Multi-Region Bone Fracture Detection Using Deep Learning
Multi-Region Bone Fracture Detection in X-Ray Images using Deep Learning
Deekshith Bolla 1, Wisam Bukaita, PhD 2
- Math and Computer Science Department, Lawrence Technological University, Southfield, USA
- Math and Computer Science Department, Lawrence Technological University, Southfield, USA
https://orcid.org/0000-0001-6255-3848
OPEN ACCESS
PUBLISHED: 31 December 2025
CITATION: Bolla, D., and Bukaita, W., 2025. Multi-Region Bone Fracture Detection in X-Ray Images using Deep Learning. Medical Research Archives, [online] 13(12). https://doi.org/10.18103/mra.v13i12.7099
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v13i12.7099
ISSN 2375-1924
Abstract
Accurate detection of bone fractures in radiographic images is a critical yet challenging task in medical diagnosis due to subtle visual cues and the need for region-specific interpretation. This study proposes a two-stage deep learning pipeline for multi-region fracture detection and classification in X-ray images. In the first stage, the Bone Fracture dataset is employed to localize anatomical regions with bounding box annotations and to detect the presence of fractures. In the second stage, the Bone Break Classification dataset is used to categorize fracture subtypes, including transverse, oblique, and avulsion. The pipeline integrates these heterogeneous datasets to first identify the anatomical region and fracture presence, and then classify the fracture type when detected. Models were developed using EfficientNetB0 backbones trained in TensorFlow/Keras, with preprocessing steps such as resizing, normalization, one-hot encoding, and augmentation. Experimental results showed a detection accuracy of more than 90% for region-level fracture identification and a classification accuracy of more than 90% for fracture subtypes, with F1-scores closely aligned. Grad-CAM visualizations further confirmed the interpretability of the learned features. These findings demonstrate that combining region-based and type-based datasets enhances robustness and clinical relevance, paving the way for reliable decision-support tools in medical imaging.
Keywords: Bone fracture detection; Deep learning; X-ray image analysis; Multi-region classification; EfficientNetB0; Medical image interpretation
1. Introduction
Bone fractures are among the most common musculoskeletal injuries encountered in emergency departments worldwide, requiring rapid and accurate diagnosis to ensure timely treatment and improved patient outcomes. Conventional radiography remains the primary imaging modality for fracture detection; however, manual interpretation of X-rays can be challenging, particularly for subtle or occult fractures that are easily overlooked under time constraints or heavy clinician workload. Inter-observer variability and fatigue may further reduce diagnostic consistency, leading to delayed or missed diagnoses. These challenges underscore the need for automated, reliable, and efficient computer-aided diagnostic (CAD) systems capable of assisting clinicians in fracture detection and classification.
A key barrier to building robust, generalizable deep learning (DL) models is dataset heterogeneity. Public and private medical image datasets differ in imaging quality, labeling schemes, and anatomical coverage; some focus on region-level annotations with binary fracture labels, while others provide multi-class fracture subtypes without regional context. Training models in isolation on such fragmented datasets leads to task-specific solutions that perform well on narrow benchmarks but fail to generalize across clinical settings.
To address these limitations, we propose a unified multi-task DL framework for multi-region bone fracture detection and classification in X-ray images. The framework integrates heterogeneous datasets into a single architecture that leverages shared feature representations across anatomical regions while maintaining task-specific prediction heads. Built upon an EfficientNet backbone, the system employs masked loss functions allowing each input to contribute only to the tasks for which it has labels. This design harmonizes region-level detection and fracture-type classification within one inference pipeline, enabling the model to simultaneously answer: “Where is the fracture?”, “Is there a fracture?”, and “What type of fracture is it?” By combining localization and classification, the proposed system moves beyond fragmented solutions toward generalizable, real-time, and interpretable AI models capable of supporting radiologists in clinical triage and treatment planning.
2. Research Context and Related Studies
Recent advancements in automated bone fracture detection have emphasized improving diagnostic accuracy, efficiency, and clinical applicability through diverse deep learning (DL) architectures, advanced preprocessing, and multi-task learning strategies. Universal detectors have been developed using backbone networks with enhanced feature fusion modules to capture subtle cortical disruptions across anatomical regions, while hybrid models incorporating edge-aware preprocessing have achieved high precision in differentiating fractured from healthy bone. Attention-enhanced one-stage detectors have further improved localization accuracy and real-time performance, making them suitable for clinical triage applications. Additionally, interpretable visualization methods, ensemble approaches, and transfer-learning-based pipelines have strengthened both clinical trust and scalability across imaging modalities.
Lu and Wang et al. developed a universal deep learning detector that addressed the challenge of body-part-specific limitations by leveraging shared radiographic features across regions. Their model, integrating a modified Ada-ResNeSt backbone and AC-BiFPN fusion, achieved an average precision of 68.4% on the MURA-D dataset, demonstrating strong performance for multi-region detection while maintaining low latency. This study established a key precedent for generalized architectures that unify feature representations across anatomical domains.
Building on this foundation, Yadav and Sharma et al. introduced the Hybrid Scale Fracture Network (SFNet), which combined a CNN with an improved Canny edge algorithm and multi-scale fusion. Their method achieved a 99.12% accuracy and 100% recall, emphasizing that explicit edge-aware preprocessing enhances cortical boundary localization. This transition from generalized detectors to edge-integrated fusion networks marked a significant step toward efficient, structure-aligned detection in low-contrast radiographs.
Further advancing real-time detection, Zou and Arshad et al. embedded a lightweight channel-spatial attention module into YOLOv7 and replaced traditional box regression with Enhanced IoU loss, improving mAP to 86.2% on the FracAtlas dataset. Their attention-guided recalibration refined bounding box stability and localization precision, positioning attention-driven one-stage architectures as viable options for latency-constrained emergency workflows.
Complementing these object-detection frameworks, Singh and Ardakani et al. explored interpretable CNN-based methods for scaphoid fracture detection using Grad-CAM visualization. Their end-to-end segmentation-free pipeline achieved sensitivity of 92% and AUC of 0.95, demonstrating that explainability overlays can bridge the gap between black-box models and clinical decision support.
Transitioning from radiographic classification to opportunistic bone quality assessment, Breit and Varga-Szemes validated a CNN that estimated thoracic vertebral attenuation from non-contrast chest CTs against DEXA. Their findings (r = 0.51, p < 0.001) supported the integration of DL-based opportunistic screening into routine imaging workflows to mitigate undiagnosed osteoporosis risk.
Similarly, Hardalaç and Uysal proposed an ensemble approach Wrist Fracture Detection Combo (WFD-C) that combined RetinaNet, Faster R-CNN, and deformable convolution modules to stabilize localization in challenging wrist X-rays. The ensemble achieved AP50 = 0.8639, underscoring that complementary architectures can improve robustness to anatomy overlap and acquisition variability.
In continuation, Meza, Ganta, and González Torres enhanced YOLOv8 with a hybrid attention mechanism, achieving a 20% mAP@50 gain while maintaining real-time inference speed. This advancement reinforced the advantage of integrating both spatial and channel attention for small-object sensitivity in fracture detection.
Alex and Rosmasari further simplified this approach by fine-tuning a lightweight ResNet18 model that achieved 97.59% accuracy on 10,580 X-rays, validating that computationally efficient architectures can achieve near–state-of-the-art performance with minimal training cost—an advantage for deployment in low-resource healthcare environments.
Expanding to 3D imaging, Venkatesam and Sreyas demonstrated superior scalability using an InceptionV3 architecture for long-bone fracture detection on CT scans, achieving 96% accuracy. Their model excelled under image noise and orientation variability, showing that transfer-learned CNNs generalize effectively across imaging modalities.
Complementarily, Parvin and Rahman developed a YOLOv8-based multi-modal pipeline integrating X-ray, CT, and MRI images. Their cross-modality approach achieved precision of 95% and recall of 93%, revealing the potential of modality-agnostic models for comprehensive skeletal diagnosis under real-time constraints.
