Metadichol®: Inhibiting Zoonotic Viruses In Vitro
Metadichol® Inhibits Zoonotic Viruses Nipah, Lassa, and Rabies In Vitro: Evidence for the Vitamin D Receptor–MYC–SP1–GSPT1 Axis and Nuclear Receptor Modulation as Core Antiviral Mechanisms
P.R. Raghavan¹
- Nanorx, Inc., PO Box 131, Chappaqua, NY 10514, USA
Email: [email protected]
OPEN ACCESS
PUBLISHED: 31 March 2026
CITATION: Raghavan, P.R., 2026. Metadichol® Inhibits Zoonotic Viruses Nipah, Lassa, and Rabies In Vitro: Evidence for the Vitamin D Receptor–MYC–SP1–GSPT1 Axis and Nuclear Receptor Modulation as Core Antiviral Mechanisms. Medical Research Archives, [online] 14(3).
COPYRIGHT: © 2026 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
ISSN 2375-1924
Abstract
Emerging and re-emerging zoonotic viruses, including Nipah virus, Lassa virus, and rabies virus, represent a persistent and escalating threat to global public health, for which no broadly effective antiviral agents currently exist. Metadichol®, a nanoemulsion of long-chain alcohols (policosanol), has previously demonstrated significant in vitro efficacy against diverse viruses including severe acute respiratory syndrome coronavirus 2 and Ebola virus. In this study, we present new in vitro data showing that Metadichol potently inhibits Lassa, Nipah, and rabies virus entry using a novel alphavirus-based pseudovirus platform, with half-maximal inhibitory concentration values of 831.7, 2455, and 2621 ng/mL, respectively, and no significant cytotoxicity in human embryonic kidney 293T cells. We focus our mechanistic analysis on two core pathways that underpin this broad-spectrum activity. First, we describe the vitamin D receptor–MYC–specificity protein 1–G1 to S phase transition 1 axis, in which Metadichol activates the vitamin D receptor, leading to MYC-mediated repression of specificity protein 1 and consequent downregulation of G1 to S phase transition 1 (also known as eukaryotic release factor 3a), a translation termination factor recently validated as a druggable host dependency factor essential for the replication of Lassa, Ebola, and other viruses. Second, we examine Metadichol’s modulation of nuclear receptors—including neuron-derived orphan receptor 1, liver X receptor alpha, and peroxisome proliferator-activated receptor gamma—which suppress interferon regulatory factors and the interferon-beta promoter, thereby fine-tuning the innate immune response to prevent immunopathology while maintaining antiviral defense. Together, these two mechanisms—direct replication blockade via G1 to S phase transition 1 inhibition and immune optimization via nuclear receptor modulation—provide a mechanistic framework for Metadichol’s observed broad-spectrum efficacy. Given that Metadichol is commercially available and non-toxic (median lethal dose > 5000 mg/kg in rats), these findings support its further clinical investigation as a host-directed antiviral agent.
Keywords
Nipah virus, rabies virus, Lassa virus, Metadichol, GSPT1, vitamin D receptor, host-directed antiviral, nuclear receptors, peroxisome proliferator-activated receptor gamma, liver X receptor, broad-spectrum antiviral
Abbreviations
| Abbreviation | Definition |
|---|---|
| 4PL | Four-parameter logistic model |
| ABCA1 | ATP-binding cassette transporter A1 |
| BMAL1 | Brain and muscle ARNT-like 1 |
| BSL-4 | Biosafety level 4 |
| eRF1/eRF3a | Eukaryotic release factor 1 / 3a |
| FOX | Forkhead box transcription factor family |
| FXR | Farnesoid X receptor |
| GDF11 | Growth differentiation factor 11 |
| GSPT1 | G1 to S phase transition 1 (also known as eRF3a) |
| HCV | Hepatitis C virus |
| HDT | Host-directed therapy |
| HEK293T | Human embryonic kidney 293T cells |
| HIV | Human immunodeficiency virus |
| HNF4α | Hepatocyte nuclear factor 4 alpha |
| IC₅₀ | Half-maximal inhibitory concentration |
| IFN | Interferon |
| IFNβ | Interferon beta |
| IRF | Interferon regulatory factor |
| JEV | Japanese encephalitis virus |
| KLF | Krüppel-like factor |
| LasV | Lassa virus |
| LD₅₀ | Median lethal dose |
| LXR/LXRα | Liver X receptor / liver X receptor alpha |
| mTOR | Mechanistic target of rapamycin |
| MXD1 | MAX dimerization protein 1 |
| NF-κB | Nuclear factor kappa-light-chain-enhancer of activated B cells |
| NiV | Nipah virus |
| NOR1 (NR4A3) | Neuron-derived orphan receptor 1 |
| NR | Nuclear receptor |
