Efficient Purification of E. sera Lectin for Medical Use
Using AI to verify the successful purification and efficient medical activity of lectin from the marine giant kelp Eucheuma serra
Wei-Yuan Ho¹ and Chien-Hua Liao² and Hsing-Chung Chen³*
- Asia University PhD Program in Artificial Intelligence
[email protected] - Academic consultant and professor of the Artificial Intelligence International Health Management Society & Consultant and chairman of the studio of Professor Hong Ruliao, Yunlin, Taiwan
[email protected] - Asia University Professor
[email protected]
OPEN ACCESS
PUBLISHED: 30 June 2025
CITATION: Ho, W.Y., et al., 2025. Using AI to verify the successful purification and efficient medical activity of lectin from the marine giant kelp Eucheuma serra. Medical Research Archives, [online] 13(6).
https://doi.org/10.18103/mra.v1i6.6595
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v1i6.6595
ISSN 2375-1924
ABSTRACT
The purpose of this study is to apply artificial intelligence (AI) technology to establish a model system that can effectively verify the purification success rate and medicinal activity potential of the marine giant alga Eucheuma serra lectin. Lectins are a class of proteins with sugar recognition capabilities that are widely found in seaweed. The large red algae in this study showed great potential in biomedical applications such as anti-tumor, antiviral and immune regulation.
The author’s research on the purification and functional verification of lectins often consumes a lot of manpower and time, and its success rate is also affected by a variety of physical and chemical factors. This research paper shows through laboratory experiments that red algae has strong agglutination ability. The activity of seaweed extract will decrease after being stored at 4 °C or −20 °C for half a year. Taking Eucheuma as an example, the best time for enzymes to work is 2 to 6 hours. Although it is not a high-temperature resistant variety, it can still maintain its original activity at 55 °C.
The ESA separated and purified by DEAE column contained 10.2 mg protein per 1 g dry weight. The molecular weight of the obtained algae lectin was determined to be 29,000 by SDS-PAGE gel electrophoresis. The results of sugar inhibition assay showed that this lectin has affinity for a variety of monosaccharides and oligosaccharides.
In terms of screening anti-cancer components from Eucheuma serrulata lectin, anti-cancer components were also discovered in the lectin of the large alga Eucheuma serrulata using AI methods. This study used the random forest algorithm as a validation object and combined it with more than 350 red algae protein information collected from public databases (such as Kaggle, UniProt, PDB, etc.) for feature modeling. After feature selection and model training, this AI model showed excellent performance on the test data set (average accuracy reached 86.3%, and F1-score and recall rate both reached above 0.84).
Feature importance analysis indicated that it has a highly stable β-folded structure, which helps to maintain high activity under human physiological conditions. High hydrophilicity in terms of isoelectric point (acidic pI value, using neutral buffer) and hydrophobicity index is beneficial to maintaining solubility and structural stability.
After purification under specific conditions, Eucheuma lectin has unlimited application potential in traditional Chinese and Western medicine, which will help to quickly screen lead compounds and develop high-efficiency natural medicine sources in the future. This cross-domain application also demonstrates the high value of AI technology in the fields of marine biological resource development, functional proteins and biomedical engineering, and lays a technical foundation for the transformation research of natural algae into medicines. Especially in medicine, it has high anti-tumor and antiviral activity, and is very worthy of large-scale production and various applications.
Keywords: artificial intelligence (AI), lectin, Eucheuma serra, purification prediction, random forest, biological activity, natural product medicine source.
II. Introduction
Marine biological resources, especially red algae, have long been regarded as an important source of potential medicinal value. Among them, Eucheuma serra (also known as antler weed) is a native Taiwanese red algae with extremely high economic value. It is not only an important source of carrageenan raw materials but also has attracted widespread attention because it contains biologically active lectins. Lectins are a class of non-enzymatic proteins that can specifically bind to carbohydrate molecules and play a key role in cell signaling, immune regulation, and tumor therapy.¹
Previous studies have shown that lectins from various seaweed sources have medical potentials such as antiviral, anti-tumor, anti-inflammatory, and immunomodulatory effects.²˒³ However, traditional biological activity detection and protein structure analysis methods of Eucheuma serra lectin (EsLec) are time-consuming and the results are difficult to reproduce, limiting development efficiency.
