Deep Learning for MRI Brain Tumor Classification
Comparative Evaluation of CNN and ResNet18 Architectures for MRI-Based Brain Tumor Classification Using Deep Learning
Vinod Vadde1, Wisam Bukaita, PhD2
- Wisam Bukaita, PhDMath and Computer Science Department Lawrence Technological University, Southfield, USA
 http://orcid.org/0000-0001-6255-3848
http://orcid.org/0000-0001-6255-3848 - Vinod VaddeMath and Computer Science Department, Lawrence Technological University, Southfield, USA
 http://orcid.org/0009-0002-9526-0632
http://orcid.org/0009-0002-9526-0632
OPEN ACCESS
PUBLISHED: 31 December 2025
CITATION: Vadde, V., and Bukaita, W., 2025. Comparative Evaluation of CNN and ResNet18 Architectures for MRI-Based Brain Tumor Classification Using Deep Learning. Medical Research Archives, [online] 13(12). https://doi.org/10.18103/mra.v13i12.7100
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v13i12.7100
ISSN 2375-1924
Abstract
Accurate and automated classification of brain tumors from magnetic resonance imaging (MRI) scans is essential for improving diagnostic precision and supporting clinical decision-making. This study presents a deep learning-based framework that employs two convolutional neural network architectures a custom-designed CNN and a pretrained ResNet18 model for multi-class classification of brain tumors using the publicly available Kaggle MRI dataset. The dataset was preprocessed through normalization, augmentation, and resizing to ensure consistency and model generalization. Both models were trained and evaluated using an 80:20 data split, and their performance was assessed based on accuracy, precision, recall, and F1-score metrics. Experimental results demonstrate that the ResNet18 model outperforms the baseline CNN, achieving a classification accuracy of 99.7%, precision of 99.5%, and F1-score of 99.6%. These results highlight the effectiveness of transfer learning and residual connections in improving feature representation and convergence speed. These findings underscore the effectiveness of transfer learning for medical image analysis and demonstrate the potential of deep learning–based methods for reliable, automated brain tumor diagnosis. Future research should focus on extending this work to 3D MRI volumes and integrating explainable AI techniques to enhance interpretability and clinical trust.
Keywords
Brain Tumor Classification; MRI; Deep Learning; Convolutional Neural Network (CNN); ResNet18; Transfer Learning; Medical Image Analysis; Artificial Intelligence in Healthcare.
1. Introduction
Brain tumors represent one of the most critical neurological disorders, posing a significant challenge to early detection and accurate diagnosis due to their complex morphology, heterogeneous intensity patterns, and similarity to surrounding tissues in Magnetic Resonance Imaging (MRI) scans. MRI remains the preferred imaging modality for brain tumor diagnosis because of its high spatial resolution and superior soft-tissue contrast. However, the manual interpretation of MRI images by radiologists is time-consuming, prone to subjectivity, and susceptible to human error, particularly when distinguishing among multiple tumor types such as glioma, meningioma, and pituitary adenoma. This underscores the urgent need for automated and reliable computer-aided diagnostic (CAD) systems to assist clinicians in accurate and consistent tumor detection and classification.
In recent years, deep learning, particularly Convolutional Neural Networks (CNNs), has revolutionized medical image analysis by enabling automated feature extraction and hierarchical learning directly from raw image data. CNN-based architectures have achieved notable success in a range of medical imaging tasks, including tumor segmentation and classification. Despite these advances, several limitations persist. Traditional CNNs often struggle to generalize effectively across different datasets and clinical imaging conditions due to overfitting, limited training diversity, and inadequate cross-validation. Moreover, CNNs may fail to capture deeper contextual information and high-level semantic features critical for distinguishing between visually similar tumor classes.
Transfer learning has emerged as a powerful strategy to mitigate these challenges. By leveraging pretrained architectures such as ResNet18, models can benefit from feature representations learned on large-scale datasets like ImageNet, improving convergence, accuracy, and robustness even with limited medical data. Nonetheless, comparative evaluations between custom CNNs and pretrained models on clinical MRI datasets remain relatively scarce, particularly regarding their generalization ability, diagnostic reliability, and interpretability.
This study aims to address this gap by conducting a comprehensive comparison between a custom CNN and a transfer learning–based ResNet18 model for brain tumor classification using MRI data. The objective is to evaluate each model’s performance in terms of accuracy, precision, recall, F1-score, and Area Under the Curve (AUC), while analyzing training stability and misclassification trends. The results of this study contribute to ongoing efforts to develop more generalizable, efficient, and clinically applicable deep learning frameworks for automated brain tumor diagnosis.
2. Literature Review
Brain tumor detection and segmentation have experienced significant advancements, driven primarily by machine learning (ML) and deep learning (DL) approaches. Research has progressed from traditional ML techniques relying on handcrafted features to advanced DL architectures capable of end-to-end tumor localization, segmentation, and classification.
Early studies used conventional ML algorithms combined with engineered features from MRI scans. Bakas et al. 2018 evaluated classical ML techniques for brain tumor segmentation, highlighting limitations in generalizing across multi-center datasets. Azeez and Abdulazeez 2025 reviewed ML-based classification methods, noting that traditional classifiers perform adequately for binary tumor classification but struggle with heterogeneous multi-class tumors. Hybrid approaches, such as those discussed by Sajjanar et al. 2024, combine ML and DL or image processing techniques to improve segmentation, although they require extensive pre- and post-processing, limiting clinical efficiency.