| PAMP | Pathogen-associated molecular pattern |
| PPARα/PPARγ | Peroxisome proliferator-activated receptor alpha / gamma |
| RabV | Rabies virus |
| RdRp | RNA-dependent RNA polymerase |
| RFU | Relative fluorescence units |
| RLU | Relative light units |
| RXRα | Retinoid X receptor alpha |
| SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 |
| SIRT | Sirtuin |
| SP1 | Specificity protein 1 |
| TLR | Toll-like receptor |
| VDR | Vitamin D receptor |
Introduction
Zoonotic viruses—those transmitted from animal reservoirs to humans—account for the majority of newly emerging infectious diseases and represent one of the most significant threats to global health security. The recent decades have witnessed repeated outbreaks caused by highly pathogenic zoonotic viruses, including Nipah virus, Lassa virus, Ebola virus, and coronaviruses, each with the potential to cause severe morbidity, high case fatality rates, and regional or global disruption. The epidemiology of zoonotic viral diseases is shaped by complex interactions among animal reservoirs, environmental change, human encroachment on wildlife habitats, and global travel, which together create conditions favorable for cross-species transmission and pandemic emergence. A One Health approach integrating human, animal, and environmental surveillance is essential, but is insufficient without effective therapeutic countermeasures.
Among the most concerning zoonotic pathogens are Nipah virus (NiV), Lassa virus, and rabies virus (RabV), all of which are associated with high morbidity and limited therapeutic options. Nipah virus, a paramyxovirus first recognized during a 1999 outbreak among pig farmers in Malaysia and Singapore, causes encephalitis with case fatality rates estimated at 40–75%. Transmission occurs via contact with infected animals (particularly bats and pigs), consumption of contaminated food, or direct human-to-human spread. Since its initial identification, NiV has caused nearly annual outbreaks in Bangladesh and India, and no approved vaccine or specific antiviral treatment exists, although monoclonal antibodies and remdesivir are under investigation. Lassa virus, an arenavirus endemic to West Africa, causes hemorrhagic fever in an estimated 100,000–300,000 individuals annually, with a case fatality rate of approximately 1% overall but up to 15% in hospitalized patients. Ribavirin has shown limited efficacy when administered early, but there are no approved vaccines, and new therapeutics are urgently needed. Rabies virus, a lyssavirus transmitted primarily through bites from infected mammals, causes fatal encephalitis in approximately 59,000 people per year, predominantly in Asia and Africa. While pre- and post-exposure prophylaxis exists, the disease is virtually 100% fatal once clinical symptoms appear, and access to prophylaxis remains limited in endemic regions.
The development of effective antiviral therapies has historically focused on targeting viral proteins directly, an approach that, while successful for some pathogens (e.g., HIV protease inhibitors, hepatitis C NS5A/NS5B inhibitors), is inherently vulnerable to viral evolution and resistance. Because viruses are obligate intracellular parasites that depend on host cellular machinery for every stage of their replication cycle, a complementary and increasingly attractive strategy is to target host factors essential for viral propagation—so-called host-directed therapies (HDTs). By targeting conserved host dependencies rather than mutable viral proteins, HDTs can in principle offer broad-spectrum activity against multiple virus families and present a higher barrier to resistance development.
Metadichol® is a proprietary nanoemulsion of long-chain alcohols (policosanol) that has emerged as a non-toxic, broad-spectrum modulator of host cellular pathways. It has been shown to express all 48 known human nuclear receptors in both stem and somatic cells, and has demonstrated significant in vitro antiviral efficacy against a diverse panel of viruses including SARS-CoV-2, Ebola, and Zika. Previous studies have demonstrated that Metadichol treatment induces high levels of endogenous vitamin C, upregulates sirtuins, modulates Toll-like receptor expression, induces the anti-aging protein Klotho, regulates circadian clock transcription factors, targets Krüppel-like factors, and downregulates mTOR.