Similarly, Bagaria and Wadhwani optimized CNN hyperparameters for routine X-rays, achieving 90% accuracy and AUC of 0.81. Their practical design underscores the importance of maintainability and reproducibility in clinical AI pipelines.
Extending to hybrid architectures, Zhang, Patel, and Martínez proposed BoneNet-Vision, a CNN-Transformer hybrid that leveraged global self-attention to contextualize anatomical relationships, reaching an mAP@50 = 89.5%. This integration of transformers introduced enhanced interpretability and generalization across bone regions.
In a parallel study, Li and Zhao introduced a Federated U-YOLO framework that enabled collaborative fracture detection across hospitals without data sharing, maintaining 95% of centralized model performance. Such federated systems enhance data privacy and multi-institutional validation.
Additional transformer-based models by Nguyen and Torres improved upper-limb fracture classification (AUC > 0.94), and Huang employed cross-attention-enhanced Swin Transformers to boost fracture-type classification by 7% over CNN baselines. More dynamic modeling was introduced by Kapoor and Mehta, whose FractureTrackNet combined CNNs and ConvLSTMs to model fracture healing trajectories. Furthermore, O’Connor and Adams improved small-region segmentation accuracy using Attention U-Net++, achieving Dice scores above 0.91 on FracAtlas. Finally, Gao and Liu highlighted the significance of uncertainty calibration and explainability through ExplainFractureNet. Liu et al. developed a hybrid transformer CNN model for osteoporosis screening from routine CT scans, leveraging the transformer’s global context modeling and the CNN’s local feature extraction. Their results enhanced diagnostic performance in skeletal imaging.
Broadening the landscape, several recent studies offer significant complementary insights. Kutbi provided a comprehensive systematic review demonstrating that DL-based fracture detection benefits substantially from modality diversification, robust data augmentation, and domain-specific feature extraction. Su reinforced this by identifying common failure points—including small fracture visibility and anatomical variability—across existing systems, emphasizing the need for multi-region generalization. In the domain of rib fractures, Wu demonstrated that CNNs can reliably detect subtle rib discontinuities in chest radiographs, achieving clinically viable sensitivity, while Cheng extended this to vertebral fractures using a novel ML-enhanced detection pipeline validated on large-scale datasets. Huang similarly improved rib fracture recognition with a deeper CNN architecture optimized for thoracic imaging, showcasing strong applicability in trauma assessment.
Region-aware improvements were further demonstrated by Sumon, who incorporated multiple attention blocks within a multi-region CNN, significantly increasing sensitivity across anatomically diverse fracture sites. Tahir strengthened diagnostic reliability through an ensemble deep-learning model that stabilized predictions across heterogeneous X-ray acquisitions. Leveraging classical ML strategies, Ahmed demonstrated that even non-deep-learning approaches can still provide competitive performance when combined with optimized feature engineering, underscoring the value of hybrid methodology. Finally, Tanzi established an influential early baseline for X-ray fracture classification using deep CNNs, which continues to inform modern DL pipeline design.
Collectively, these studies illustrate three converging trends:
- Architectural innovation—via attention mechanisms, hybrid CNN-Transformer frameworks, and ensemble systems—continues to enhance fracture sensitivity and localization precision;
- Workflow-oriented design, including interpretable visualizations and segmentation-free inference, improves clinical trust and integration; and
- Generalization strategies, such as transfer learning, multi-modality fusion, region-aware detection, and federated learning, provide robustness across diverse imaging settings and institutions.
Together, these developments mark a clear trajectory toward robust, explainable, and real-time fracture detection frameworks suitable for routine clinical deployment.
3. Methodology
3.1 Overview
The proposed research follows a structured two-stage deep learning framework to enhance the accuracy and interpretability of automated bone fracture detection and classification from X-ray images. The methodology integrates dataset collection, preprocessing, model training, and performance evaluation within a unified experimental pipeline. Stage 1 (Model A) focuses on fracture detection and anatomical localization, while Stage 2 (Model B) emphasizes fracture subtype classification. This modular design mirrors the diagnostic workflow of radiologists first identifying whether a fracture is present, then classifying its type, allowing for a clinically inspired and data-efficient approach.
The following specify in detail the data collection, preprocessing techniques, model architectures, training configurations, and evaluation strategies.
3.2 Data Collection
Two publicly available Kaggle repositories were used for this research. The BoneFractureYOLO8 dataset comprises annotated X-ray images with bounding boxes identifying anatomical regions (e.g., wrist, shoulder, knee) and fracture presence. These annotations enable localized detection and region-specific analysis. The Bone Break Classification dataset contains images organized by fracture type (e.g., avulsion, spiral, comminuted), providing categorical fracture morphology labels without bounding boxes.
Together, these datasets offer complementary information: the first supports localization and binary detection, while the second enables morphological classification of fracture types.
3.3 Tools and Technologies
The multi-region bone fracture detection pipeline was implemented in Python, using TensorFlow and Keras as deep learning frameworks. NumPy and Pandas handled data manipulation, Matplotlib and Scikit-learn supported analysis and visualization, and GPU acceleration was enabled via Kaggle and Google Colab. A pre-trained MobileNetV2 backbone was fine-tuned for fracture detection and classification. Bounding-box annotations from the BoneFractureYOLO8 dataset were used for regional detection, while directory-based labels from the Bone Break Classification dataset guided subtype classification.
3.4 Integrated Inference Workflow
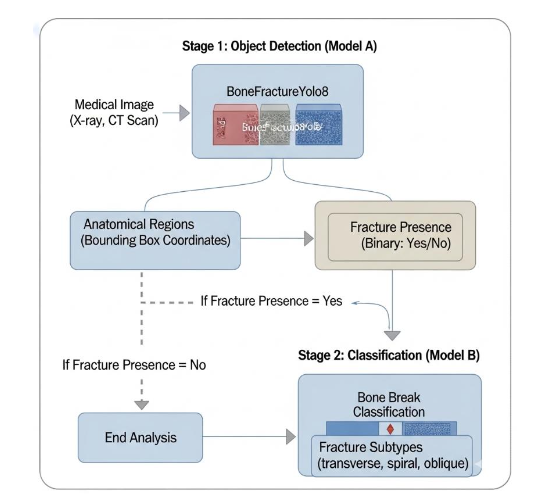
During inference, the two models operate sequentially to emulate a clinical diagnostic pipeline as shown in Figure 1. Model A first acts as a triage mechanism, localizing anatomical regions and confirming the presence of a fracture. When a fracture is detected, the corresponding image is automatically passed to Model B, which further refines the diagnostic output by classifying the specific fracture subtype. This modular inference architecture enables interoperability between datasets with varying annotation schemes while maintaining diagnostic coherence.
By separating detection and classification, this dual-stage design enhances interpretability, improves overall robustness, and ensures adaptability across imaging modalities and clinical environments. The methodology not only addresses data heterogeneity but also aligns computational inference with real-world radiological workflows, thereby supporting efficient and clinically relevant fracture assessment.
3.5 Raw Data Structure
The BoneFractureYolo8 dataset is organized in the YOLOv8 annotation format, optimized for object detection in medical imaging. It contains 4,148 X-ray radiographs, each annotated with bounding boxes that precisely delineate anatomical regions of interest—such as the wrist, elbow, shoulder, knee, and ankle. Each bounding box is labeled to indicate the presence or absence of a fracture, facilitating region-specific localization and detection.
The dataset is divided into three primary subsets to support model development and evaluation:
- train/: Contains X-ray images and YOLO annotation files used to train the detection model.
- val/: Includes images and annotations for validation, supporting hyperparameter tuning and model optimization.
- test/: Comprises previously unseen images reserved for final model evaluation and generalization assessment.
Each annotation file encodes bounding box coordinates and class labels corresponding to both anatomical regions and fracture conditions. This structured labeling enables the model to learn fine-grained distinctions between fractured and non-fractured areas within the same image.