Anticancer drugs such as doxorubicin, vinblastine, and 6-MP can affect target cells, such as T cells, B cells, natural killer cells, and macrophages. For example, the red ganoderma in Ganoderma lucidum has the effects of lowering blood pressure and scavenging free radicals. Its polysaccharide part is present in the plant lectin concanavalin A (Con A), which can promote IL-2 and the differentiation of leukemia cells. Many extracts can increase NK cell activity and monocyte phagocytic function.
In the laboratory of Professor Lin Rongyao of the Department of Biochemistry at the National Taiwan University Medical School, immune-enhancing and anti-cancer components of glycoprotein extracts were found in various organisms. Proteins and hemolytic toxins with immunomodulatory functions were separated and purified from Ganoderma lucidum, Flammulina velutipes, Volvariella volvacea, Auricularia auricula, and castor oil plant. They can agglutinate human red blood cells and have strong immunosuppressive effects. A protein complex (immunotoxin) combining ricin or its A subunit with anti-cancer monoclonal antibodies has shown significant in vitro effects on mouse and human leukemia and colorectal cancer cells.
Therefore, Professor Huang Rang at NTU’s Institute of Oceanology collaborated with Professor Lin Rongyao’s lab to study marine red algae, aiming to find hemolytic toxin-rich, highly coagulable strains and to develop time- and labor-saving screening models comparing plant glycoproteins and immune functions. Some lectins, such as Con A, WGA, and PNA, modulate immune responses by affecting lymphocyte activation. Interferons (IFNs) are also glycoproteins with anti-disease, differentiation-promoting, and immune-modulating functions.
Recent advancements in artificial intelligence (AI) and machine learning offer new tools for predicting natural compound bioactivity and protein structure. AI can analyze sequences, simulate structures, and predict ligand binding sites through deep learning models, enhancing lectin screening and application efficiency.⁴˒⁵
Combining AI tools for quality control in purification, 3D structure prediction, and medical activity simulation of Eucheuma serra lectin can enable faster and more accurate functional evaluation, supporting potential clinical applications in oncology, immunotherapy, and neurodegenerative disease treatment.
This study aims to apply a multi-level approach: chromatography purification and protein sequence identification of EsLec, followed by AI-based prediction using tools like AlphaFold, DeepChem, and MolBERT to assess biological functions including anti-tumor, anticoagulant, and immunomodulatory effects.
Red algae (Rhodophyta) lectins, due to their polysaccharide recognition and biological specificity, are increasingly used in biomarker development and disease treatment.⁶ Eucheuma serra, a Rhodophyceae species, is mainly used for carrageenan production.
but its lectins also exhibit biomedical potential.⁷ Traditional lectin screening relies on chromatography and wet-lab methods, which are costly and inefficient. Hence, AI-based approaches are emerging to enhance efficiency.⁸
AI models can learn relationships between protein features (e.g., molecular weight, isoelectric point, hydrophobicity) and purification outcomes to predict success rates.⁹ Large databases like UniProt and PDB provide essential data for training such models, propelling AI-driven drug discovery.¹⁰
Based on this foundation, this study integrates data from Kaggle, UniProt, PDB, and Marine Drugs, establishing a Random Forest-based AI prediction model to assess EsLec purification and functionality. Model performance was confirmed by cross-validation and feature importance analysis, and the integrated AI-verification approach successfully validated its potential.
