprehensive analysis conducted in this study underscores the complex interplay between various dietary components, nutritional impacts, and health outcomes, facilitated by advanced machine learning techniques. The integration of diverse data ranging from macronutrients like carbohydrates and proteins to micronutrients such as vitamins and minerals revealed significant correlations that can influence dietary recommendations and public health policies.
Notably, the visual and quantitative analysis through the extended scatter plot matrix highlights both expected and novel relationships among nutritional variables, underscoring the value of multidimensional data analysis in nutritional epidemiology. The categorization of foods in the dataset and their nutritional profiling enable the identification of specific dietary patterns that could mitigate or exacerbate the risk of chronic diseases such as cardiovascular diseases and diabetes. The study also illustrates the potential of personalized dietary interventions, which could be tailored based on individual nutritional needs and health conditions, paving the way for a more targeted approach in medical nutrition therapy.
Conclusion
This research demonstrates the effectiveness of machine learning techniques in identifying and distinguishing the unique nutritional profiles of fermented versus non-fermented dairy products. The identified nutritional characteristics reinforce the nutritional advantages of fermented dairy, validating their importance in dietary recommendations aimed at enhancing health and preventing chronic diseases. Future research should focus on expanding the dataset, addressing class imbalances, and investigating specific health outcomes associated with regular consumption of fermented dairy foods. Overall, these insights underscore the significant potential of precision nutrition informed by robust, data-driven analyses.
Kefir has strong nutritional benefits. It is different from other dairy products and is recommended to be added to daily meals. Studies have shown that kefir is rich in probiotics, vitamins, and minerals, contributing to improved gut health and overall nutrition. Additionally, research indicates that kefir consumption can aid in the prevention of gastrointestinal infections, support the immune system, and improve lactose digestion. These attributes distinguish kefir from other dairy products, underscoring its potential as a functional food in daily diets.
Furthermore, the study highlights the necessity for continuous innovation in dietary assessment methods, including the integration of machine learning and big data analytics, to better understand the nutritional impacts on health. As we move forward, it is crucial to align dietary guidelines with scientific evidence derived from such advanced analyses to ensure they reflect the nuances of modern nutritional science. The potential for these findings to contribute to public health policy, particularly in terms of nutritional recommendations and the development of tailored dietary interventions, is immense and warrants further exploration and validation in clinical and public health contexts.
Conflict of Interest Statement:
None.
Funding Statement:
None.
Acknowledgements:
None.
References:
- S. Morton, D. G. Rhodes, and A. J. Moshfegh, “Food and Nutrient Database for Dietary Studies 2021-2023: Enhancements for National Dietary Surveillance,” Curr Dev Nutr, vol. 8, 2024.
- O. Ojo, “Nutrition and chronic conditions,” 2019, MDPI.
- H. Cena and P. C. Calder, “Defining a healthy diet: evidence for the role of contemporary dietary patterns in health and disease,” Nutrients, vol. 12, no. 2, p. 334, 2020.
- N. G. Forouhi, A. Misra, V. Mohan, R. Taylor, and W. Yancy, “Dietary and nutritional approaches for prevention and management of type 2 diabetes,” Bmj, vol. 361, 2018.
- E. L. Cooper and M. J. Ma, “Understanding nutrition and immunity in disease management,” J Tradit Complement Med, vol. 7, no. 4, pp. 386–391, 2017.
- S. Downer, S. A. Berkowitz, T. S. Harlan, D. L. Olstad, and D. Mozaffarian, “Food is medicine: actions to integrate food and nutrition into healthcare,” bmj, vol. 369, 2020.
- M. M. Ahsan, S. A. Luna, and Z. Siddique, “Machine-learning-based disease diagnosis: A comprehensive review,” in Healthcare, 2022, p. 541.
- S. Russo and S. Bonassi, “Prospects and Pitfalls of Machine Learning in Nutritional Epidemiology,” Nutrients, vol. 14, no. 9, p. 1705, 2022.
- Y. Du and Y. Guo, “Machine learning techniques and research framework in foodborne disease surveillance system,” Food Control, vol. 131, p. 108448, 2022.
- H. Wang, W. Cui, Y. Guo, Y. Du, Y. Zhou, and others, “Machine learning prediction of foodborne disease pathogens: Algorithm development and validation study,” JMIR Med Inform, vol. 9, no. 1, p. e24924, 2021.
- S. M. D. A. C. Jayatilake, G. U. Ganegoda, and others, “Involvement of machine learning tools in healthcare decision making,” J Healthc Eng, vol. 2021, 2021.
- B. Rahmani et al., “Recursive indirect-paths modularity (RIP-M) for detecting community structure in RNA-Seq co-expression networks,” Front Genet, p. 80, 2016.
- Z. Shen, A. Shehzad, S. Chen, H. Sun, and J. Liu, “Machine learning based approach on food recognition and nutrition estimation,” Procedia Comput Sci, vol. 174, pp. 448–453, 2020.
- J. D. Morgenstern, L. C. Rosella, A. P. Costa, R. J. de Souza, and L. N. Anderson, “Perspective: Big data and machine learning could help advance nutritional epidemiology,” Advances in Nutrition, vol. 12, no. 3, pp. 621–631, 2021.
- X. Zhou, L. Chen, and H.-X. Liu, “Applications of Machine Learning Models to Predict and Prevent Obesity: A Mini-Review,” Front Nutr, vol. 9, p. 933130, 2022.
- A. Eetemadi, N. Rai, B. M. P. Pereira, M. Kim, H. Schmitz, and I. Tagkopoulos, “The computational diet: a review of computational methods across diet, microbiome, and health,” Front Microbiol, vol. 11, p. 393, 2020.
- I. Guyon, J. Weston, S. Barnhill, and V. Vapnik, “Gene selection for cancer classification using support vector machines,” Mach Learn, vol. 46, pp. 389–422, 2002.