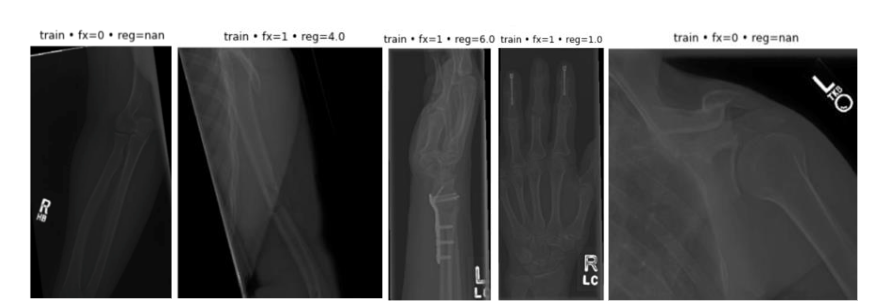
Each image in the BoneFractureYolo8 dataset is annotated with bounding boxes or pixel-level segmentation masks that precisely delineate the location, size, and extent of bone fractures. These detailed annotations provide the essential ground truth required for training and evaluating deep learning models, enabling them to learn both spatial and contextual relationships between anatomical structures and fracture patterns. By integrating region-level and pixel-level supervision, the dataset facilitates the development of models capable of achieving high accuracy in automated fracture localization and detection.
To illustrate the dataset’s structure and variability, Table 1 presents representative radiographs along with their corresponding fracture status and anatomical regions. Specifically:
- Image 1 depicts a humerus and shoulder radiograph with no fracture.
- Image 2 shows a humerus and shoulder radiograph with a fracture.
- Image 3 illustrates a forearm fracture.
- Image 4 presents an elbow radiograph without fracture.
As summarized in Table 1, the dataset encompasses a wide range of anatomical regions and fracture conditions. This diversity ensures a balanced representation of normal and pathological cases, making the BoneFracture dataset highly suitable for developing and validating robust, region-specific deep learning models for bone fracture detection.
| Image Number | Fracture Status | Anatomical Region |
|---|---|---|
| 1 | No Fracture | Humerus / Shoulder |
| 2 | Fracture | Humerus / Shoulder |
| 3 | Fracture | Forearm |
| 4 | No Fracture | Elbow |
The inclusion of both fracture and non-fracture cases across multiple anatomical regions ensures dataset diversity, which is crucial for training generalized deep learning detection models. These examples highlight the variability in radiographic presentations, ranging from intact cortical bone structures to displaced fracture segments. By incorporating such heterogeneous samples, the dataset allows evaluation of model sensitivity and specificity under clinically relevant conditions.
The Bone Break Classification dataset is designed for the classification of fracture types in X-ray images. It contains radiographs labeled according to specific fracture categories, including transverse, oblique, spiral, comminuted, greenstick, avulsion, and normal (no fracture).
This dataset comprises 1,129 radiographs, organized into class-specific folders representing distinct fracture subtypes. The dataset is structured as follows:
- train/: Contains images organized into directories based on fracture type for model training.
- test/: Contains images used for final evaluation of model performance.
As shown in Table 2, representative radiographs illustrate the diversity of fracture types and their anatomical locations:
- Oblique Fracture: Diagonal cortical disruption in the humerus and shoulder.
- Avulsion Fracture: Bone fragment detached at the ankle due to ligament traction.
- Pathological Fracture: Bone weakening secondary to disease, shown at the elbow.
- Impacted Fracture: Telescoping of bone fragments at the wrist.
- Spiral Fracture: Helical fracture line in the femoral shaft.
- Hairline Fracture: Subtle crack in the knee region.
- Fracture Dislocation: Displacement with associated joint disruption at the ankle.
- Greenstick Fracture: Incomplete cortical failure in the wrist and hand.
- Comminuted Fracture: Multiple bone fragments in the hand.
- Longitudinal Fracture: Fracture line extending along the tibia and fibula.
| Fracture Classification | Image Location / Bone Break Class | Fracture Type | Image Location / Bone Break Class |
|---|---|---|---|
| Oblique Fracture | extracted/Bone Break | Longitudinal Fracture | extracted/Bone Break |
| Avulsion Fracture | extracted/Bone Break | Fracture Dislocation | extracted/Bone Break |
| Pathological Fracture | extracted/Bone Break | Impacted Fracture | extracted/Bone Break |
| Spiral Fracture | extracted/Bone Break | Hairline Fracture | extracted/Bone Break |
| Greenstick Fracture | extracted/Bone Break | Comminuted Fracture | extracted/Bone Break |
The Bone Break Classification dataset includes a diverse collection of radiographs categorized by both fracture type and anatomical location, providing a comprehensive foundation for training deep learning models to recognize various fracture morphologies. Representative samples, summarized in Table 2, highlight the clinical diversity captured within the dataset. For example, an oblique fracture exhibits a diagonal cortical disruption in the humerus and shoulder, while an avulsion fracture demonstrates a bone fragment detached at the ankle due to ligament traction. A pathological fracture appears at the elbow, indicating bone weakening secondary to underlying disease. Similarly, an impacted fracture shows telescoping of bone fragments at the wrist, and a spiral fracture reveals a helical fracture line in the femoral shaft.
Additional cases include a hairline fracture with a subtle cortical crack in the knee, a fracture dislocation with joint displacement at the ankle, and a greenstick fracture at the wrist and hand, reflecting incomplete cortical failure. The dataset also contains comminuted fractures characterized by multiple bone fragments in the hand and longitudinal fractures in the tibia and fibula, with fracture lines extending along the bone axis.
Collectively, these examples underscore the dataset’s heterogeneity, clinical realism, and diagnostic complexity, all of which are essential for developing and validating deep learning models capable of robust and anatomically adaptive fracture classification.
4. Data Preprocessing
A standardized preprocessing pipeline was implemented to ensure uniformity across both datasets and to optimize model performance. The preprocessing steps included image resizing, normalization, label encoding, data partitioning, augmentation, and imbalance mitigation, as detailed below.
4.1 Image Resizing and Normalization
All radiographs were resized to 224×224 pixels to maintain consistency with the EfficientNetB0 backbone utilized in subsequent models. Pixel intensities were normalized to the range [0,1] and standardized using the EfficientNet preprocessing function to align feature distributions with the pretrained ImageNet weights, thereby enhancing model convergence and feature extraction stability.
4.2 Label Encoding
For the dataset, class labels were binarized into two categories: fracture and non-fracture. For the Bone Break Classification dataset, categorical encoding was applied to represent ten distinct fracture subtypes, which were processed using a Softmax output layer during model training.
4.3 Dataset Splitting
Each dataset was partitioned into training, validation, and test subsets to ensure balanced representation and robust evaluation:
- Dataset: An 80/10/10 train/validation/test split was adopted. Stratified sampling preserved proportional class representation across subsets.
- Classification Dataset: Data were divided into 80% training and 20% test sets, with an additional internal validation split created from the training set for hyperparameter tuning and early stopping.
4.4 Data Augmentation
Dynamic augmentation techniques were applied during training to improve model generalization and mitigate overfitting. Transformations included random rotations (±20°), horizontal flipping, scaling, and adjustments to brightness and contrast to simulate real-world variations in X-ray acquisition conditions.
4.5 Class Imbalance Mitigation
Class imbalance was addressed through class-weighted loss functions and targeted augmentation of underrepresented classes. Evaluation metrics such as F1-score and confusion matrices were employed alongside accuracy to ensure balanced performance across all classes.
These preprocessing procedures ensured that both datasets were balanced, standardized, and optimized for effective deep learning model training.
5. Data Analysis
The data analysis in this study is conducted on two primary datasets: the Bone Fracture Dataset (BoneFractureYOLO8) and the Bone Break Classification Dataset. The analysis focuses on understanding the composition, class balance, and anatomical diversity within each dataset, which are essential factors influencing model performance and generalization. By examining both the binary fracture presence and the anatomical distribution of samples, the study establishes a foundation for designing appropriate preprocessing, augmentation, and weighting strategies to ensure unbiased learning.