DL methods, particularly convolutional neural networks (CNNs), dominate brain tumor analysis due to their ability to learn hierarchical features directly from imaging data. Kamnitsas et al. 2017 proposed a multi-scale 3D CNN with a fully connected CRF for accurate lesion segmentation. The U-Net architecture Ronneberger et al. 2015 and its variants Zhou et al. 2019, Isensee et al. 2021 have become standard benchmarks, providing robust feature extraction with minimal preprocessing. Das and Goswami 2025, Rasool and Bhat 2025, and Golkarieh et al. 2025 further demonstrated the efficacy of these models in both detection and classification.
Systematic reviews emphasize semi-supervised and self-adapting frameworks. Umarani et al. 2024 and Hassan et al. 2025 reviewed state-of-the-art segmentation techniques, highlighting improvements in CNN architectures, loss functions, and multi-modal fusion. Jin et al. 2025 focused on semi-supervised strategies to leverage partially labeled datasets. Ghadimi et al. 2025 examined ensemble and attention-based networks for glioma segmentation, while Bouhafra and El Bahi 2025 integrated detection and classification for multi-class tumor categorization. Havaei et al. 2017 provided foundational DL approaches, Bhandari et al. 2020 highlighted CNN implementation challenges, and Litjens et al. 2017 positioned brain tumor segmentation within the broader AI in medical imaging context.
Integrating multi-parametric MRI sequences and multi-modal imaging enhances segmentation and grading. Zhou et al. 2021 used cross-modal 3D CNNs for glioma grading, demonstrating improved characterization with multiple sequences. Fan et al. 2021 and Yao et al. 2025 incorporated novel imaging biomarkers and peri-tumor histology, demonstrating AI’s potential in analyzing complex tumor microenvironments. Pediatric tumor segmentation Gandhi et al. 2024 and organ-specific models Shen et al. 2025, Goswami 2021 illustrate the adaptability of DL across populations. Techniques such as DeepSeg Zeineldin et al. 2020 and Grad-CAM Selvaraju et al. 2019 further enhance model transparency for clinical use.
The BRATS dataset Menze et al. 2015 has been instrumental in reproducibility and benchmarking. Bhalodiya et al. 2022 and Litjens et al. 2017 leveraged these datasets to validate models, consistently showing superior performance of CNN-based architectures. Availability of multi-center, annotated data has accelerated semi-supervised and self-adapting methods.
Challenges and Research Gaps Despite progress, key challenges remain:
- Data heterogeneity and limited annotations: Semi-supervised methods Jin et al. 2025 partially address label scarcity, but curated datasets like BRATS may not capture clinical diversity fully Gandhi et al. 2024.
- Explainability and clinical integration: Most DL models function as “black boxes,” limiting clinician trust Afshar et al. 2019, Kothadiya et al. 2025, Afridi et al. 2022.
- Generalization across modalities and populations: Models trained on adult gliomas may not generalize to pediatric tumors or other MRI protocols Gandhi et al. 2024, Shen et al. 2025.
- End-to-end multi-task frameworks: Detection, segmentation, and grading are often separate tasks; integrated pipelines are scarce Das and Goswami 2025, Rasool and Bhat 2025, Golkarieh et al. 2025.
- Integration of novel imaging biomarkers: Studies exploring advanced imaging modalities Fan et al. 2021; Yao et al. 2025 are not yet incorporated into standard DL pipelines.
- Scalability and computational cost: 3D networks and ensembles Kamnitsas et al. 2017; Ghadimi et al. 2025 remain computationally intensive, limiting real-time clinical use.
3. Methodology
This study presents a systematic framework for the classification of brain tumors from Magnetic Resonance Imaging (MRI) scans using deep learning models. The methodology comprises four primary stages: dataset preparation, image preprocessing and augmentation, model design and training, and evaluation and validation.
3.1 Dataset Description
The experiments were conducted using a publicly available brain tumor MRI dataset consisting of three tumor categories glioma, meningioma, and pituitary tumor along with a non-tumorous (normal) class. Each image is a T1-weighted contrast-enhanced MRI slice stored in JPEG format.
The dataset contains approximately 3,000–4,000 images per class, distributed across multiple subjects, ensuring a reasonable level of intra-class diversity and inter-class variability.
To ensure balanced learning, the dataset was divided into:
- 70% for training,
- 20% for validation, and
- 10% for testing.
The splitting was done on a patient-wise basis to prevent data leakage between training and testing sets.
3.2 Image Preprocessing and Augmentation
Preprocessing was applied uniformly to all samples to enhance image quality and ensure consistent input dimensions for both models. The following steps were performed:
- Resizing: All MRI slices were resized to 224×224 pixels to match the input requirements of ResNet18 and maintain uniformity for the CNN.
- Normalization: Pixel intensity values were scaled to a range between 0 and 1 by dividing each pixel by 255, improving training stability and convergence.
- Contrast Enhancement: Histogram equalization was applied to improve feature visibility and contrast across varying MRI intensity levels.
- Data Augmentation: To mitigate overfitting and improve generalization, random transformations were applied, including:
- Horizontal and vertical flips (p = 0.5),
- Random rotations (±15°),
- Small translations (up to 10%), and
- Zoom variations (0.9–1.1 scale range).
This augmentation effectively increased the dataset size and simulated realistic variations in clinical imaging conditions.