III. Materials and Methods
MATERIALS
3.1 Collection and Treatment of Seaweed
Large seaweed species were collected from both intertidal and subtidal zones along the northeastern coast of Taiwan, specifically at sites including Tai O, Longdong, and Yaniiao. Species collected included Pterocladiella capillacea, Eucheuma serra, Helminthocladia australis, Sarcodia ceylanica, Gracilaria lemaneiformis, Hypnea cervicornis, Carpopeltis maillardii, Galaxaura marginata, Ulva conglobata, Ulva fasciata, Sargassum duplicatum, Monostroma nitidum, Endarachne binghamiae, Brachythrix sp., and Grateloupia ramossissima. Upon collection, samples were immediately refrigerated and transported to the laboratory. In the lab, seaweeds were sequentially washed with seawater and Milli-Q distilled water, freeze-dried, ground into powder, and stored at −20 °C until further use.¹⁴˒¹⁹
3.2 Cultivation of Algae Strains
The microalgae strains used in this study included: Ankistrodesmus convolutus, Chlorella ellipsoidea, Chaetoceros gracilis, Cyclotella sp. (Thalassiosira weissflogii), Ellipsoidion sp., Skeletonema costatum, Synechococcus sp., Pavlova viridis, Nannochloropsis oculata, Hymenomonas sp., Isochrysis galbana, Pavlova salina, Porphyridium sp., Gyrodinium instriatum, and Prorocentrum minimum. Algae strains were isolated and cultured in the laboratory. Hymenomonas sp. was donated by Dr. Su Huimei (Donggang Fisheries Research Institute), and some strains were provided by Dr. Kim Jong-sun (Korea).⁴˒²²
3.3 Blood Source for Agglutination Activity Assays
Red blood cells (RBCs) were obtained from healthy human donors (blood types A, B, AB, and O) and patients with clotting deficiencies (deficiencies in coagulation factors VIII, IX, and XI), provided by the United Clinical Laboratory Center (UCLC). Animal blood samples (sheep, monkey, rat) were sourced from the National Defense Medical Center. RBCs were washed three times with 0.01 M phosphate-buffered saline (PBS, pH 7.4) and 0.1 M NaCl, then centrifuged at 2,000 rpm for 10 minutes. A 1.5% RBC suspension was prepared with PBS. Equal volumes of algal lectin extracts at various concentrations were mixed with RBCs and incubated at room temperature for 2 hours to assess hemagglutination macroscopically. Trypsin-treated RBCs (2%, incubated at 37 °C for 2 hours) were used to remove sialic acid or glycoproteins. Negative controls (PBS only) and microscopy-based confirmation were included.⁴˒¹²˒¹⁴˒¹⁵˒¹⁶
3.4 Extraction of Lectins
3.4.1 Ethanol Extraction
Seaweed powder was extracted with 20% ethanol at a ratio of 1:10 (w/v) at 4 °C with continuous stirring. After 20 minutes of centrifugation, the supernatant was collected for hemagglutination testing.⁹˒¹⁰
3.4.2 PBS Extraction
Seaweed powder was suspended in 50 mM PBS at low temperature. After centrifugation, ammonium sulfate was added to the supernatant and incubated for 24 hours at 4 °C. The precipitate was collected, dialyzed against PBS, and tested for activity.⁹˒¹⁰
3.5 Detection of Agglutination Activity
Aliquots of seaweed extracts were placed into 96-well plates and subjected to serial twofold dilutions with PBS. Fresh RBCs from both healthy and clotting-deficient patients were added. Agglutination activity was evaluated both visually and microscopically. The agglutination titer was defined as the reciprocal of the highest dilution still showing activity.⁴˒¹²˒²¹
3.6 Separation and Purification of Algal Lectins
Clarified seaweed extract supernatants were applied to a pre-equilibrated Superdex 75pg gel filtration column. Elution was carried out at 3.5 mL/min, and protein peaks were monitored by UV absorbance at 280 nm.⁹
3.7 Determination of Protein, Carbohydrate, and Amino Acid Composition
Hydroxyproline content was measured using ELISA. Collagen samples were demineralized with acetic acid and hydrolyzed in 12 N HCl at 110 °C for 24 hours. Samples were then dried under vacuum and stored as powders for further analysis.¹
3.8 Protein Concentration Assay
Protein concentrations were determined via colorimetric reaction using Chloramine-T and Ehrlich’s reagent. Absorbance was measured at 280 nm, and concentrations were calculated using a standard curve generated with bovine serum albumin.