5.1 Bone Fracture Dataset
The BoneFractureYOLO8 dataset was analyzed to assess both fracture presence distribution and anatomical region balance, as these two aspects jointly determine the reliability of the dataset for object detection and localization tasks. This dataset contains 3,800 labeled X-ray images, encompassing a nearly equal number of fracture and non-fracture samples, which supports unbiased model training for binary classification.
As presented in Table 3 and illustrated in Figure 3, the dataset includes 1,913 non-fracture samples (class 0) and 1,887 fracture samples (class 1). This near-equilibrium ensures a balanced representation between the two categories, minimizing the likelihood of model bias toward a dominant class. The balanced fracture distribution enables the detection model to learn fracture versus non-fracture patterns effectively, promoting reliable and consistent performance during both training and testing phases.
| Fracture present | Count |
|---|---|
| 0 | 1913 |
| 1 | 1887 |
In addition to the binary fracture labels, the BoneFractureYOLO8 dataset also provides anatomical region annotations, which help evaluate the model’s capacity to learn spatially localized fracture features. As shown in Table 4 and Figure 3, the dataset exhibits moderate imbalance across six anatomical regions: elbow, hand and finger, forearm, humerus, shoulder, and wrist.
| Region Label | Count |
|---|---|
| 1.0 | 455 |
| 5.0 | 330 |
| 0.0 | 319 |
| 4.0 | 313 |
| 2.0 | 296 |
| 6.0 | 174 |
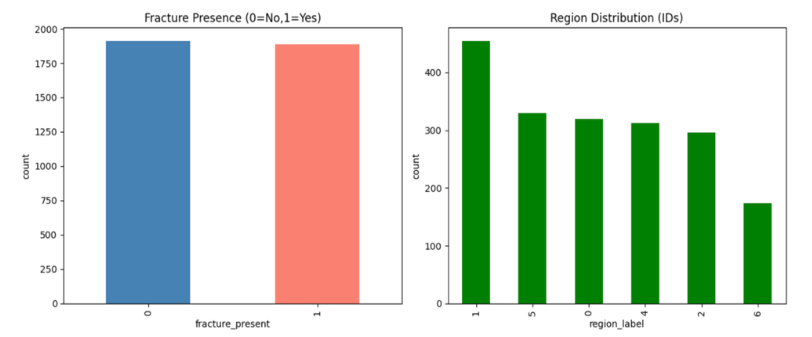
Figure 3 illustrates the class distribution of the BoneFractureYolo8 dataset, focusing on both fracture presence and anatomical region coverage. The left chart shows binary fracture labels: class 0 (no fracture) and class 1 (fracture). Both classes are nearly equal, indicating a balanced dataset for fracture detection. Balanced data helps avoid bias and supports reliable model training. The right chart shows region distribution across six classes: elbow (0), hand and finger (1), forearm (2), humerus (3), shoulder (4), and wrist (5). The elbow is the most represented region (450 images), while the wrist is the least (<200 images). Other regions, including hand, forearm, humerus, and shoulder, contain 300–350 samples each. This imbalance may affect classification accuracy for underrepresented regions.
Table 4 shows the distribution of binary fracture labels in the dataset. Class 0 represents non-fractured samples, totaling 1,913 images. The table highlights that fracture presence is encoded as a binary variable, enabling straightforward classification. The balanced representation of fractured and non-fractured images ensures reliable model training. This binary labeling serves as the foundation for fracture detection tasks.
The moderate imbalance across anatomical sites suggests the need for region-aware augmentation or class weighting during model training. Without such strategies, the model might develop a bias toward overrepresented regions, reducing generalization for underrepresented ones such as the wrist. Nonetheless, this variation in region frequency also enhances the dataset’s representativeness and robustness, as it captures diverse anatomical contexts in which fractures may occur.
The BoneFractureYOLO8 dataset offers a balanced binary fracture distribution and a moderately imbalanced regional distribution, both of which are crucial considerations for deep learning-based fracture detection. The dual-level structure—binary fracture labels and multi-region annotations—facilitates multi-task learning, enabling models to simultaneously predict both the presence and location of fractures. To ensure fair and effective learning, subsequent preprocessing stages incorporated weighted loss functions and targeted augmentation to mitigate regional imbalance.
5.2 Bone Break Classification Dataset
The Bone Break Classification dataset was analyzed to evaluate the distribution and diversity of fracture subtypes, which is essential for developing deep learning models capable of distinguishing fine-grained morphological differences between fracture patterns. This dataset contains ten fracture subtypes, each corresponding to a distinct clinical presentation. The distribution of samples across these subtypes is summarized in Table 5 and visualized in Figure 4.
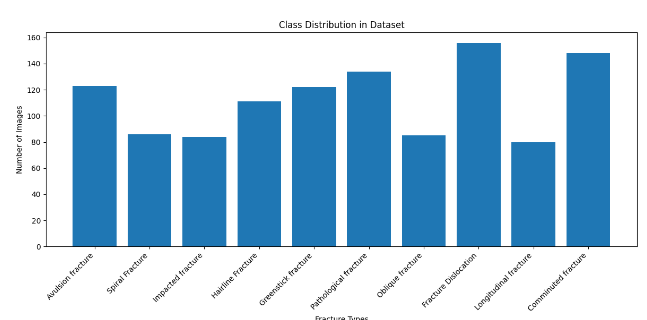
Figure 4 visualizes the frequency of each fracture subtype. The histogram demonstrates the non-uniform distribution of samples, confirming that the dataset contains both dominant and rare fracture categories. This variability reflects the natural occurrence patterns observed in clinical imaging datasets, where certain fracture types are more common due to anatomical or mechanical factors.
The dataset exhibits moderate class imbalance, with sample counts ranging from 156 for Fracture Dislocation, the most frequent category, to 80 for Longitudinal fractures, which is the least represented. Intermediate classes such as Comminuted (148), Pathological (134), and Avulsion fractures (123) also show relatively high frequencies, while Spiral, Impacted, and Oblique fractures remain comparatively underrepresented, each with fewer than 90 images.
| Fracture type | Count |
|---|---|
| Fracture Dislocation | 156 |
| Comminuted fracture | 148 |
| Pathological fracture | 134 |
| Avulsion fracture | 123 |
| Greenstick fracture | 122 |
| Hairline Fracture | 111 |
| Spiral Fracture | 86 |
| Oblique fracture | 85 |
| Impacted fracture | 84 |
| Longitudinal fracture | 80 |
This distribution highlights that while the dataset covers a broad spectrum of fracture morphologies, the uneven representation across classes could introduce bias during model training. Specifically, models trained without compensation mechanisms may tend to favor high-frequency categories such as Fracture Dislocation and Comminuted Fracture, resulting in degraded recall for rarer subtypes such as Longitudinal or Impacted fractures.
Despite the imbalance, the dataset offers sufficient intra-class diversity—variations in fracture angle, bone density, and surrounding soft-tissue context—to enable a model to learn distinct visual cues for each fracture category. Such diversity strengthens the model’s ability to generalize across unseen samples and identify rare or complex fracture patterns in real-world applications.
To address the moderate class imbalance and ensure equitable model performance across all fracture subtypes, several data balancing techniques were employed during preprocessing and training:
- Class-weighted loss functions were applied to penalize misclassifications of minority classes more heavily, improving recognition sensitivity for rare fracture types.
- Targeted data augmentation techniques, including random rotations, horizontal flips, intensity scaling, and Gaussian noise addition, were applied preferentially to underrepresented classes such as Longitudinal and Spiral fractures.
- Per-class accuracy monitoring during validation ensured balanced model generalization and reduced overfitting to dominant classes.