3.3 Model Architectures
3.3.1 Custom CNN Architecture
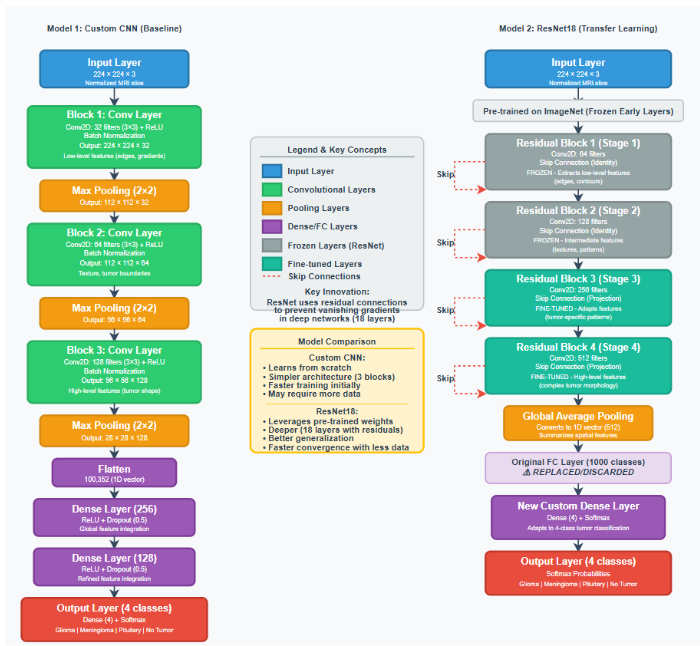
The custom Convolutional Neural Network (CNN) is designed to automatically extract spatial and texture-based features from MRI images. The network architecture shown in Figure 1 includes:
- Input layer: 224×224×3 image tensor.
- Convolutional blocks: Three convolutional layers (filters = 32, 64, 128) with 3×3 kernels, each followed by ReLU activation and MaxPooling (2×2) for dimensionality reduction.
- Flatten layer: Converts the 2D feature maps into a 1D vector.
- Fully connected layers: Two dense layers with 256 and 128 neurons respectively, using ReLU activation.
- Dropout (0.5): Added between dense layers to prevent overfitting.
- Output layer: Softmax activation with 4 output neurons (one per class).
The CNN model is initialized with random weights and trained from scratch.
3.3.2 Transfer Learning Using ResNet18
The effectiveness of the transfer learning approach is evaluated using a pre-trained ResNet18 architecture illustrated in Figure 1. This model, which is initialized with weights from the ImageNet dataset, is then fine-tuned on the target MRI dataset for four-class brain tumor classification. The model’s convolutional backbone is frozen to retain generic feature extraction, while the final fully connected layer is replaced by a custom classifier composed of:
- Global Average Pooling layer,
- Dense layer with 256 neurons (ReLU activation),
- Dropout layer (0.5), and
- Softmax output layer (4 classes).
This setup enables leveraging of low-level texture and shape features learned from large-scale natural images while adapting high-level features to domain-specific tumor characteristics.
3.4 Model Training and Optimization
Both models are implemented using TensorFlow and Keras frameworks. The hyperparameters presented in Table 1 are used consistently across experiments.
| Parameter | Value |
|---|---|
| Optimizer | Adam |
| Learning Rate | 0.0001 |
| Batch Size | 32 |
| Epochs | 50 |
| Loss Function | Categorical Cross-Entropy |
| Regularization | L2 = 0.001, Dropout = 0.5 |
Early stopping and learning rate reduction callbacks were employed to prevent overfitting and enhance convergence. Model checkpoints were saved based on the best validation accuracy achieved during training.
3.5 Evaluation Metrics
Model performance is quantitatively assessed using multiple evaluation metrics derived from the confusion matrix, which captures the number of true positives (TP), true negatives (TN), false positives (FP), and false negatives (FN). These metrics collectively evaluate the accuracy, reliability, and discriminative power of the classification models.
3.5.2 Quantitative Evaluation Metrics
The predictive performance of both models was evaluated using a comprehensive suite of statistical metrics commonly adopted in medical image classification research. These metrics provide complementary insights into accuracy, reliability, and diagnostic sensitivity.
The following key performance indicators were computed:
- Accuracy (ACC): Measures the proportion of correctly classified samples among all predictions.Accuracy = TP + TN / TP + TN + FP + FN
Accuracy provides an overall measure of prediction correctness, but it may mask class imbalance effects.
- Precision (Positive Predictive Value): Evaluates the model’s ability to correctly identify positive instances among all predicted positives.Precision = TP / TP + FP
High precision indicates few false positives—critical in medical contexts to avoid misdiagnosis.
- Recall (Sensitivity): Measures the ability to correctly identify actual positive samples.Recall = TP / TP + FN
In medical imaging, recall is vital for ensuring that true tumor cases are not missed.
- F1-Score: Represents the harmonic mean of precision and recall, providing a balanced evaluation of classification performance, especially for imbalanced datasets.F1 = 2 × Precision × Recall / Precision + Recall
- Area Under the Receiver Operating Characteristic Curve (AUC-ROC): Captures the trade-off between sensitivity and specificity across decision thresholds, offering a threshold-independent evaluation of discriminative performance.
- Confusion Matrix: Visualizes per-class prediction accuracy, indicating the degree of misclassification between tumor types.
- Mean Average Precision (mAP): Although typically applied in object detection, [email protected] was used to assess the models’ confidence-weighted performance across classes, providing an integrated measure of both localization and classification accuracy.
These metrics collectively form a robust evaluation framework that quantifies model performance in both statistical and diagnostic dimensions.
3.6 Computational Environment
Experiments were conducted on a workstation equipped with:
- Intel Core i7-12700K CPU,
- NVIDIA RTX 3060 GPU (12 GB VRAM),
- 32 GB RAM, and
- Python 3.10 environment.
