VSV-Based Vaccine Strategies for Future Pandemic Preparedness
Preparedness for Future Pandemics Using a Highly Effective Recombinant Vesicular Stomatitis Virus-Based Vaccine Platform Technology: Strategies for Developing Superior Vaccines
Gyoung Nyoun Kim1, Kunyu Wu1, and Chil-Yong Kang1
- Department of Microbiology and Immunology, Schulich School of Medicine and Dentistry, The University of Western Ontario, London, Ontario, Canada N6G 2V4
*Corresponding author: [email protected]
OPEN ACCESS
PUBLISHED: 31 December 2025
CITATION: Kim, GN., Wu, K., et al., 2025. Preparedness for Future Pandemics Using a Highly Effective Recombinant Vesicular Stomatitis Virus-Based Vaccine Platform Technology: Strategies for Developing Superior Vaccines. Medical Research Archives, [online] 13(12). https://doi.org/10.18103/mra.v13i12.7143
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v13i12.7143
ISSN 2375-1924
ABSTRACT
We have developed highly effective, avirulent vesicular stomatitis virus (VSV) vectors to create potent recombinant viral vector-based vaccines. These vaccines induce both humoral and cellular immune responses. For developing a safe and effective viral vector vaccine, we chose VSV because of its broad host range and efficient replication. To enhance the safety of the rVSV vector, we introduced mutations in the M gene to attenuate it, since the VSV M protein is responsible for VSV-induced pathogenesis. The combined mutations G21E, M51R, and L111F (GML) in the M protein of the Indiana serotype of VSV (VSVInd-GML), along with mutations G22E, M48R, and M51R (GMM), and G22E, M48R, M51R, and L110F (GMML) in the M protein of VSV New Jersey serotype (VSVNJ), were used to generate VSVNJ-GMM and VSVNJ-GMML, respectively. The rVSVInd-GML, rVSVNJ-GMM, and rVSVNJ-GMML exhibited reduced cytopathic effects in vitro and across various animal species. Animals injected with up to 5 billion live M gene mutant VSV showed no significant adverse effects, whereas only 1,000 wild-type VSV were enough to kill mice within four days. All future pandemics are likely caused by airborne enveloped RNA viruses that feature surface glycoproteins on their virions. An effective signal peptide at the N-terminus of all glycoproteins is crucial for efficient synthesis, proper processing, glycosylation, intracellular transport, and secretion. Therefore, we replaced the natural signal peptides of viral glycoproteins with a highly efficient signal peptide from honeybee melittin, known for its effectiveness in glycoprotein biosynthesis and intracellular transport. Additionally, we attached the transmembrane domain and cytoplasmic tail of VSV G protein (Gtc) to the C-terminus of the target glycoprotein to enhance its incorporation into pseudotype virions. To prevent vector immunity in booster immunization, we utilised two different serotypes of VSV, along with pseudotype virions carrying the VSV G protein and the glycoproteins of the target virus. These pseudovirions will bind to the VSV receptor, low-density lipoprotein (LDL) receptor, and the receptors of the target surface glycoprotein to initiate infection. The M gene mutants of VSVInd and VSVNJ vectors, which carry surface glycoprotein genes from target viruses, stimulate strong humoral and cellular immune responses and protect animals from challenges with wild-type viruses. These M gene mutant vectors are ideal for developing vaccines to fight future pandemics. This article explains how to develop an effective vaccine for future pandemics caused by enveloped RNA viruses.
Keywords
Vesicular stomatitis virus, vaccine development, recombinant viral vectors, pandemic preparedness, immune response
INTRODUCTION
Humans have encountered numerous pandemics caused by various viruses throughout history. The earliest recorded variola virus pandemic, known as the Antonine Plague, occurred in Iraq between 165 and 180 AD, resulting in the deaths of over 5 million people. The largest viral pandemic was caused by the H1N1 influenza A virus, known as the Spanish Flu, which led to 40 to 50 million deaths in 1918-1919. Other, less severe influenza pandemics, such as the Asian flu and the Hong Kong flu, occurred in 1957 and 1968, respectively. In the early 1980s, human immunodeficiency virus (HIV) infections resulted in acquired immunodeficiency syndrome (AIDS), claiming approximately 36 million lives between 1981 and 2020. Additionally, minor pandemics such as SARS, MERS, Ebola hemorrhagic fever, and swine flu occurred before the most recent COVID-19 pandemic. SARS-CoV-2 has infected over 700 million people worldwide in the past five years and has caused roughly 7 million deaths.