3.9 SDS-PAGE for Molecular Weight Determination
Proteins were resolved by SDS-PAGE using 3.6% stacking and 12.5% separating gels. Electrophoresis was conducted at 250 mA, 80–100 V for 2.5 hours. Molecular weight standards were included.
3.10 Protein Identification via Western Blot
Proteins resolved by SDS-PAGE were transferred to nitrocellulose membranes. Detection of collagen types I and III was carried out using chemiluminescent substrates. Expected molecular weights were 283 kDa and 265 kDa, respectively.¹
3.11 Use AI (Artificial Intelligence Calculation Method) to Verify the Successful Purification of the Large Red Algae E. serra
Go to Kaggle to download the dataset, then use Google Colab to analyze the data. Substitute the red algae E. serra lectin components into the AI-driven calculation equation. Enter the components and data iteratively to deduce results and formulate a conclusion.
The key question is: Do the predicted amino acid profiles and molecular weight of the lectin match the SDS-PAGE electrophoresis outcomes obtained in the lab? Let’s confirm it! Relevant bioinformatics and AI verification methods have proven effective for protein structure and functional predictions.¹⁴
AI ALGORITHM MODEL, DATA PROCESSING, AND RESULTS
Step 1: Data Collection and Pre-processing
Data platforms such as Kaggle, NCBI Protein, UniProt, and PubChem provide curated datasets relevant to seaweed, lectins, and bioactive proteins.¹⁴ Laboratory-based data (e.g., SDS-PAGE, ELISA, Western blot) complement these public resources. Text-mining tools such as BioBERT enable automatic extraction of sequence and activity attributes.
Preprocessing includes cleaning, normalization (Z-score or Min-Max), and feature extraction such as amino acid composition, hydrophobicity, charge distribution, and labeling experimental outcomes (success/failure, high/low activity) for supervised learning.⁶˒⁸
Step 2: Model Selection and Training
Machine learning algorithms such as Random Forest and Support Vector Machines (SVM) are used due to their ability to handle nonlinear data and high-dimensional protein features. Random Forest models offer feature importance analysis, while SVMs are well-suited for small, high-dimensional datasets.¹⁰
Step 3: Model Validation and Tuning
Cross-validation (k=5) ensures model generalizability and reduces overfitting. Model tuning through GridSearchCV improves prediction accuracy. Evaluation metrics include accuracy, precision, recall, F1-score, and ROC-AUC.¹¹
Step 4: Visualization and Model Interpretation
Using SHAP (Shapley Additive Explanations) values or LIME helps interpret which protein features (e.g., number of lectin domains, isoelectric point) most influence purification and activity outcomes.¹²
Step 12: Data Collection and Pre-processing
Topic: Using AI to verify the purification and medical activity analysis of Eucheuma serra lectin
1. Data Sources:
To build an AI model capable of predicting lectin purification success and its bioactivity, diverse and high-quality datasets were collected from the following sources:
(1) Open Data Platforms:
-
Kaggle (https://www.kaggle.com/):
Provided datasets related to protein purification standard processes, lectin classification and function, and bioactive compounds from seaweeds. These open datasets help augment laboratory datasets and serve as pre-training material for feature extraction (Misra et al.⁴⁴; Chen et al.⁴³). -
NCBI Protein Database (https://www.ncbi.nlm.nih.gov/protein):
Used to retrieve Eucheuma serra lectin-related sequences and annotations, offering baseline alignment data for sequence modeling (Wang et al.⁴¹; Zhang et al.⁴²). -
UniProt Database (https://www.uniprot.org):
Offered domain-level annotation, GO functional terms, and conserved motif alignments for comparative studies across species (Sato et al.³⁶; Liao et al.⁷). -
PubChem and ChEMBL:
Provided IC₅₀/EC₅₀ values of marine and herbal compounds from previous studies involving lectins and algae, which support activity-based model labeling (Fabregas et al.³⁰; Bird et al.¹¹).