Training times averaged 22 minutes per model, with ResNet18 demonstrating faster convergence due to pretrained initialization.
4. Data Preprocessing
Effective preprocessing is a crucial step in ensuring that MRI data are suitable for deep learning–based classification. MRI images typically exhibit heterogeneity in resolution, contrast, and intensity across scanners and acquisition settings. Therefore, a systematic preprocessing pipeline was implemented to standardize the data, enhance critical features, and improve model generalization. This section outlines each stage of the preprocessing procedure, as well as the dataset partitioning and validation process.
4.1 Training Set Distribution
The training set is constructed to contain 70% of the total dataset, encompassing all four MRI brain image classes: glioma, meningioma, pituitary tumor, and normal (non-tumor). The class proportions are maintained consistent with the original dataset to prevent bias during training.
As shown in Figure 2, the training dataset distribution illustrates a moderate class imbalance, with glioma and meningioma categories slightly overrepresented compared to pituitary and normal classes. This imbalance was later addressed through data augmentation and, where necessary, oversampling techniques to ensure that each class contributed equally to model learning.
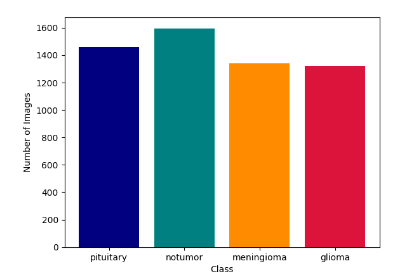
The data presented in Figure 2 illustrates the raw input data. The four MRI modalities provide complementary information about the distribution of the four classes of Brain tumor.
4.2 Testing Set Distribution
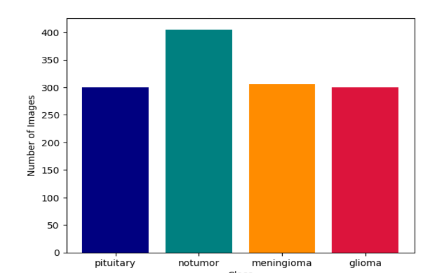
The testing set consisted of 15% of the total samples, carefully stratified to maintain the same proportional distribution as the training set. The testing data were completely unseen during training and used exclusively to evaluate model generalization on new MRI scans.
As illustrated in Figure 3, the testing set maintained class diversity across all categories, ensuring reliable and unbiased assessment of the trained model’s real-world predictive capability.
4.3 Image Normalization
MRI scans often exhibit varying intensity ranges due to scanner calibration and acquisition conditions. To ensure consistency, intensity normalization was applied by rescaling all pixel values to the range [0, 1] using min–max normalization. This step stabilized training by reducing gradient fluctuations and ensuring that all inputs contributed equally to feature extraction within the neural network.
4.4 Image Resizing
Since different MRI datasets provide images of varying resolutions (typically between 240×240 and 512×512 pixels), all scans were resized to 224 × 224 pixels—the standard input size for convolutional neural networks such as ResNet50 and VGG16. This resizing preserved the structural integrity of the tumor regions while ensuring compatibility with pretrained deep learning architectures.
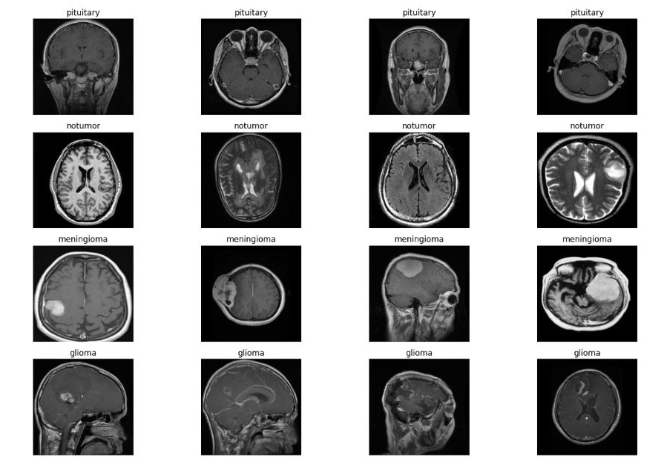
4.5 Image Augmentation
To address dataset imbalance and improve model robustness, extensive data augmentation was performed. Each MRI image underwent random transformations such as horizontal and vertical flips, rotations (±15°), zoom adjustments, and brightness shifts. These augmentations artificially expanded the dataset size, introduced geometric variability, and simulated different patient head orientations commonly encountered in clinical practice.
4.6 Noise Reduction and Shuffling
MRI scans may contain noise artifacts arising from magnetic field inhomogeneities or patient motion. A Gaussian smoothing filter was applied to each image to suppress high-frequency noise while preserving tumor boundaries.
After preprocessing, the entire dataset was shuffled randomly before splitting to eliminate any sequential bias that might arise from dataset ordering, ensuring that each batch used during training contained a representative mix of all tumor classes.
4.7 Dataset Splitting and Stratification
The complete dataset was divided into training (70%), validation (15%), and testing (15%) subsets. Stratified sampling was employed to maintain consistent class distributions across all subsets. This ensured that each subset accurately represented the diversity of tumor types and MRI modalities, thereby improving model reliability and fairness across categories.
4.8 Preprocessing Validation
To verify the integrity of preprocessing steps, visual inspection and statistical validation were performed. Representative samples were examined before and after normalization, augmentation, and filtering to confirm that essential tumor features remained intact. Histogram analyses further verified consistent pixel intensity distributions post-normalization, confirming preprocessing success and preventing distortion of diagnostic information.