A reliable vaccine supply is essential for preventing future pandemics. Five different vaccine strategies have been used to fight various viral diseases. Traditional methods include attenuated live-virus vaccines and inactivated whole-virus vaccines. Advances in biotechnology have enabled the development of second-generation vaccines, which involve expressing target protein genes from different viruses to produce virus-like particles or pseudotype viruses that carry these antigens. Hepatitis B virus 22-nm virus-like particles (VLPs) have been successfully generated by expressing the hepatitis B virus envelope protein gene. Additionally, human papillomavirus vaccines have been developed using VLPs, which are produced by expressing the virus’s major capsid protein (L1) in various cell lines. These VLPs are non-infectious but closely resemble the native HPV structure, prompting strong immune responses.
Recently, new vaccine strategies have emerged, including messenger RNA (mRNA) vaccines and recombinant viral vector vaccines. The mRNA vaccines against SARS-CoV-2 tend to elicit weaker immune responses, and the protection they offer does not last long. Multiple booster doses of COVID-19 vaccines have failed to fully prevent breakthrough infections, and immunity has lasted only a short time. In contrast, vaccines based on recombinant viral vectors produce strong protective immunity.
Advantages and Limitations of Recombinant Viral Vector-Based Vaccine Approaches
Several viral vectors have been developed to create effective recombinant viral vector-based vaccines, including adenoviruses, integrase-defective lentivirus, poxviruses, insect-specific viruses, and vesicular stomatitis virus.
The vesicular stomatitis virus (VSV) (Figure 1) has a broad host range because it utilises the LDL receptor for infection, produces high yields of virus in vitro, and is relatively easy to recover recombinant viruses through reverse genetics in a T7 bacteriophage RNA polymerase-expressing cell line. VSV is a single-stranded, negative-sense RNA virus with a genome of 11,161 bases. The virion contains five structural proteins. Infection with VSV causes vesicular stomatitis in domestic mammals. There are two main antigenically distinct serotypes: the Indiana serotype (VSVInd) and the New Jersey serotype (VSVNJ). Antibodies against VSVInd do not neutralize VSVNJ, making it suitable for prime-boost vaccine vectors. VSV can produce very high viral yields, exceeding >109 PFU/ml within 18 hours of incubation of infected cells in vitro. The VSV genome can carry up to 6,000 bases of foreign genes and expresses high levels of these inserted genes, forming pseudotype virions.
Introduction to various VSV vector systems
Several VSV vectors have been developed (Figure 2). These include VSV ΔG with or without inserted other viral glycoprotein genes, VSV G gene translocation, VSV N4CT1, VSV Mtd L mutant, and VSV M gene mutants. The VSV ΔG vector has been used to develop a commercial Ebola virus vaccine that prevents Ebola virus infection. However, the yield of the ΔG-vector-based VSV-Ebola vaccine is relatively low, making large-scale production challenging. The VSV G gene translocation vector produced low levels of neutralizing antibodies, despite a virus yield of approximately 3 × 108 PFU/ml. The VSV N4CT1 vector was well tolerated at all tested dose levels and elicited an immune response; however, the vaccine yield was low due to translocation of the N gene between the M and G genes, as well as the gene of interest’s position at the 3’ end of the genome. The VSV Mtd L mutant vector demonstrated safety but elicited low neutralizing antibody responses. In contrast, the M gene mutants of VSVInd and VSVNJ induced high levels of neutralizing antibodies and cytotoxic T-cell responses.
Generation of M gene mutants of VSV as avirulent vaccine vectors
Live recombinant viral vectors for vaccine development must be safe at all doses and capable of efficiently expressing inserted foreign genes. VSV infection causes cytopathic effects (CPE), leading to cell death through the expression of the matrix (M) protein during VSV replication. It has been demonstrated that the VSV M protein inhibits host cell protein expression by blocking cellular transcription and preventing the nuclear export of various cellular RNAs, including spliced mRNAs. The M protein induces CPE through both direct and indirect mechanisms, typically inhibiting host gene expression. Consequently, the M protein is responsible for cell death. The matrix proteins of both VSVInd and VSVNJ consist of 229 amino acids, with approximately 60% sequence homology. The M proteins of both VSVInd and VSVNJ contain methionine at position 51 at the N-terminus.