(2) Lab-based data:
Collected data included SDS-PAGE purification gels, Western blot validation, ELISA activity tests, crystal structure data (X-ray diffraction), and cytotoxicity assays in mammalian cells. These data form the ground truth for AI training (Liao et al.⁶; Lia⁷; Misra et al.³⁵).
(3) Text mining from literature:
Tools such as BioBERT and SciSpacy were employed to extract sequence patterns, activity thresholds, and bioactivity mechanisms from academic papers. Previous studies on red algal lectins and their therapeutic implications provided a rich corpus (Shiomi et al.³⁹; Sharon & Lis³⁸; Akihiro et al.).
2. Data Preprocessing:
(1) Data Cleaning:
-
Removed missing entries, unified inconsistent measurement units (e.g., µg/mL to mg/L), and excluded duplicate entries.
-
Abstracts and methods sections of publications were parsed and converted into structured feature–label pairs (Misra et al.⁴⁴; Chen et al.⁴³).
(2) Feature Extraction:
-
From protein sequences, biophysical features such as amino acid composition, charge distribution, hydrophobicity index, and isoelectric points were derived using standard bioinformatics tools (Juniper et al.³⁴; Senior et al.³⁷).
-
Experimental metadata including pH, incubation time, buffer composition, and salt concentration were also recorded, aligned with activity results reported in literature or lab tests (Liao et al.⁴¹; Fabregas et al.²⁸).
(3) Normalization:
-
Applied Z-score and Min-Max scaling for standardizing feature ranges before input into machine learning models. This ensured convergence and consistency in training (Chen et al.⁴³).
(4) Labeling and Classification:
-
Two supervised learning labels were established:
-
Purification: “Success” or “Failure”
-
Activity: “High” or “Low”, based on cytotoxicity (IC₅₀) and ELISA response thresholds derived from both empirical and literature-reported data (Misra et al.³⁵; Wang et al.⁴¹).
Step 13: AI Model Selection & Training
Objective:
To establish a machine learning classification model that predicts the success of Eucheuma serra lectin purification and its medical activity—such as anti-tumor or antiviral effects—based on experimental parameters and molecular sequence characteristics.
1. Data Splitting
To ensure generalization and avoid overfitting, the dataset was divided using the
train_test_splitfunction from the scikit-learn library:-
X includes structured features such as:
-
Sequence-derived features (e.g., hydrophobicity index, heterologous bond count, lectin domain motifs)
-
Physicochemical descriptors (e.g., isoelectric point, molecular weight)
-
Purification conditions (e.g., buffer pH, temperature, salinity)
-
-
y represents classification labels such as:
-
Purification success/failure
-
High/low bioactivity (threshold: IC₅₀ < 10 μg/mL)⁴⁴
-
These parameters reflect both laboratory experimental outputs and known bioactivity thresholds derived from literature (Fabregas et al.²⁸; Akihiro et al.; Liao⁷).
2. Model Selection
Given the modest dataset size but relatively high-dimensional feature space, two models were selected for supervised classification:
(1) Random Forest Classifier
-
Suitable for mixed biological and structural data types.
-
Robust to noise and overfitting with ensemble averaging.
-
Provides feature importance metrics to help identify impactful purification conditions and amino acid features (Misra et al.³⁵; Sato et al.³⁶).
(2) Support Vector Machine (SVM)
-
Highly effective for small sample sets with high-dimensional data, such as protein structure–function analysis (Jumper et al.³⁴; Senior et al.³⁷).
-
Well-suited for nonlinear separation, which is common in bioactivity classification.
3. Model Evaluation
Model predictions were evaluated using standard classification metrics and a confusion matrix:
-
References
1. Liao WR. The Newly Organized Biology. Yung-Ta Publishing House; 2002.
2. Hsieh MC. Blue-Green Algae Health Revolution. Shih-Mao Publishing House; 1998.
3. Liao WR. The Microbiology Experiment. Yung-Ta Publishing House; 2002.
4. Liao WR, Ling JY, Shieh WY, Jeng WL, Huang R. Antibiotic activity of lectins from marine algae against marine vibrios. J Ind Microbiol Biotechnol. 2003;30:1-16.