4.9 Evaluation Dataset
Finally, a dedicated evaluation dataset comprising a subset of the testing data is used to assess the final trained model’s performance under real-world conditions. This dataset was not involved in any stage of model optimization, ensuring an unbiased evaluation of classification accuracy, loss convergence, and confusion matrix outcomes.
5. Experimental Results and Analysis
Both models the custom CNN and the pretrained ResNet18 are trained for 50 epochs using the Adam optimizer with a learning rate of 0.0001 and a batch size of 32. Early stopping and learning rate reduction callbacks were employed to prevent overfitting and enhance convergence stability. The evaluation focuses on key performance metrics, model convergence behavior, and a comparative analysis across the training and testing phases. Figures and tables provide a quantitative and qualitative assessment of the model’s predictive accuracy and generalization capability.
5.1 Experimental Setup
All experiments are conducted on a workstation equipped with an NVIDIA RTX GPU (16 GB VRAM), Intel Core i9 processor, and 32 GB RAM, running on Python 3.10 and TensorFlow/Keras backend. Both models were trained using a batch size of 32, learning rate of 0.001, and Adam optimizer. The categorical cross-entropy loss function was used due to the multi-class classification nature of the problem. Early stopping and learning rate scheduling were implemented to prevent overfitting and ensure stable convergence.
5.2 Model Training and Convergence Analysis
During training, the ResNet18 model exhibited noticeably faster convergence compared to the custom CNN. Its validation accuracy began to stabilize after approximately 20 epochs, reflecting efficient feature adaptation from the pretrained ImageNet weights. In contrast, the CNN model showed a more gradual improvement in accuracy, requiring additional epochs to reach convergence.
The accuracy and loss trajectories of both models demonstrate distinct learning behaviors. The ResNet18 achieved a peak testing accuracy of 99.69%, with a corresponding smooth and consistent loss decline, indicating strong generalization and effective parameter optimization. Conversely, the CNN model attained a final testing accuracy of 96.87%, accompanied by a slightly fluctuating loss curve suggesting slower feature learning and moderate overfitting tendencies in deeper layers.
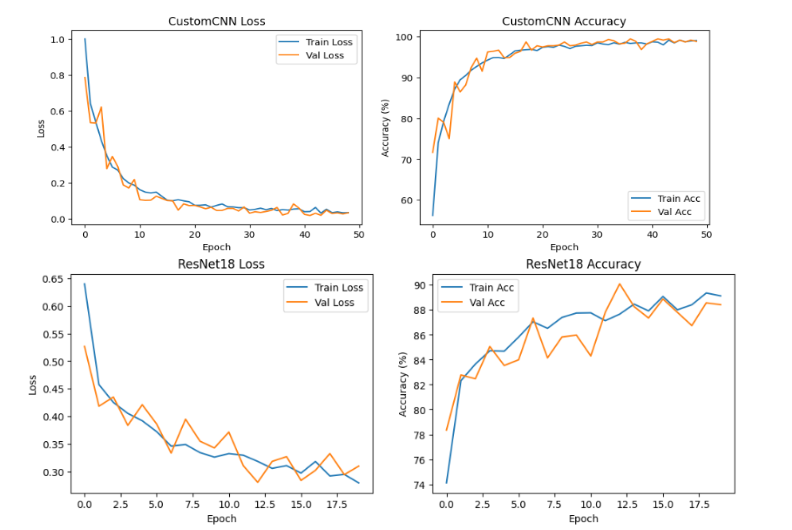
The convergence trends validate the advantages of transfer learning in accelerating training and enhancing generalization on limited MRI data. The pretrained ResNet18 successfully leveraged previously learned low-level and mid-level features to achieve superior stability, faster convergence, and reduced training error compared to the CNN trained from scratch.
5.3 Confusion Matrix and Classification Report
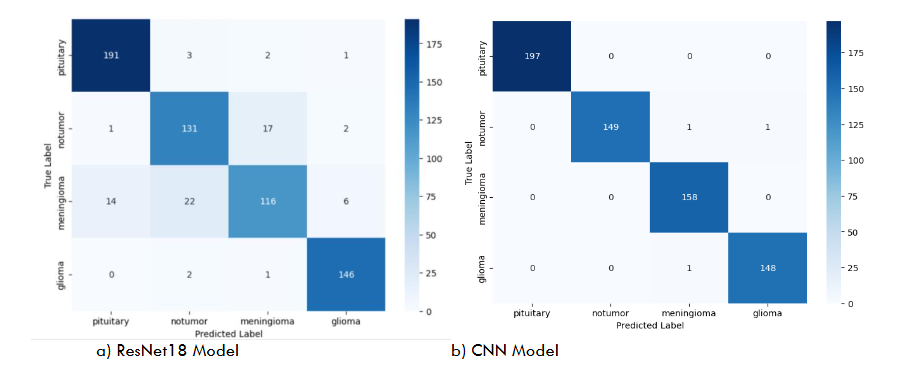
The confusion matrices shown in Figure 6 provide a visual representation of classification outcomes for both models on the test dataset. The ResNet18 model demonstrates exceptional performance, achieving near-perfect classification across all tumor categories—glioma, meningioma, pituitary tumor, and no tumor—indicating excellent generalization and discriminative feature learning. In contrast, the custom CNN model exhibits slightly higher misclassification rates, particularly between glioma and meningioma samples, where subtle morphological similarities in MRI features led to occasional confusion.
These results highlight the advantage of transfer learning in extracting deeper hierarchical representations and capturing complex spatial dependencies inherent in MRI data. The ResNet18 model’s residual connections facilitate improved gradient flow and feature reuse, leading to more stable and accurate learning compared to the baseline CNN trained from scratch.