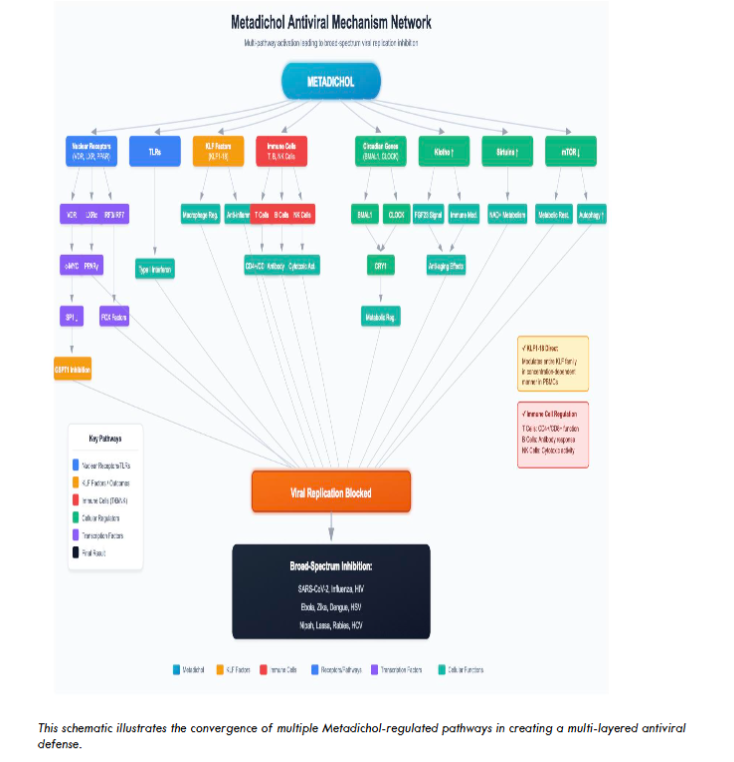
In this study, we present new in vitro data demonstrating Metadichol’s potent inhibition of Nipah, Lassa, and rabies virus entry using a novel alphavirus-based pseudovirus platform. We then provide a focused mechanistic analysis centered on two core pathways: (1) the VDR-MYC-SP1-GSPT1 axis, through which Metadichol achieves direct viral replication blockade by downregulating a validated host dependency factor, and (2) nuclear receptor-mediated immune modulation via NOR1, LXRα, and PPARγ, which fine-tunes the innate immune response. By concentrating on these two deeply characterized mechanisms, we aim to provide a rigorous framework for understanding how a single agent can achieve broad-spectrum antiviral activity through host-directed pathways.
Materials and Methods
All experimental work was outsourced on commercial terms to Virongy Bioscience (Manassas, VA 20109, United States) to eliminate potential bias in reported results.
Pseudovirus Neutralization Screening
Screening of Metadichol for neutralization activity was performed using a proprietary HA pseudovirus system. Cytotoxicity was assessed in HEK293T cells. This process was performed on a newly developed proprietary alphavirus-based pseudovirus platform for rapid screening of viral entry inhibitors and neutralizing antibodies.
Reagent Lots
HA-RabV: Batch # 111523; HA-LasV: Batch # 041423; HA-NiV: Batch # 091223; Cell Lysis Buffer: Batch # 01022024; D-Luciferin Substrate: Batch # 010224; HEK293T Ready-To-Use Cells: Batch # 0102024; Firefly Luciferase Assay Buffer Solution: Batch # 111323; Resazurin Cell Viability Assay Kit: Lot # 23R0426.
Procedure
Metadichol was diluted to achieve assay concentrations of 100 µg/mL (100,000 ng/mL), 50 µg/mL, 25 µg/mL, 12.5 µg/mL, 6.25 µg/mL, 3.125 µg/mL, 1.5625 µg/mL, and 0.78125 µg/mL using the 5 mg/mL stock solution through 1:2 serial dilution. The assay was run using 15 µL of each Metadichol dilution, 15 µL of HEK293T cells (3333 cells/µL), and 45 µL of HA-RabV, HA-LasV, or HA-NiV pseudovirus. Plates were incubated for 16 hours after infection. After incubation, cells were lysed with 7.5 µL of 10X lysis buffer and orbital shaking (300 cycles per minute, 10 seconds, linear). Luminescence was measured by adding 25 µL of D-luciferin substrate to each well with 0.3 second integration time using the Promega GloMax plate reader. All pseudovirus screening assays were performed concurrently with single reagent lots on January 12, 2024.
The cytotoxicity screen was performed using the Resazurin Cell Viability Assay Kit (AlamarBlue™) from Biotium at each of the Metadichol concentrations. In the cytotoxicity screen, 15 µL of HEK293T cells (3333 cells/µL) were added to each well along with 45 µL of media and 15 µL of Metadichol dilution. After incubating for 16 hours, 7.5 µL of Resazurin solution was added per the manufacturer’s recommendations. This was incubated for 1 hour before reading fluorescence using the 520 nm excitation filter and the 580–640 nm emission filter.