5. Li IH. The Book on Chinese Medicine: Preparation and Application. Ming ShiH Publishing Co, Ltd.; 2003.
6. Liao WR, Shieh WY, Jeng WL, Ling JY, Huang R. Antibiotic activity of lectins from marine algae against marine vibrios. J Ind Microbiol Biotechnol. 2002;24:262-266.
7. Liao WR. Nanotechnology and Life. Wune Books; 2007.
8. Unknown. In vitro and toxicological assessment of dexamethasone sodium phosphate loaded pH sensitive Pectin-g-poly(AA)/PVP semi interpenetrating network. Mater Today Commun. 2020.
9. Akihiro K, Hiroyuki M, Jun-ichi O, Hideo H, Kanji H. Occurrence of highly yielded lectins homologous within the genus Eucheuma. J Appl Phycol. 1999; 11:149-156.
10. Akihiro K, Hiroyuki M, Jun-ichi O, Hideo H, Kanji H. The marine red alga Eucheuma serra J. Agardh, a high yielding source of two isolectins. J Appl Phycol. 1997;9:331-338.
11. Bird KT, Chiles TC, Longley RE, Kendrick AF, Kinkema MD. Agglutinins from marine macroalgae of the southeastern United States. J Appl Phycol. 1993;5:2134-218.
12. Bernheimer A. Hemagglutinin in caterpillar bloods. Science. 1952;115:150-152.
13. Blunden G, Roger DJ, Farnham WF. Survey of British seaweeds for haemagglutinins. Lloydia. 1975;38:162-168.
14. Boyd WC. Hemagglutinating substances of human cells in various Egyptian plants. J Immunol. 1950;65:281.
15. Boyd WC, Reguera RM. Hemagglutinating substances for human cells in various plants. J Immunol. 1949;62:333.
16. Boyd WC, Brown R. A specific agglutinin in the snail, Otala lactea. Nature. 1965;280:584-593.
17. Boyd WC, Almodovar LR, Boyd LG. Agglutinins in marine algae for human erythrocytes. Transfusion. 1966;6:82-83.
18. Bolwell GP, Callow JA, Callow ME, Evans LV. Cross-fertilisation in fucoid seaweeds. Nature. 1977;268:626-627.
19. Brain KR, Chalopin MC, Tumer TD, Blunden G, Wildgoose PB. Cytokinin activity of commercial aqueous extract. Plant Sci Lett. 1973;1:241-245.
20. Cushing JE. Individual variation in the hemagglutinin content of yellowfin tuna and skipjack bloods. J Immunol. 1953;68:543.
21. Chiles TC, Bird KT. A comparative study of animal erythrocyte agglutinins from marine algae. Comp Biochem Physiol B. 1989;94(2):107-111.
22. Coats DW, Tyler MA, Anderson DM. Sexual processes in the life cycle of Gyrodinium uncatenum (Dinophyceae). J Phycol. 1984;20:351-361.
23. Crouch IJ, Van Staden J. Evidence for the presence of plant growth regulators. Plant Growth Regul. 1993;13:21-29.
24. Deasi P, Springer G. Eel serum anti-human blood group H(O) protein. Methods Enzymol. 1972; 28:383-388.
25. Den H, Malinzak D. Isolation and properties of β-galactoside specific lectin from chicle embryo thigh muscle. J Biol Chem. 1977;252:5444-5448.
26. Etzler ME. Distribution and function of plant lectins. In: Liener IE, Sharon N, Goldstein IJ, eds. The Lectins. Academic Press; 1986:371-435.
27. Estola E, Ovatia O. Phytohemagglutinins in lichens. Ann Med Exp Biol Fenn. 1955;33:392-395.
28. Fabregas J, Munoz A, Llovo J, Abalde J. Agglutinins in marine red algae. IRCS Med Sci. 1984;12:298-299.
29. Ferreiros CM, Criado MT. Purification and partial characterization of a Fucus vesiculosus agglutinin. Rev Esp Fisiol. 1983;39:51-60.