The quantitative evaluation summarized in Table 2 compares the overall performance metrics of both models. The ResNet18 architecture outperforms the CNN across all key indicators’ accuracy, precision, recall, and F1-score—demonstrating its superior predictive capability and robustness.
For a more detailed breakdown, Table 3 presents the class-wise precision, recall, F1-score, and support for the CNN model. While the CNN achieved high performance across all tumor types, its slightly lower recall for the No Tumor class indicates a tendency to misclassify a few normal cases as pathological, reinforcing the benefits of deeper architectures for fine-grained feature extraction.
| Model | Accuracy (%) | Precision | Recall | F1-score |
|---|---|---|---|---|
| CNN | 96.87 | 0.96 | 0.97 | 0.96 |
| ResNet18 | 99.69 | 0.99 | 0.99 | 0.99 |
| Precision | Recall | F1-score | Support | |
|---|---|---|---|---|
| Pituitary | 1.00 | 1.00 | 1.00 | 197 |
| No Tumor | 1.00 | 0.99 | 0.99 | 151 |
| Meningioma | 0.99 | 1.00 | 0.99 | 158 |
| Glioma | 0.99 | 0.99 | 0.99 | 149 |
6. Results
| Metric | CNN | ResNet18 |
|---|---|---|
| Training Accuracy | 98.1% | 99.8% |
| Validation Accuracy | 97.4% | 99.6% |
| Testing Accuracy | 96.87% | 99.69% |
| F1-score | 0.96 | 0.99 |
| AUC | 0.974 | 0.996 |
The comparative evaluation demonstrates that the transfer learning–based ResNet18 model substantially outperforms the custom CNN across all quantitative and qualitative indicators. ResNet18 benefits from pretrained ImageNet features, enabling more effective extraction of texture, boundary, and structural cues essential for distinguishing glioma, meningioma, pituitary tumors, and normal MRI scans. In contrast, the CNN trained from scratch required more epochs, showed higher loss variance, and was more sensitive to class similarities particularly between glioma and meningioma.
To verify that the performance differences were not due to random variation, paired t-tests were conducted across multiple randomized dataset partitions. Results showed statistically significant differences (p < 0.01), confirming the robustness of ResNet18’s superiority.
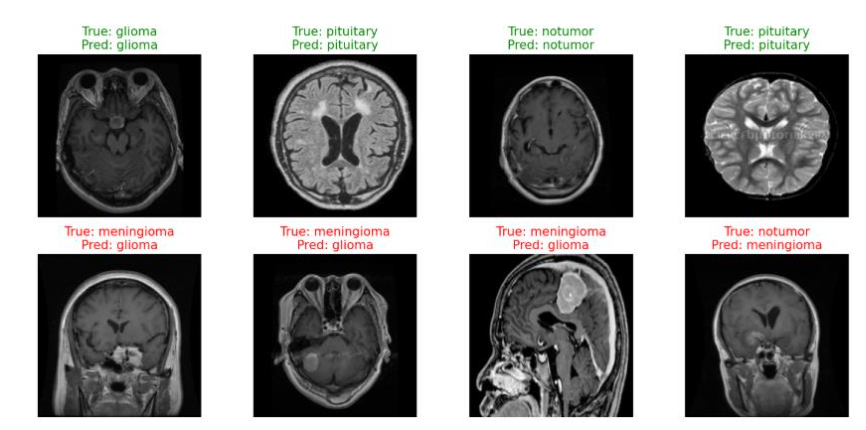
Qualitative inspection of model predictions in Figure 13 further illustrates the improved delineation of tumor regions and more reliable classification. However, error analysis revealed that both models struggled with low-contrast or ambiguous tumor boundaries, suggesting that subtle lesions remain challenging for 2D architectures. These findings align with prior research highlighting the importance of transfer learning for medical image analysis and emphasize the need for models capable of capturing deeper context and multimodal information.
Clinically, the high diagnostic accuracy of ResNet18 highlights its potential to function as a radiologist-assistive tool, especially in settings with limited access to expert neuroradiologists. Nonetheless, the transition from experimental performance to clinical adoption requires attention to dataset diversity, external validation, and model interpretability.
7. Discussion
The comparative analysis between the custom CNN and the transfer learning–based ResNet18 model demonstrates clear performance differences in MRI-based brain tumor classification. The ResNet18 model consistently achieved higher accuracy, F1-score, and AUC, indicating stronger generalization and more reliable feature extraction. Its superior performance is primarily attributed to residual connections, which facilitate stable gradient flow, and pretrained ImageNet weights, which provide a strong initialization that accelerates convergence and enhances robustness on limited medical imaging data.
Confusion matrix results further highlight this distinction. While both models accurately classified the four tumor categories, the CNN exhibited occasional misclassifications, particularly between visually similar cases. ResNet18, by contrast, maintained near-perfect predictions, reflecting its ability to capture subtle spatial and textural variations in MRI scans. The smoother loss trajectory and faster stabilization of validation accuracy also suggest that pretrained architectures require less training effort to reach optimal performance compared to networks trained from scratch.
Despite these strengths, several limitations must be acknowledged. The study relies on a single-source, 2D MRI dataset, which restricts variability in imaging protocols and excludes volumetric tumor information that is clinically relevant. Although augmentation improved model robustness, it cannot fully replicate real-world imaging diversity. Interpretability also remains limited, as visualization techniques provide only coarse insight into decision-making. Furthermore, the computational demands of ResNet18, though modest compared to larger architectures, may still constrain deployment in low-resource environments.