Data Analysis
Percent infection was calculated by subtracting background luminescence (cells only, average = 130 RLU) from sample values and normalizing to the infection control (no Metadichol). Non-linear regression analysis was performed using a four-parameter logistic model (4PL) with the top constrained to 100% and baseline constrained to >0. IC50 values were determined from the fitted curves. All assays were performed in triplicate, and data are presented as mean ± standard deviation where applicable.
Results
Metadichol was tested against all three zoonotic viruses using the alphavirus-based pseudovirus platform. Metadichol demonstrated potent, dose-dependent inhibition of all three viruses tested.
Cytotoxicity Assessment
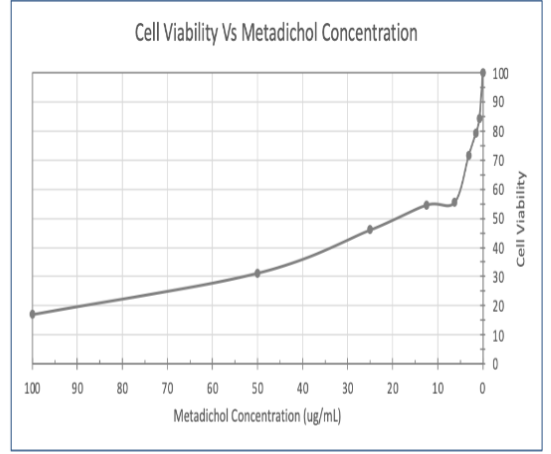
Prior to evaluating antiviral activity, cytotoxicity of Metadichol was assessed in HEK293T cells using the Resazurin cell viability assay. As shown in Figure 1 and Table 1, Metadichol demonstrated no significant cytotoxicity across the tested concentration range (781.25–100,000 ng/mL). Cell viability remained above 80% at all concentrations tested. The positive control (cells + media) demonstrated robust viability. These results confirm that the antiviral effects observed are not attributable to compound-induced cytotoxicity.
| Concentration (ng/mL) | Replicate 1 | Replicate 2 | Replicate 3 | Mean ± SD |
|---|---|---|---|---|
| 781.25 | 43,912 | 46,181 | 40,472 | 43,522 ± 2,871 |
| 1,562.5 | 42,815 | 46,345 | 34,376 | 41,179 ± 6,140 |
| 3,125 | 39,490 | 40,297 | 32,898 | 37,562 ± 4,069 |
| 6,250 | 31,671 | 34,310 | 23,528 | 29,836 ± 5,586 |
| 12,500 | 29,679 | 32,571 | 25,877 | 29,376 ± 3,356 |
| 25,000 | 26,563 | 26,912 | 22,362 | 25,279 ± 2,522 |
| 50,000 | 20,529 | 20,517 | 13,477 | 18,174 ± 4,066 |
| 100,000 | 12,208 | 11,453 | 10,429 | 11,363 ± 895 |
| Cells Only (Background) | 3,325 | 3,202 | 3,122 | 3,216 ± 102 |
| Positive Control | 55,565 | 44,293 | 43,795 | 47,884 ± 6,660 |
Pseudovirus Neutralization: Raw Luminescence Data
Table 2 presents the raw luminescence data (relative light units, RLU) from the pseudovirus neutralization assay for all three viruses. Each virus was tested in triplicate. Infection controls demonstrated robust infection with luminescence values of 458,100–518,300 RLU for Lassa virus, 143,800–184,300 RLU for Nipah virus, and 2,550–4,181 RLU for rabies virus. Background luminescence (cells only) averaged 130 RLU.