The amino acid residue in the M protein of VSVInd that is crucial for inhibiting host cell gene expression is methionine at position 51 (M51). The significance of M51 in this process was first indicated by the conditional temperature-sensitive M mutant of VSVInd, tsO82. It has been shown that M51 of the VSVInd M protein plays a key role in disrupting nucleocytoplasmic transport of mRNAs by directly binding to components of the nuclear pore complex, such as Nup98, one of the nucleoporins. Replacing M51 with arginine (R), alanine (A), leucine (L), or aspartic acid (D) diminished the M protein’s ability to inhibit the nuclear export of cellular RNA. Accordingly, we introduced mutations in the M gene of VSVInd and VSVNJ to develop safer and highly effective recombinant VSV vaccine vectors. The combined mutations G21E, M51R, and L111F in the M protein of VSVInd significantly reduced the virus’s burst size by up to 10,000-fold at 37°C without affecting protein expression levels. We also created VSV-New Jersey (VSVNJ) vaccine vectors by combining G22E, M48R, and M51R mutations with or without L110F mutations in the M gene, labelled rVSVNJ(GMM) and VSVNJ(GMML), respectively. BHK21 cells and SH-SY5Y human neuroblastoma cells infected with rVSVInd(GML), rVSVNJ(GMM), and rVSVNJ(GMML) exhibited markedly diminished cytopathic effects in vitro at 37°C. Mice injected with 1 million infectious virus particles of these mutants into the brain showed no neurological dysfunctions or other adverse effects. To improve the stability of the temperature-sensitive mutants, we replaced phenylalanine with alanine, changing all three nucleotides from UUG (leucine) to GCA (alanine). The resulting L111A mutant displayed the temperature-sensitive properties of rVSVInd(GML) and demonstrated increased stability. Twenty consecutive passages of rVSVInd(GML) with an L111A mutation did not revert to leucine (UUG) at position 111 in the M protein gene. All M gene mutants are safe in mice, rats, rabbits, and dogs, and are well tolerated at doses of up to 5 billion PFUs of recombinant vaccine viruses (Table 1). In contrast, only one thousand wild-type VSV can kill mice within four days of infection.
| Modification | Description | Repeated dose toxicity test for dose determination in rats (if required doses) | Single-dose toxicity test in rabbits | General | Neurotoxicity |
|---|---|---|---|---|---|
| VSV ΔG | G gene deleted VSV | GLP, 0.5 x 106 PFU | GLP, 10 x 106 PFU | Non-toxic | Non-toxic |
| VSV G | VSV G gene translocation | GLP, 0.5 x 106 PFU | GLP, 10 x 106 PFU | Non-toxic | Non-toxic |
| VSV N4CT1 | VSV N4CT1 | GLP, 0.5 x 106 PFU | GLP, 10 x 106 PFU | Non-toxic | Non-toxic |
| VSV Mtd L mutant | VSV Mtd L mutant | GLP, 0.5 x 106 PFU | GLP, 10 x 106 PFU | Non-toxic | Non-toxic |
| VSV M gene mutants | VSV M gene mutants | GLP, 0.5 x 106 PFU | GLP, 10 x 106 PFU | Non-toxic | Non-toxic |
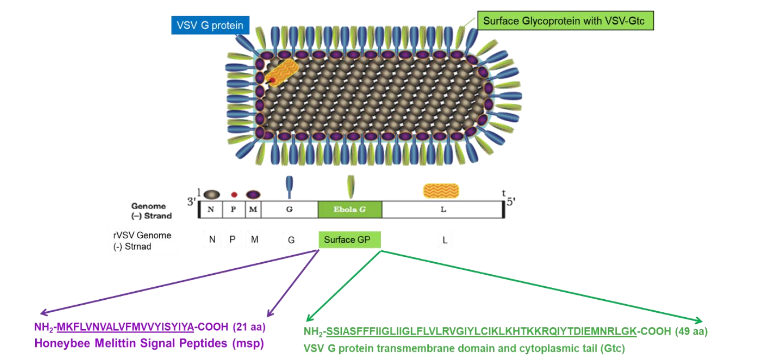