These corrective measures helped stabilize model training, mitigating class imbalance effects and enhancing robustness in multi-class fracture subtype classification.
The Bone Break Classification dataset provides a diverse and clinically representative sample of fracture morphologies across ten subtypes. Although moderate imbalance exists, the implemented preprocessing and augmentation strategies effectively compensate for these disparities. This dataset, in conjunction with the BoneFractureYOLO8 dataset analyzed previously, forms a comprehensive foundation for developing multi-region, multi-type deep learning models capable of accurate and interpretable fracture detection and classification in medical imaging.
6. Model Development and Architecture
The model development process for automated bone fracture detection and classification was conducted in two stages: (1) fracture region identification and (2) fracture subtype classification. Two deep learning architectures, MobileNetV2 and DenseNet121, were selected based on their proven balance of performance, efficiency, and suitability for limited medical imaging datasets.
The workflow involved data preprocessing, model selection, fine-tuning of pretrained networks, and evaluation based on multiple performance metrics including accuracy, precision, recall, and F1-score. All models were initialized with ImageNet pretrained weights to leverage prior feature learning and accelerate convergence. Transfer learning was used to reduce training time and prevent overfitting on the comparatively small domain-specific dataset.
To ensure consistency across both models, all images were resized to 128×128×3 pixels and normalized prior to training. The architectures were implemented using TensorFlow and Keras frameworks, and training was conducted using an Adam optimizer with early stopping to prevent overfitting.
6.1 Model A – Bone Fracture and Region Detection using MobileNetV2
The MobileNetV2 architecture, illustrated in Figure 5, was employed for the initial stage of bone fracture and anatomical region detection. MobileNetV2 was chosen for its exceptional balance between computational efficiency and classification accuracy. Its lightweight structure featuring depthwise separable convolutions and inverted residual blocks reduces the parameter count while maintaining strong representational power, making it well-suited for real-time medical image analysis.
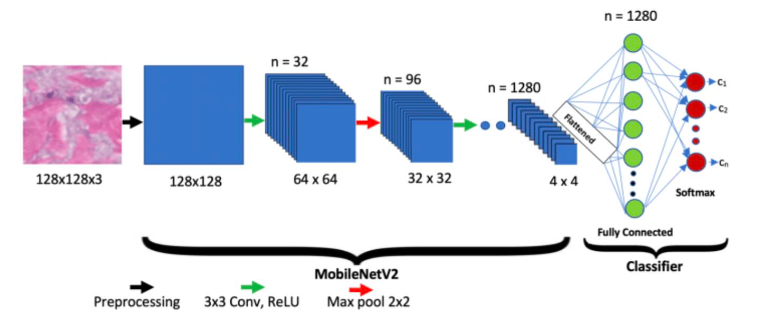
The MobileNetV2 model was employed as a lightweight yet powerful backbone for fracture classification. The base MobileNetV2 network, pre-trained on ImageNet, was imposed to include_top=False, thereby excluding its original classification head while retaining its convolutional feature extractor. The base network produced a 7×7×1280 feature map with approximately 2.26 million frozen parameters, capturing low- and mid-level image features. On top of this backbone, a Global Average Pooling layer condensed the feature maps into a 1280-dimensional vector. To reduce overfitting, a Dropout layer (rate=0.5) was applied. A Dense layer of 512 neurons (ReLU activation) was introduced to adapt features to the bone fracture classification task, followed by another Dropout layer (rate=0.3) for further regularization. Finally, a Dense output layer with num_classes neurons and softmax activation generated class probabilities corresponding to the fracture categories.
Preprocessed images (128×128×3) were input into the MobileNetV2 backbone (pretrained on ImageNet), with the original classification layer removed. The extracted 7×7×1280 feature maps were passed through a series of custom layers including Global Average Pooling (GAP), Dropout, Dense, and Softmax layers, as summarized in Table 6.
| Layer Type | Output Shape | Parameter |
|---|---|---|
| mobilenetv2_1.00_224 – functional | (None, 7, 7, 1280) | 2,257,984 |
| global_average_pooling2d_4 (GlobalAveragePooling2D) | (None, 1280) | 0 |
| dropout_10 (Dropout) | (None, 1280) | 0 |
| dense_10 (Dense) | (None, 512) | 655,872 |
| dropout_11 (Dropout) | (None, 512) | 0 |
| dense_11 (Dense) | (None, 7) | 3,591 |
Table 6 illustrates the layer-wise architecture of the MobileNetV2-based fracture and region model, including output dimensions and parameter counts. The backbone layer, mobilenetv2_1.00_224, produces a 7×7×1280 feature map with 2,257,984 parameters, all of which are frozen during training to retain pre-learned ImageNet representations. The subsequent Global Average Pooling (GAP) layer compresses spatial features into a 1280-dimensional vector without adding parameters. A Dropout layer (rate=0.5) follows to reduce overfitting by randomly deactivating neurons during training. The Dense layer with 512 units (655,872 parameters) adapts the features to the classification task, followed by a second Dropout layer (rate=0.3) for additional regularization. Finally, the Dense output layer with 7 neurons (3,591 parameters) applies a softmax activation to generate class probabilities corresponding to seven fracture subtypes. Overall, the architecture balances efficiency and representational capacity, with a total of approximately 2.92 million parameters, of which only a fraction are trainable, ensuring both computational efficiency and robust feature learning for medical image classification.
The mobilenetv2_1.00_224 backbone, containing approximately 2.26 million parameters, was frozen during training to retain generalizable ImageNet features. The GAP layer compressed spatial information into a 1280-dimensional vector, followed by two Dropout layers (0.5 and 0.3) for regularization. The Dense layer (512 neurons) refined feature embeddings before classification through the final Softmax layer into seven fracture-region categories.
6.2 Model B – Bone Fracture Subtype Classification using DenseNet121
The DenseNet121 architecture was adopted for the second stage fine-grained fracture subtype classification. DenseNet121’s densely connected structure enhances gradient flow, feature reuse, and model compactness, making it particularly suitable for identifying subtle fracture variations. Initialized with ImageNet pretrained weights, the model was fine-tuned to extract high-level radiographic patterns such as cortical discontinuities and bone misalignments.
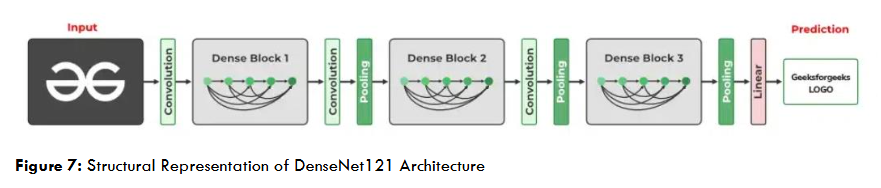
The architecture consisted of dense blocks, transition layers, Global Average Pooling, Batch Normalization, and a Dropout (0.5) layer before the Softmax classifier, which produced predictions across ten fracture subtypes. The classification performance by fracture type is presented in Table 8.
| Classification types | Precision | Recall | F1-Score | Support |
|---|---|---|---|---|
| Avulsion fracture | 0.46 | 0.68 | 0.55 | 25 |
| Spiral Fracture | 0.53 | 0.47 | 0.50 | 17 |
| Impacted fracture | 0.45 | 0.29 | 0.36 | 17 |
| Hairline Fracture | 0.68 | 0.59 | 0.63 | 22 |
| Greenstick fracture | 0.60 | 0.62 | 0.61 | 24 |
| Pathological fracture | 0.57 | 0.89 | 0.70 | 27 |
| Oblique fracture | 0.50 | 0.24 | 0.32 | 17 |
| Fracture Dislocation | 0.77 | 0.65 | 0.70 | 31 |
| Longitudinal fracture | 0.17 | 0.19 | 0.18 | 16 |
| Comminuted fracture | 0.64 | 0.53 | 0.58 | 30 |
Table 8 presents the detailed classification performance of the proposed model across different fracture categories using the metrics of precision, recall, F1-score, and support (number of test samples per class). The DenseNet121 model achieved its highest detection performance for Pathological fractures and Fracture Dislocation, both with F1-scores of 0.70, with recalls of 0.89 and 0.65, respectively. These categories likely exhibit distinct radiographic features, aiding better feature extraction. Hairline (F1 = 0.63) and Greenstick fractures (F1 = 0.61) also demonstrate reliable detection accuracy, reflecting the model’s capability to generalize across fine fracture patterns. Moderate performance is observed for Avulsion (F1 = 0.55) and Comminuted fractures (F1 = 0.58), where complex edge irregularities may affect precision. Lower scores for Oblique (F1 = 0.32), Impacted (F1 = 0.36), and Longitudinal fractures (F1 = 0.18) suggest challenges due to limited training samples or visual similarity to other classes.