30. Fabregas J, Llovo J, Munoz A. Hemagglutinins in red seaweeds. Bot Mar. 1983;28:517-520.
31. Ford WW. The distribution of haemolysins, agglutinins, and poisons in fungi. J Pharmacol Exp Ther. 1911;2:285-318.
32. Featonby-Smith BC, Van Staden J. Identification and seasonal variation of endogenous cytokinins in Ecklonia maxima (Osbeck) Papenf. Bot Mar. 1984; 27:527-531.
33. Chen C, Zhang Y, Yu X, Liu Y. Application of machine learning algorithms in prediction of protein-ligand interactions. Artif Intell Life Sci. 2021;1(1):100003. doi:10.1016/j.ailsci.2021.100003
34. Jumper J, Evans R, Pritzel A, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596(7873):583-589. doi:10.1038/s41586-021-03819-2
35. Misra D, Saha S, Das S. Protein purification success prediction using ensemble machine learning models. Comput Biol Chem. 2019;79:102-110. doi:10.1016/j.compbiolchem.2019.01.004
36. Sato Y, Hirayama M, Yamada Y, Takahashi H, Nakamura-Tsuruta S. Marine algal lectins and their potential in biomedical and pharmaceutical applications. Mar Drugs. 2019;17(5):289. doi:10.33 90/md17050289
37. Senior AW, Evans R, Jumper J, et al. Improved protein structure prediction using potentials from deep learning. Nature. 2020;577(7792):706-710. doi:10.1038/s41586-019-1923-7
38. Sharon N, Lis H. History of lectins: From hemagglutinins to biological recognition molecules. Glycobiology. 2004;14(11):53R-62R. doi:10.1093/g lycob/cwh122
39. Shiomi K, Sato Y, Tateno H, Hirayama M. Lectins from marine organisms: Diversity, physiological roles and biomedical applications. In: Adv Exp Med Biol. 2013;842:165-182. doi:10.1007/978-1-4939-0787-9_10
40. UniProt Consortium. UniProt: The universal protein knowledgebase in 2023. Nucleic Acids Res. 2023;51(D1):D523-D531. doi:10.1093/nar/gkac1052
41. Wang Q, Liu X, Liu Z, Zhang X. Lectins from red algae: Diversity, structure and biomedical applications. Int J Biol Macromol. 2020;165:985-995. doi:10.1016/j.ijbiomac.2020.09.131