Modification of selected target protein genes by replacing them with efficient glycoprotein signal peptides and attaching the VSV G protein transmembrane domain and cytoplasmic tail (Gtc) at the C-terminus to produce pseudotyped viruses for enhanced immune responses. We designed VSV genomes that incorporate foreign genes to produce avirulent rVSVs expressing the full-length surface glycoprotein of the target antigen (e.g., the SARS-CoV-2 Spike protein, the Zika virus Env protein, or the influenza A virus H1N1 glycoprotein). Including the 21-amino acid honeybee melittin signal peptide (msp) at the N-terminus of these glycoproteins enhanced their expression. Additionally, attaching the VSV G protein transmembrane domain and cytoplasmic tail (Gtc) to the C-terminus of the SARS-CoV-2 Spike protein, Zika virus Envelope protein, or influenza A virus HA protein helped incorporate them into VSV pseudovirions (Figure 3). In immunized mice, rVSV expressing the SARS-CoV-2 Spike gene, the Zika virus Envelope gene, and the influenza A virus HA gene elicited the strongest neutralizing antibody response. Our recombinant vesicular stomatitis virus-based prime-boost vaccination approach induces robust, protective neutralizing antibodies against SARS-CoV-2, and a prime-boost vaccination with the recombinant VSV-Zika virus vaccine completely protected type 1 IFN-receptor knock-out (ifnar -) mice from wild-type Zika virus challenges. Vaccination using our rVSV-based vector vaccine may be the most effective solution in the global fight against future pandemic viruses.
Strategies to overcome vector immunity for booster immunization and for immunizations against other virus vaccines after using the same rVSV vectors.
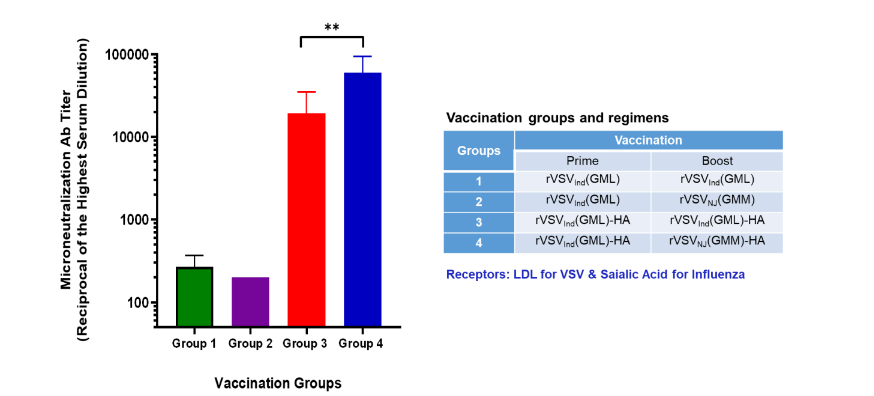
Recombinant viral vector-based vaccine strategies must consider for vector immunity when using the same vector for booster shots or immunization against another virus with the same vector. VSV has two serotypes: Indiana (VSVInd) and New Jersey (VSVNJ). Antibodies against VSVInd do not neutralize VSVNJ, and vice versa, making them suitable for VSVInd prime followed by a VSVNJ boost with the same target antigen gene. This prime-boost vaccination strategy, which employs two different serotype vectors, is highly effective. Additionally, attaching the VSV G protein transmembrane domain and cytoplasmic tail (Gtc) at the C-terminus of the target antigen aids in packaging the target protein onto pseudotype virions. These pseudotype virions can bind to two receptors: the LDL receptor for VSV and the specific receptor for the target virus, such as the ACE2 receptor for SARS-CoV-2 or the sialic acid receptor for Influenza. Figure 4 illustrates the level of neutralizing antibody production after VSVInd-Flu-HA prime vaccination, followed by either the same VSVInd-Flu-HA boost or heterotypic VSVNJ-Flu-HA. The results demonstrate that VSVInd-Flu-HA prime vaccination followed by the same VSVInd-Flu-HA boost induces marginally lower levels of neutralizing antibodies compared to heterotypic VSVNJ-Flu-HA boost vaccination. Boost immunization with the homotypic vector appears to be effective for protective immunization; however, using a heterotypic vector for boosting might be slightly more effective. This suggests the potential to employ either the same serotype VSV or to boost with heterotypic VSV and the target antigen on pseudotype virions.
Potential contributions to controlling future pandemics: An example of producing rVSV pseudotype viruses using the hemagglutinin (HA) gene(s) of the influenza virus for future pandemic preparedness.