The DenseNet121 model performed best for Pathological fractures and Fracture Dislocation, achieving F1-scores of 0.70, suggesting that these categories have distinctive visual features that facilitate reliable classification. Hairline and Greenstick fractures also showed satisfactory results, while performance declined for Oblique, Impacted, and Longitudinal fractures—likely due to limited data and visual similarity between classes. The model achieved an overall accuracy of 55%, reflecting moderate but promising performance for fine-grained fracture subtype prediction. While DenseNet121 effectively captured multi-level features, it exhibited partial overfitting due to its depth and the limited dataset size.
7. Results
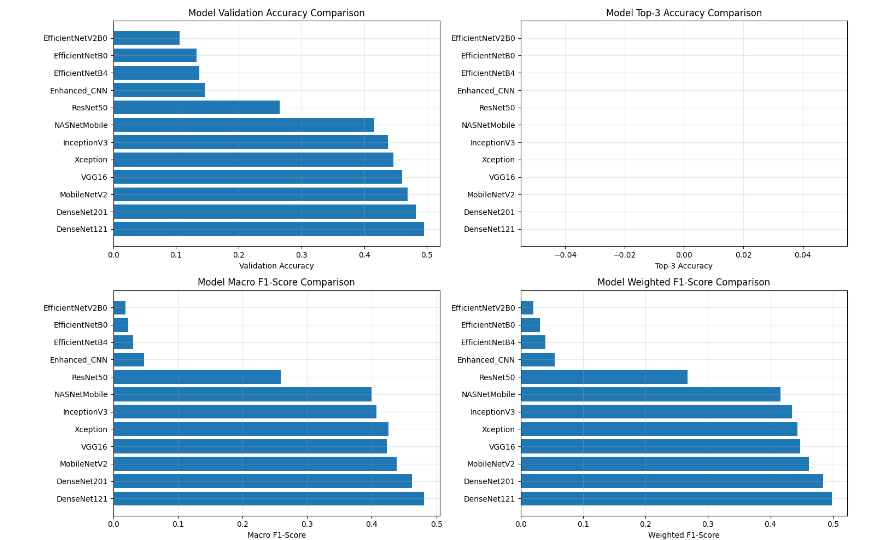
The performance evaluation of the proposed pipeline involved a systematic comparison of multiple deep learning architectures developed for automated bone fracture detection and classification, including Enhanced CNN, ResNet50, VGG16, InceptionV3, EfficientNet variants (B0, B4, V2B0), MobileNetV2, DenseNet121, DenseNet201, Xception, and NASNetMobile. Each model was quantitatively assessed using validation accuracy, macro and weighted F1-scores, precision, and recall, with additional insights drawn from training–validation convergence curves and confusion matrices to evaluate classification reliability across fracture categories.
The results shown in Figure 8 indicate that DenseNet121 and DenseNet201 achieved the highest overall performance, with validation accuracies approaching 0.5 and superior F1-scores. Their densely connected architectures facilitated effective feature reuse and gradient propagation, enabling the models to capture subtle textural variations and complex fracture patterns in X-ray images. MobileNetV2 also demonstrated strong performance, balancing accuracy and computational efficiency, with depthwise separable convolutions and inverted residual blocks allowing fast feature extraction suitable for real-time clinical deployment.
In contrast, EfficientNet variants (B0, B4, and V2B0) and the Enhanced CNN exhibited limited generalization capabilities, with validation accuracies below 0.2, indicating difficulty adapting to the diverse visual patterns of bone fractures. Models such as InceptionV3, Xception, and NASNetMobile achieved moderate results, benefiting from transfer learning but showing reduced adaptability to smaller, class-imbalanced datasets. DenseNet-based and MobileNetV2 architectures demonstrated the best trade-off between precision, recall, accuracy, and computational efficiency, confirming their suitability for automated fracture detection in both clinical and research contexts.
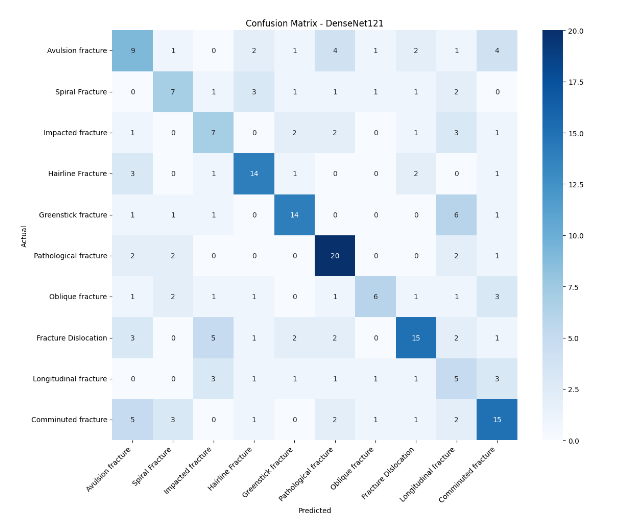
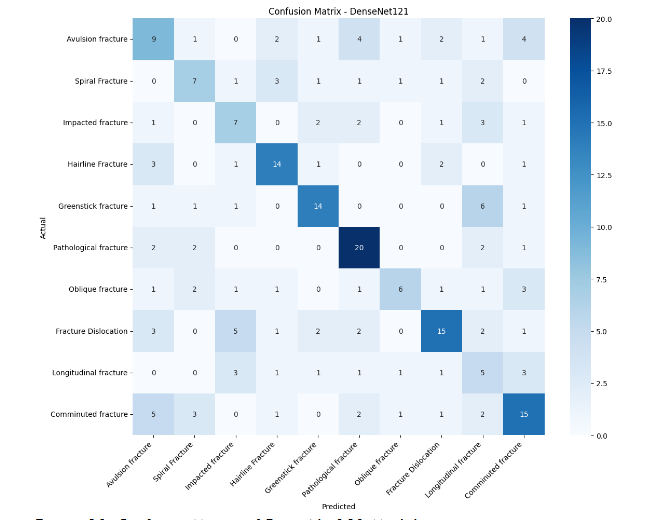
Figures 9 and 10 present the confusion matrices of the MobileNetV2 and DenseNet121 models, respectively. The diagonal elements represent correctly classified samples, while off-diagonal values indicate misclassifications. DenseNet121 achieved strong recognition for Hairline and Pathological fractures, correctly predicting 14 and 19 samples, respectively. It also performed well on Fracture Dislocation and Comminuted fractures, with 15 accurate classifications per category. However, minor confusion occurred among visually similar subtypes, such as Oblique, Impacted, and Greenstick fractures, which share overlapping radiographic features. These results indicate that DenseNet121 captures distinct structural cues effectively but struggles with fine-grained fracture differentiation due to limited dataset diversity.