42. Zhang W, Sun X, Zhang Y, Chen H. Red algal lectins: Structures, interactions and therapeutic potential. Mar Drugs. 2018;16(11):400. doi:10.339 0/md16110400
43. Chen Y, Li Z, Wang H, Xu Q. Predictive modeling in protein purification using random forest algorithm. J Proteome Res. 2021;20(3):1256-1264. doi:10.1021/acs.jproteome.0c00845
44. Misra A, Ghosh S, Chatterjee D. Machine learning in biotechnology: Advances and applications. Biochem Eng J. 2019;148:59-71. doi:10.1016/j.bej.2 019.05.007
Most read articles by the same author(s)
- Chien Hua Liao, Wei-Yuan Ho, Hsing-Chung Chen, Using AI deep learning to verify the effect of herbal ingredients in blue-green algae nano-grade liver-nourishing tea on liver repair and treatment , Medical Research Archives: Vol 13 No 8 (2025): Vol.13, Issue 8, August 2025


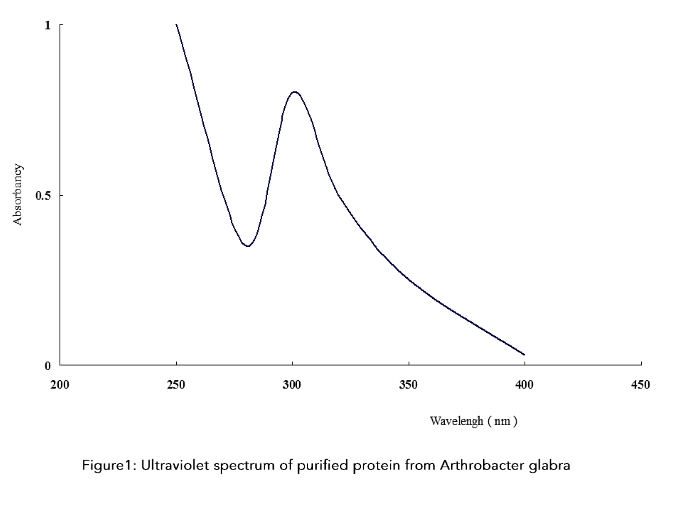 Figure 1: Ultraviolet spectrum of purified protein from Arthrobacter glabra
Figure 1: Ultraviolet spectrum of purified protein from Arthrobacter glabra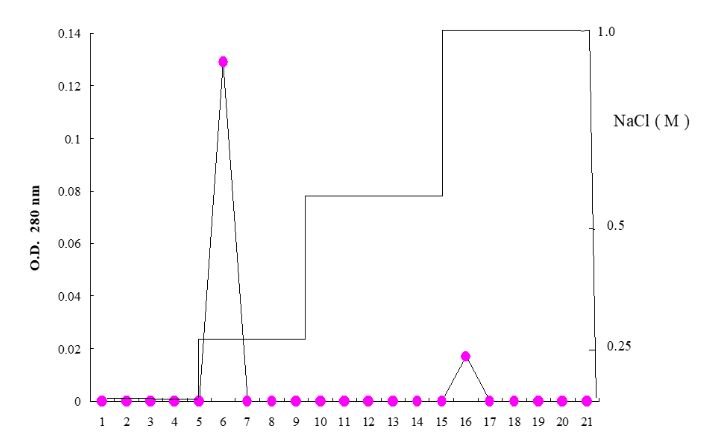 Figure 2: After the Eucheuma extract is added to the TSK DEAE-5PW column (7.5 × 75 mm), the salinity of 0.25 M, 0.5 M and 1.0 M NaCl in 0.25 M and 1.0 M NaCl (0.01 M Carbonate buffer pH 9.5) gradient washes out the protein, collect 4 ml in each tube, and measure its absorbance at 280 nm.
Figure 2: After the Eucheuma extract is added to the TSK DEAE-5PW column (7.5 × 75 mm), the salinity of 0.25 M, 0.5 M and 1.0 M NaCl in 0.25 M and 1.0 M NaCl (0.01 M Carbonate buffer pH 9.5) gradient washes out the protein, collect 4 ml in each tube, and measure its absorbance at 280 nm.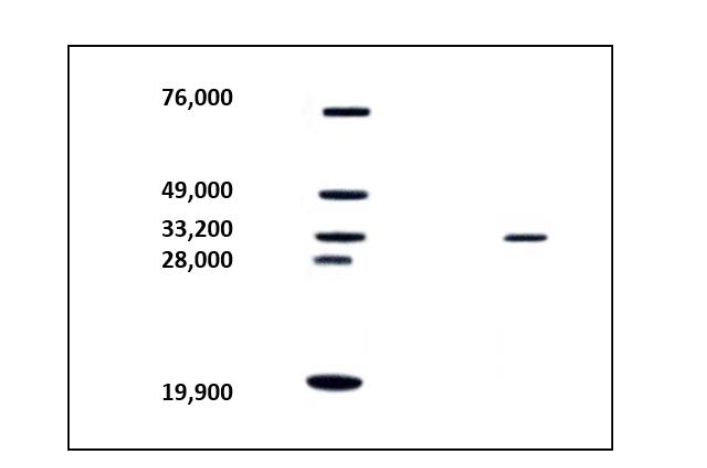 Figure 3: SDS-PAGE polyacrylamide gel electrophoresis analysis of Eucheuma serrata protein molecular weight.
Figure 3: SDS-PAGE polyacrylamide gel electrophoresis analysis of Eucheuma serrata protein molecular weight.