VSV vectors have been shown to induce a robust immune response, particularly involving both B-cell and T-cell immunity. This could provide broader protection against influenza. The modified HA protein will be displayed on the pseudotype VSV in a manner that resembles the native viral surface, allowing the immune system to recognise and target influenza virus strains. rVSV-vector based vaccines are generally easier and quicker to produce than traditional egg-based vaccines, potentially enabling faster responses during a pandemic. Overall, our rVSV-based SARS-CoV-2 vaccine produced significantly higher levels of neutralizing antibodies compared to other COVID vaccines.
The World Health Organization recommends two strains of Influenza Virus A and two strains of Influenza Virus B each year to produce a quadrivalent influenza vaccine. The M gene mutant of the rVSV vector system can be used to develop influenza vaccines annually, using strains selected by WHO to ensure vaccine efficacy. Each rVSV can carry two HA genes from the influenza virus, with each HA gene approximately 1,800 nucleotides long. The rVSV vector can accommodate up to 6,000 nucleotides and efficiently express the inserted gene. The M gene mutant vaccine vectors of VSVInd and VSVNJ, along with modifications to the target glycoprotein genes of future pandemic viruses, will support the development of effective recombinant viral vector vaccines to address upcoming pandemics.
Conclusion:
The M gene mutants of VSVInd and VSVNJ vectors, which carry surface glycoprotein gene(s) from target viruses, elicit strong humoral and cellular immune responses and protect animals from wild-type virus challenges. These M gene mutant vectors are ideal for developing vaccines to combat future pandemics.
Conflict of interest statement: The authors declare no conflicts of interest.
Funding statement: Our vaccine research is supported by the Canadian Institute of Health Research (CIHR) and Sumagen Canada Inc.
Acknowledgements: We thank CreoSG of Korea for conducting pre-clinical studies on our M gene mutants of the VSV vectors.
References
- Sampath S, Khedr A, Qamar S, et al. Pandemics Throughout the History. Cureus. Sep 2021;13(9):e18136. doi:10.7759/cureus.18136
- Hilleman MR, Ellis R. Vaccines made from recombinant yeast cells. Vaccine. Jun 1986;4(2):75-6. doi:10.1016/0264-410x(86)90040-x
- Ho JK, Jeevan-Raj B, Netter HJ. Hepatitis B Virus (HBV) Subviral Particles as Protective Vaccines and Vaccine Platforms. Viruses. Jan 21 2020;12(2)doi:10.3390/v12020126
- Cheng L, Wang Y, Du J. Human Papillomavirus Vaccines: An Updated Review. Vaccines (Basel). Jul 16 2020;8(3)doi:10.3390/vaccines8030391
- Buchholz UJ, Finke S, Conzelmann KK. Generation of bovine respiratory syncytial virus (BRSV) from cDNA: BRSV NS2 is not essential for virus replication in tissue culture, and the human RSV leader region acts as a functional BRSV genome promoter. J Virol. Jan 1999;73(1):251-9. doi:10.1128/JVI.73.1.251-259.1999
- Kim GN, Wu, K., and Kang, C.-Y. Establishment of Chinese Hamster ovary (CHO) cells expressing T7 RNA polymerase constitutively. Manuscript in preparation. 2025;
- Lyles DS, Rupprecht, C.E. . Fields Virology vol 1. Rhabdoviridae. Lippincott Williams & Wilkins, a Wolters Kluwer business; 2007.