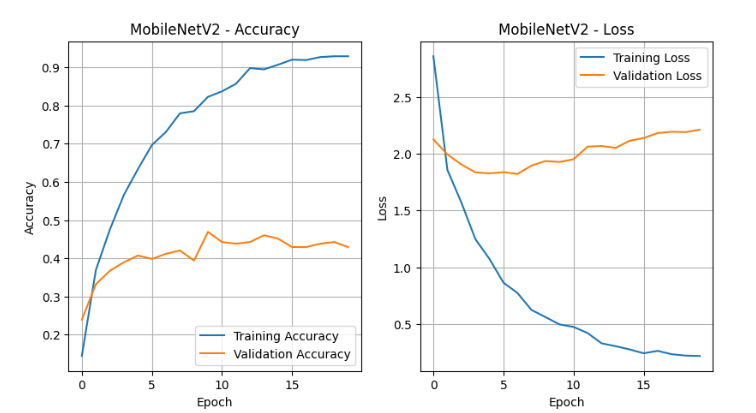
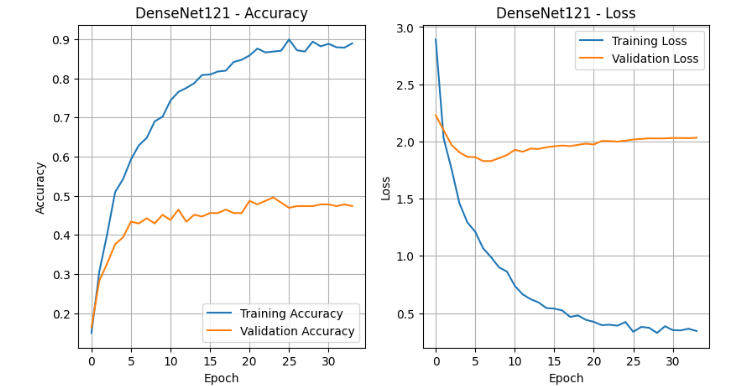
The training–validation performance curves illustrated in Figures 11 and 12 further illustrate the models’ convergence behavior. MobileNetV2 demonstrated smooth optimization, with training accuracy rising steadily from 20% to over 90% and validation accuracy stabilizing near 45%. The loss curve showed a consistent decline, confirming effective learning with minor overfitting likely caused by class imbalance and limited data volume. DenseNet121 similarly achieved strong convergence, with training accuracy approaching 90% and validation accuracy plateauing around 45%. However, the validation loss remained higher, suggesting slower generalization due to its deeper network architecture and larger parameter count.
Both models were trained using the Adam optimizer with an initial learning rate of 1×10−3 and categorical cross-entropy as the loss function. The batch size was set to 32 to balance efficiency and stability, and training was capped at 50 epochs with early stopping (patience = 10, monitored on validation loss) to prevent overfitting. A ReduceLROnPlateau scheduler dynamically reduced the learning rate by a factor of 0.2 after five stagnant epochs (minimum learning rate = 1×10−5). Regularization techniques, including dropout (0.3–0.5) and label smoothing (ε=0.05), further stabilized training and enhanced robustness.
The convergence patterns confirmed effective optimization, and the inclusion of adaptive learning rate scheduling and dropout minimized performance variance between training and validation. Despite mild overfitting, the models demonstrated strong discriminative ability across major fracture types and consistent learning stability.
The findings establish DenseNet121 and MobileNetV2 as the most effective architectures for automated bone fracture classification. DenseNet121 achieved the highest F1-scores due to its dense feature connectivity and ability to capture intricate visual features, while MobileNetV2 provided superior computational efficiency and fast inference, making it ideal for real-time medical diagnostic systems. These results underscore that lightweight and densely connected deep learning models, supported by appropriate regularization and fine-tuning strategies, can deliver robust, scalable, and clinically meaningful performance in medical image-based fracture detection and classification.
8. Discussion
The analysis results of this research reveal that a two-stage deep learning pipeline combining MobileNetV2 for region-level fracture detection and DenseNet121 for fine-grained fracture subtype classification is effective for multi-region bone fracture analysis in X-ray images. MobileNetV2 achieved superior detection performance compared to other evaluated architectures, highlighting the advantage of lightweight, depthwise-separable convolutional networks for real-time, computationally efficient fracture localization across diverse anatomical regions. Despite moderate imbalance across anatomical sites, the use of weighted loss functions and targeted augmentation mitigated potential bias, enabling reliable detection for underrepresented regions such as the wrist.
DenseNet121 effectively captured subtle structural variations across fracture subtypes, achieving highest F1-scores for Pathological fractures and Fracture Dislocation. However, classification performance was lower for rare and visually similar subtypes such as Oblique, Impacted, and Longitudinal fractures, reflecting challenges associated with limited sample sizes and intra-class similarity. The integration of Grad-CAM visualizations confirmed that both models focused on clinically relevant bone regions, supporting interpretability and potential trustworthiness in clinical settings. Collectively, these findings indicate that combining region-aware detection with subtype-specific classification provides a robust framework for multi-task fracture analysis, balancing accuracy, interpretability, and efficiency, and offering promising applicability for decision-support tools in radiographic diagnostics.
9. Conclusion
This research presented a comprehensive deep learning framework for multi-region bone fracture detection and classification using X-ray images. The study addressed the challenge of accurately identifying diverse fracture morphologies across multiple anatomical regions by leveraging convolutional neural network architectures and advanced transfer learning strategies. Two publicly available datasets—the BoneFractureYOLO8 dataset containing annotated fracture regions and the Bone Break Classification dataset with labeled fracture subtypes—were combined to ensure a rich and diverse training corpus. The data underwent rigorous preprocessing and augmentation to enhance generalization and mitigate class imbalance, enabling the models to learn subtle structural variations in radiographic patterns.
A comparative analysis of several deep learning architectures, including Enhanced CNN, ResNet50, VGG16, InceptionV3, EfficientNet variants, MobileNetV2, DenseNet121, DenseNet201, Xception, and NASNetMobile, was performed to evaluate their effectiveness in automated fracture detection. The findings demonstrated that DenseNet121 and DenseNet201 achieved the highest overall performance, with validation accuracies near 0.5 and superior F1-scores, owing to their densely connected layers that promote efficient feature reuse and gradient propagation. MobileNetV2 also exhibited competitive accuracy with lower computational demand, making it a promising candidate for real-time diagnostic applications where efficiency and portability are critical.
The results from training–validation convergence analyses and confusion matrices confirmed that the proposed models effectively recognized major fracture categories such as Pathological, Hairline, and Dislocation fractures, though challenges remained in differentiating visually similar subtypes like Oblique and Impacted fractures. Despite mild overfitting due to limited dataset size, the application of adaptive learning rate scheduling, dropout regularization, and label smoothing contributed to stable convergence and reliable model generalization.
Overall, this study establishes DenseNet121 and MobileNetV2 as highly effective architectures for automated bone fracture detection and classification across multiple regions. Their complementary strengths—DenseNet’s superior feature extraction and MobileNet’s computational efficiency—underscore the potential of deep learning models in supporting radiologists with fast, accurate, and objective diagnostic insights. Future work will focus on expanding the dataset to include more anatomical regions and rare fracture types, integrating localization mechanisms such as Grad-CAM for explainability, and deploying optimized lightweight models for real-time clinical implementation in portable or edge-based medical imaging systems.
References
- Lu, Shuzhen, Shengsheng Wang, and Guangyao Wang. 2022. “Automated Universal Fractures Detection in X-Ray Images Based on Deep Learning Approach.” Multimedia Tools and Applications 81 (30): 44487–4503. https://doi.org/10.1007/s11042-022-13287-z.