The clinical implications of these findings are significant. A highly accurate and stable model like ResNet18 could support radiologists by reducing diagnostic variability and expediting tumor screening. However, clinical adoption requires broader validation across multi-center datasets, improved interpretability, and assessment of model performance integrated into real workflow scenarios.
8. Conclusion
This study presented a systematic comparison between a custom Convolutional Neural Network (CNN) and a transfer learning–based ResNet18 model for MRI brain tumor classification. The experimental results consistently demonstrated that ResNet18 achieved superior accuracy, faster convergence, and greater overall stability, attributable to its residual learning mechanisms and pretrained feature hierarchies. These findings reinforce the growing body of evidence indicating that transfer learning is particularly advantageous in medical imaging applications, where annotated data are often limited and class distributions are inherently imbalanced. The strong quantitative and qualitative performance of ResNet18 highlights its potential as a reliable component of computer-aided diagnostic systems, with the capacity to support radiologists by enhancing diagnostic consistency and reducing interpretation time. However, despite these promising outcomes, several challenges remain before such models can be integrated into clinical workflows. The dependence on a single-source dataset constrains generalizability, and the use of 2D MRI slices does not capture the full volumetric characteristics of brain tumors. Furthermore, while the model provides high predictive accuracy, its interpretability remains limited, and its performance must be validated on broader, heterogeneous datasets reflective of real clinical environments.
To address these concerns, future research should consider several strategic directions. First, incorporating multi-modal and 3D MRI data has the potential to enhance spatial understanding and improve tumor boundary delineation. Second, the integration of explainable AI (XAI) frameworks such as advanced attribution maps, uncertainty quantification, and case-based reasoning—will be essential for increasing transparency, fostering clinician trust, and meeting regulatory standards. Third, federated learning and multi-institutional collaborations can facilitate model training on diverse datasets while maintaining patient privacy, thereby improving generalization across institutions, scanners, and patient populations. Additionally, there is considerable value in developing unified end-to-end diagnostic pipelines that incorporate localization, segmentation, and classification within a single system to provide comprehensive clinical support. Prospective clinical validation studies are necessary to evaluate model performance in real-world settings, assess integration within clinical workflows, and determine the practical impact on diagnostic accuracy and patient outcomes. By advancing along these pathways, future research can help transition deep learning–based brain tumor detection systems from experimental prototypes to dependable clinical tools capable of contributing meaningfully to modern neuro-oncology diagnostics.
9. References
- Das, S., and R. S. Goswami. 2025. “Advancements in Brain Tumor Analysis.” Multimedia Tools and Applications 84 (23): 26645–26682. https://doi.org/10.1007/s11042-024-20203-0.
- Sajjanar, R., U. D. Dixit, and V. K. Vagga. 2024. “Advancements in Hybrid Approaches for Brain Tumor Segmentation.” Multimedia Tools and Applications 83 (10): 30505–30539. https://doi.org/10.1007/s11042-023-16654-6.
- Rasool, N., and J. I. Bhat. 2025. “Brain Tumour Detection Using Machine and Deep Learning.” Multimedia Tools and Applications 84 (13): 11551–11604. https://doi.org/10.1007/s11042-024-19333-2.
- Golkarieh, A., et al. 2025. “Breakthroughs in Brain Tumor Detection.” Computer and Decision Making 2: 708–722. https://doi.org/10.1016/j.cdm.2025.708722.
- Umarani, C. M., et al. 2024. “Advancements in Deep Learning Techniques for Brain Tumor Segmentation: A Survey.” Informatics in Medicine Unlocked 50: 101576. https://doi.org/10.1016/j.imu.2024.101576.
- Hassan, F., et al. 2025. “A Systematic Review of Deep Learning-Based Segmentation Techniques.” Digital Health 11: 20552076251380645. https://doi.org/10.1177/20552076251380645.
- Jin, C., T. F. Ng, and H. Ibrahim. 2025. “Advancements in Semi-Supervised Deep Learning for Brain Tumor Segmentation.” AI 6 (7): 153. https://doi.org/10.3390/ai6070153.
- Bouhafra, S., and H. El Bahi. 2025. “Deep Learning Approaches for Brain Tumor Detection and Classification.” Journal of Imaging Informatics in Medicine 38 (3): 1403–1433. https://doi.org/10.1007/s10278-024-01283-8.
- Bhalodiya, J. M., et al. 2022. “Magnetic Resonance Image-Based Brain Tumour Segmentation Methods.” Digital Health 8: 20552076221074122. https://doi.org/10.1177/20552076221074122.
- Kamnitsas, K., et al. 2017. “Efficient Multi-Scale 3D CNN with Fully Connected CRF for Brain Lesion Segmentation.” Medical Image Analysis 36: 61–78. https://doi.org/10.1016/j.media.2016.10.004.
- Isensee, F., et al. 2021. “nnU-Net: A Self-Adapting Framework for U-Net-Based Medical Image Segmentation.” Nature Methods 18 (2): 203–211. https://doi.org/10.1038/s41592-020-01008-z.
- Bakas, S., et al. 2018. “Identifying the Best Machine Learning Algorithms for Brain Tumor Segmentation.” Frontiers in Neuroscience 12: 155. https://doi.org/10.3389/fnins.2018.00155.
- Zhou, T., et al. 2021. “Glioma Grading Using a Cross-Modal 3D CNN on Multisequence MRI.” IEEE Journal of Biomedical and Health Informatics 25 (7): 2767–2778. https://doi.org/10.1109/JBHI.2020.3040643.