- An HY, Kim GN, Wu K, Kang CY. Genetically modified VSV(NJ) vector is capable of accommodating a large foreign gene insert and allows high level gene expression. Virus Res. Jan 2013;171(1):168-77. doi:10.1016/j.virusres.2012.11.007
- Whitt MA. Generation of VSV pseudotypes using recombinant DeltaG-VSV for studies on virus entry, identification of entry inhibitors, and immune responses to vaccines. J Virol Methods. Nov 2010;169(2):365-74. doi:10.1016/j.jviromet.2010.08.006
- Miyanohara A. Preparation of vesicular stomatitis virus-G (VSV-G) conjugate and its use in gene transfer. Cold Spring Harb Protoc. Apr 1 2012;2012(4):453-6. doi:10.1101/pdb.prot068528
- Clarke DK, Nasar F, Lee M, et al. Synergistic attenuation of vesicular stomatitis virus by combination of specific G gene truncations and N gene translocations. J Virol. Feb 2007;81(4):2056-64. doi:10.1128/JVI.01911-06
- Lu M, Zhang Y, Dravid P, et al. A Methyltransferase-Defective Vesicular Stomatitis Virus-Based SARS-CoV-2 Vaccine Candidate Provides Complete Protection against SARS-CoV-2 Infection in Hamsters. J Virol. Sep 27 2021;95(20):e0059221. doi:10.1128/JVI.00592-21
- Choi JA, Wu K, Kim GN, et al. Induction of protective immune responses against a lethal Zika virus challenge post-vaccination with a dual serotype of recombinant vesicular stomatitis virus carrying the genetically modified Zika virus E protein gene. J Gen Virol. Apr 2021;102(4)doi:10.1099/jgv.0.001588
- Kim GN, Choi JA, Wu K, et al. A vesicular stomatitis virus-based prime-boost vaccination strategy induces potent and protective neutralizing antibodies against SARS-CoV-2. PLoS Pathog. Dec 2021;17(12): e1010092. doi:10.1371/journal.ppat.1010092
- Marzi A, Ebihara H, Callison J, et al. Vesicular stomatitis virus-based Ebola vaccines with improved cross-protective efficacy. J Infect Dis. Nov 2011;204 Suppl 3(Suppl 3):S1066-74. doi:10.1093/infdis/jir348
- Black BL, Lyles DS. Vesicular stomatitis virus matrix protein inhibits host cell-directed transcription of target genes in vivo. J Virol. Jul 1992;66(7):4058-64. doi:10.1128/JVI.66.7.4058-4064.1992
- Her LS, Lund E, Dahlberg JE. Inhibition of Ran guanosine triphosphatase-dependent nuclear transport by the matrix protein of vesicular stomatitis virus. Science. Jun 20 1997;276(5320):1845-8. doi:10.1126/science.276.5320.1845
- Blondel D, Harmison GG, Schubert M. Role of matrix protein in cytopathogenesis of vesicular stomatitis virus. J Virol. Apr 1990;64(4):1716-25. doi:10.1128/JVI.64.4.1716-1725.1990
- Melki R, Gaudin Y, Blondel D. Interaction between tubulin and the viral matrix protein of vesicular stomatitis virus: possible implications in the viral cytopathic effect. Virology. Jul 1994;202(1):339-47. doi:10.1006/viro.1994.1350
- Kopecky SA, Lyles DS. The cell-rounding activity of the vesicular stomatitis virus matrix protein is due to the induction of cell death. J Virol. May 2003;77(9):5524-8. doi:10.1128/jvi.77.9.5524-5528.2003
- Gill DS, Banerjee AK. Complete nucleotide sequence of the matrix protein mRNA of vesicular stomatitis virus (New Jersey serotype). Virology. Apr 15 1986;150(1):308-12. doi:10.1016/0042-6822(86)90293-x
- Coulon P, Deutsch V, Lafay F, et al. Genetic evidence for multiple functions of the matrix protein of vesicular stomatitis virus. J Gen Virol. Apr 1990;71 ( Pt 4):991-6. doi:10.1099/0022-1317-71-4-991
- Petersen JM, Her LS, Varvel V, Lund E, Dahlberg JE. The matrix protein of vesicular stomatitis virus inhibits nucleocytoplasmic transport when it is in the nucleus and associated with nuclear pore complexes. Mol Cell Biol. Nov 2000;20(22):8590-601. doi:10.1128/MCB.20.22.8590-8601.2000
- von Kobbe C, van Deursen JM, Rodrigues JP, et al. Vesicular stomatitis virus matrix protein inhibits host cell gene expression by targeting the nucleoporin Nup98. Mol Cell. Nov 2000;6(5):1243-52. doi:10.1016/s1097-2765(00)00120-9
- Petersen JM, Her LS, Dahlberg JE. Multiple vesiculoviral matrix proteins inhibit both nuclear export and import. Proc Natl Acad Sci U S A. Jul 17 2001;98(15):8590-5. doi:10.1073/pnas.151240998
- Kim GN, Wu K, Hong JP, Awamleh Z, Kang CY. Creation of matrix protein gene variants of two serotypes of vesicular stomatitis virus as prime-boost vaccine vectors. J Virol. Jun 2015;89(12):6338-51. doi:10.1128/JVI.00222-15