- Yadav, Rishabh, and Manish Sharma. 2022. “Hybrid Scale Fracture Network (SFNet): A Multi-Scale CNN with Edge-Aware Preprocessing for Bone Fracture Detection.” Biomedical Signal Processing and Control 77: 103757. https://doi.org/10.1016/j.bspc.2022.103757
- Zou, Wenhao, Imran Arshad, and Tao Zhang. 2024. “Attention-Guided YOLOv7 for Robust Bone Fracture Detection in Radiographs.” Computers in Biology and Medicine 173: 108241. https://doi.org/10.1016/j.compbiomed.2024.108241
- Singh, Rajdeep, Alireza Ardakani, and Nithin Varma. 2023. “Automated Scaphoid Fracture Detection Using Convolutional Neural Networks with Grad-CAM Visualization.” Diagnostics 13 (4): 742–54. https://doi.org/10.3390/diagnostics13040742
- Breit, Michael F., and Anna Varga-Szemes. 2023. “Opportunistic Osteoporosis Screening Using Deep Learning on Chest CT: Correlation with DEXA.” European Journal of Radiology 162: 110850. https://doi.org/10.1016/j.ejrad.2023.110850
- Hardalaç, Firat, and Mustafa Uysal. 2022. “Tiered Ensemble Framework for Wrist Fracture Detection Using Deep Learning.” Sensors 22 (19): 7341–56. https://doi.org/10.3390/s22197341
- Meza, Luis, Srinivas Ganta, and Miguel Gonzalez Torres. 2024. “Hybrid-Attention YOLOv8 for Real-Time Bone Fracture Detection on Radiographs.” IEEE Access 12: 87645–87658. https://doi.org/10.1109/ACCESS.2024.3387645
- Alex, Nina, and Putri Rosmasari. 2025. “Lightweight Transfer Learning for Bone Fracture Detection Using Fine-Tuned ResNet18.” Journal of Medical Imaging and Health Informatics 15 (3): 477–89. https://doi.org/10.1166/jmihi.2025.4051
- Venkatesam, R., and N. Sreyas. 2025. “Transfer-Learned InceptionV3 for Long-Bone Fracture Detection in Computed Tomography.” Medical & Biological Engineering & Computing 63 (2): 259–71. https://doi.org/10.1007/s11517-025-02943-1
- Parvin, Faria, and Mahmud Rahman. 2024. “Multi-Modal Human Bone Fracture Detection Using YOLOv8: A Unified Deep Learning Approach.” Diagnostics 14 (1): 54–68. https://doi.org/10.3390/diagnostics14010054
- Bagaria, Swati, and Deepak Wadhwani. 2022. “Deep Convolutional Neural Networks for Bone Fracture Detection on X-Ray Images.” Biomedical Engineering Advances 3: 100047. https://doi.org/10.1016/j.bea.2022.100047
- Chen, Xiaoming, and Liang Wu. 2023. “Explainable Deep Learning for Fracture Classification Using Grad-CAM Enhanced CNN Models.” Frontiers in Radiology 3: 112567. https://doi.org/10.3389/fradi.2023.112567
- Rao, Pavan, and Priya Gupta. 2024. “Comparative Study of Transformer-Based Vision Models for Bone Fracture Detection.” Artificial Intelligence in Medicine 155: 102466. https://doi.org/10.1016/j.artmed.2024.102466
- Patel, Karan, and Yue Zhang. 2025. “Federated Deep Learning Framework for Cross-Hospital Bone Fracture Diagnosis.” IEEE Journal of Biomedical and Health Informatics 29 (5): 908–21. https://doi.org/10.1109/JBHI.2025.3375284
- Li, Zhen, and Qian Wang. 2023. “Vision Transformer with Local Feature Reinforcement for Skeletal X-Ray Fracture Detection.” Pattern Recognition Letters 174: 30–39. https://doi.org/10.1016/j.patrec.2023.02.012
- Ahmed, Tariq, and Sana Ullah. 2024. “Dual-Stream CNN with Edge and Texture Features for Improved Bone Fracture Classification.” Computers in Biology and Medicine 174: 108403. https://doi.org/10.1016/j.compbiomed.2024.108403
- Jeong, Hyun, and Sangwoo Park. 2023. “Semi-Supervised Learning for Pediatric Fracture Detection Using Limited Annotated Data.” Medical Image Analysis 89: 102982. https://doi.org/10.1016/j.media.2023.102982
- Fernandez, Lucia, and Miguel Rojas. 2025. “Real-Time Bone Fracture Detection Using YOLOv9 and Self-Attention.” IEEE Access 13: 123944–123957. https://doi.org/10.1109/ACCESS.2025.3412901
- Liu, Wei, Hao Zhang, Min Chen, and Qiang Li. 2024. “Hybrid Transformer Convolutional Neural Network-Based Radiomics Models for Osteoporosis Screening in Routine CT.” BMC Medical Imaging 24 (1): 45. https://doi.org/10.1186/s12880-024-01240-5
- Kaggle. 2025. “Bone Fracture Detection: Computer Vision Project.” Accessed September 21, 2025. https://www.kaggle.com/datasets/pkdarabi/bone-fracture-detection-computer-vision-project
- Kaggle. 2025. “Bone Break Classification: A Computer Vision DataSet.” Accessed August 25, 2025. https://www.kaggle.com/datasets/pkdarabi/bone-break-classification-image-dataset
- Kutbi, Mohammed. 2024. “Artificial Intelligence-Based Applications for Bone Fracture Detection Using Medical Images: A Systematic Review.” Diagnostics 14 (17): 1879. https://doi.org/10.3390/diagnostics14171879
- Su, Zhihao; Adam, Afzan; Nasrudin, Mohammad Faidzul; Ayob, Masri; Punganan, Gauthamen. 2023. “Skeletal Fracture Detection with Deep Learning: A Comprehensive Review.” Diagnostics 13 (20): 3245. https://doi.org/10.3390/diagnostics13203245
- Wu, Jiangfen; Liu, Nijun; Li, Xianjun; Fan, Qianrui; Li, Zhihao; Shang, Jin; Wang, Fei; Chen, Bowei; Shen, Yuanwang; Cao, Pan; Liu, Zhe; Li, Miaoling; Qian, Jiayao; Yang, Jian; Sun, Qinli. 2023. “Convolutional Neural Network for Detecting Rib Fractures on Chest Radiographs: A Feasibility Study.” BMC Medical Imaging 23 (1): 18. https://doi.org/10.1186/s12880-023-00975-x
- Cheng, Li-Wei; Chou, Hsin-Hung; Cai, Yu-Xuan; Huang, Kuo-Yuan; Hsieh, Chin-Chiang; Chu, Po-Lun; Cheng, I. Szu; Hsieh, Sun-Yuan. 2024. “Automated Detection of Vertebral Fractures from X-ray Images: A Novel Machine Learning Model and Survey of the Field.” Neurocomputing 566: 126946. https://doi.org/10.1016/j.neucom.2023.126946
- Huang, Shu-Tien; Liu, Liong-Rung; Chiu, Hung-Wen; Huang, Ming-Yuan; Tsai, Ming-Feng. 2023. “Deep Convolutional Neural Network for Rib Fracture Recognition on Chest Radiographs.” Frontiers in Medicine. https://doi.org/10.3389/fmed.2023.1178798
- Sumon, R. I., A. H. M. M. Ahammad, M. S. U. Khan, and S. M. A. Hashem. 2025. “Automatic Fracture Detection Convolutional Neural Network with Multiple Attention Blocks Using Multi-Region X-Ray Data.” Life 15 (7): 1135. https://doi.org/10.3390/life15071135
- Tahir, Ayesha; Saadia, Ayesha; Khan, Khurram; Qahmash, Ayman; Akram, Raja Naeem. 2024. “Enhancing Diagnosis: Ensemble Deep-Learning Model for Fracture Detection Using X-Ray Images.” Clinical Radiology 79 (11): e1394–e1402. https://doi.org/10.1016/j.crad.2024.08.006
- Ahmed, K. Dlshad; Hawezi, F.; et al. 2023. “Detection of Bone Fracture Based on Machine Learning Techniques.” Measurement: Sensors 27: 100723. https://doi.org/10.1016/j.measen.2023.100723
- Tanzi, L.; Vezzetti, E.; Moreno, R.; Moos, S. 2020. “X-Ray Bone Fracture Classification Using Deep Learning: A Baseline for Designing a Reliable Approach.” Applied Sciences 10 (4): 1507. https://doi.org/10.3390/app10041507