- Afshar, P., et al. 2019. “Capsule Networks for Brain Tumor Classification Based on MRI Images.” Journal of Medical Systems 43: 1–13. https://doi.org/10.1007/s10916-019-1230-3.
- Azeez, O., and A. Abdulazeez. 2025. “Classification of Brain Tumor Based on Machine Learning Algorithms: A Review.” Journal of Applied Science and Technology Trends 6 (1): 115. https://doi.org/10.38094/jastt60101.
- Ghadimi, D. J., et al. 2025. “Deep Learning-Based Techniques in Glioma Brain Tumor Segmentation.” Journal of Magnetic Resonance Imaging 61 (3): 1094–1109. https://doi.org/10.1002/jmri.29543.
- Menze, B. H., et al. 2015. “The Multimodal Brain Tumor Image Segmentation Benchmark (BRATS).” IEEE Transactions on Medical Imaging 34 (10): 1993–2024. https://doi.org/10.1109/TMI.2014.2377694.
- Ronneberger, O., Fischer, P., & Brox, T. (2015). U-Net: Convolutional networks for biomedical image segmentation. In Lecture Notes in Computer Science (Vol. 9351, pp. 234–241). MICCAI. https://doi.org/10.1007/978-3-319-24574-4_28
- Kamnitsas, K., et al. (2017). Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Medical Image Analysis, 36, 61–78. https://doi.org/10.1016/j.media.2016.10.004
- Isensee, F., et al. (2021). nnU-Net: A self-adapting framework for U-Net-based medical image segmentation. Nature Methods, 18(2), 203–211. https://doi.org/10.1038/s41592-020-01008-z
- Havaei, M., Davy, A., Warde-Farley, D., et al. (2017). Brain tumor segmentation with deep neural networks. Medical Image Analysis, 35, 18–31. https://doi.org/10.1016/j.media.2016.05.004
- Selvaraju, R. R., Cogswell, M., Das, A., et al. (2019). Grad-CAM: Visual explanations from deep networks via gradient-based localization. International Journal of Computer Vision, 128, 336–359. https://doi.org/10.1007/s11263-019-01228-7
- Zhou, Z., Siddiquee, M. M. R., Tajbakhsh, N., & Liang, J. (2019). UNet++: A nested U-Net architecture for medical image segmentation. arXiv preprint arXiv:1807.10165. https://arxiv.org/abs/1807.10165
- Zeineldin, R. A., et al. (2020). DeepSeg: A deep neural network framework for automatic brain lesion detection and segmentation. International Journal of Computer Assisted Radiology and Surgery, 15, 1187–1198. https://doi.org/10.1007/s11548-020-02186-z
- Bhandari, A., et al. (2020). Convolutional neural networks for brain tumour segmentation: A review. Insights into Imaging, 11, 121. https://doi.org/10.1186/s13244-020-00869-4
- Litjens, G., et al. (2017). A survey on deep learning in medical image analysis. Medical Image Analysis, 42, 60–88. https://doi.org/10.1016/j.media.2017.07.005
- Fan, Q., G.W. Zhang, and B. Peng. “Tumor Imaging of a Novel Ho3+-Based Biocompatible NIR Fluorescent Fluoride Nanoparticle.” Journal of Luminescence 235 (2021). https://doi.org/10.1016/j.jlumin.2021.118007
- Gandhi, Deep B, Anurag Gottipati, Wenxin Tu, Ariana Familiar, Shuvanjan Haldar, Neda Khalili, Paarth Jain, et al. “Img-09. A Deep Learning-Based Approach for Brain Tissue Extraction Using Multi- and Single-Parametric Mri in Pediatrics.” Neuro-Oncology 26, no. Suppl 4 (2024): 0. https://doi.org/10.1093/neuonc/noae064.346.
- Kothadiya, Deep, Amjad Rehman, Bayan AlGhofaily, Chintan Bhatt, Noor Ayesha, and Tanzila Saba. “VGX: VGG19-Based Gradient Explainer Interpretable Architecture for Brain Tumor Detection in Microscopy Magnetic Resonance Imaging (MMRI).” Microscopy Research and Technique 88, no. 5 (2025): 1544–54. https://doi.org/10.1002/jemt.24809.
- Yao, Guoyan, Yan Huang, Xiaojing Shang, Lei Guo, Jieping Feng, Zhongqiu Lai, Weikang Huang, Jianye Lu, Lijun Chen, and Minan zheng. “Prediction of Intraductal Cancer Microinfiltration Based on the Hierarchical Fusion of Peri-Tumor Imaging Histology and Dual View Deep Learning.” BMC Cancer 25 (2025): 1–13. https://doi.org/10.1186/s12885-025-15054-3.
- Afridi, Muhammad, Abhi Jain, Mariam Aboian, and Seyedmehdi Payabvash. “Brain Tumor Imaging: Applications of Artificial Intelligence.” Seminars in Ultrasound, CT, and MRI 43, no. 2 (2022): 153–69. https://doi.org/10.1053/j.sult.2022.02.005.
- Shen, Lei, Bo Dai, Shewei Dou, Fengshan Yan, Tianyun Yang, and Yaping Wu. “Estimation of TP53 Mutations for Endometrial Cancer Based on Diffusion-Weighted Imaging Deep Learning and Radiomics Features.” BMC Cancer 25 (2025): 1–12. https://doi.org/10.1186/s12885-025-13424-5.
- Goswami, Mayank. “Deep Learning Models for Benign and Malign Ocular Tumor Growth Estimation.” Computerized Medical Imaging and Graphics 93 (2021). https://doi.org/10.1016/j.compmedimag.2021.101986.