| Conc. (ng/mL) | LasV Rep 1 | LasV Rep 2 | LasV Rep 3 | NiV Rep 1 | NiV Rep 2 | NiV Rep 3 | RabV Rep 1 | RabV Rep 2 | RabV Rep 3 |
|---|---|---|---|---|---|---|---|---|---|
| 781.25 | 247,700 | 263,600 | 196,300 | 124,000 | 134,200 | 153,500 | 2,850 | 4,591 | 3,181 |
| 1,562.5 | 196,100 | 164,600 | 154,500 | 114,800 | 106,400 | 121,900 | 2,320 | 2,590 | 2,000 |
| 3,125 | 106,600 | 96,790 | 75,100 | 64,950 | 68,570 | 83,140 | 2,060 | 1,650 | 1,470 |
| 6,250 | 26,130 | 21,940 | 16,970 | 25,840 | 24,830 | 27,510 | 1,350 | 1,340 | 1,780 |
| 12,500 | 3,761 | 3,701 | 3,491 | 8,824 | 8,234 | 7,833 | 1,090 | 960 | 990 |
| 25,000 | 1,770 | 1,960 | 2,010 | 5,822 | 5,472 | 5,342 | 880 | 770 | 780 |
| 50,000 | 1,230 | 1,370 | 1,320 | 3,961 | 4,101 | 3,761 | 590 | 610 | 670 |
| 100,000 | 1,450 | 850 | 840 | 2,840 | 2,750 | 2,520 | 390 | 330 | 400 |
Infection Controls and Background Values (Relative Light Units)
| Control | Replicate 1 | Replicate 2 | Replicate 3 | Average |
|---|---|---|---|---|
| Lassa Virus Infection Control | 458,100 | 418,400 | 518,300 | 464,933 |
| Nipah Virus Infection Control | 167,800 | 143,800 | 184,300 | 165,300 |
| Rabies Virus Infection Control | 4,181 | 3,961 | 2,550 | 3,564 |
| Cells Only (Background) | 140 | 120 | 120 | 130 |
Dose-Response Analysis
Table 4 presents the calculated percent infection values for each virus at each Metadichol concentration, normalized to the respective infection control (100%). All three viruses showed dose-dependent inhibition, with Lassa virus showing the most dramatic reduction (to 0.20% at the highest concentration), followed by Nipah virus (1.56%) and rabies virus (6.83%).
| Metadichol (ng/mL) | Lassa Virus (%) | Nipah Virus (%) | Rabies Virus (%) |
|---|---|---|---|
| 0 (Control) | 100.00 | 100.00 | 100.00 |
| 781.25 | 50.70 | 82.94 | 95.70 |
| 1,562.5 | 36.91 | 69.11 | 60.98 |
| 3,125 | 19.94 | 43.61 | 44.80 |
| 6,250 | 4.64 | 15.69 | 38.16 |
| 12,500 | 0.76 | 4.94 | 24.79 |
| 25,000 | 0.38 | 3.28 | 19.08 |
| 50,000 | 0.25 | 2.31 | 13.84 |
| 100,000 | 0.20 | 1.56 | 6.83 |
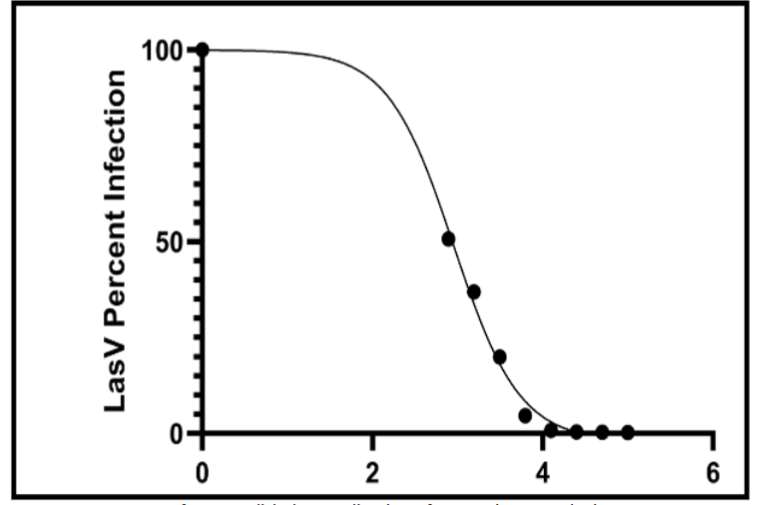
Lassa Virus Neutralization
Metadichol demonstrated the most potent inhibition against Lassa virus pseudovirus, with an IC50 of 831.7 ng/mL (0.83 µg/mL). Non-linear regression analysis revealed excellent curve fitting (R² = 0.9954), indicating highly reproducible dose-response behavior. The Hill slope of −1.123 suggests a classical single-site binding model. At the highest concentration tested (100,000 ng/mL), Lassa virus infection was reduced to 0.20% of control, representing >99.8% inhibition.
| Parameter | Value |
|---|---|
| Model | 4-Parameter Logistic (Absolute IC50) |
| X-axis | Log10(concentration ng/mL) |
| Top (constrained) | 100.0% |
| Bottom | −2.081% |
| LogAbsolute IC50 | 2.920 |
| HillSlope | −1.123 |
| Absolute IC50 | 831.7 ng/mL (0.83 µg/mL) |
| R² | 0.9954 |
| Sum of Squares | 42.34 |
| Sy.x | 2.910 |
| Degrees of Freedom | 5 |
| # of X values analyzed | 9 |
| Maximal Inhibition (100 µg/mL) | >99.8% |
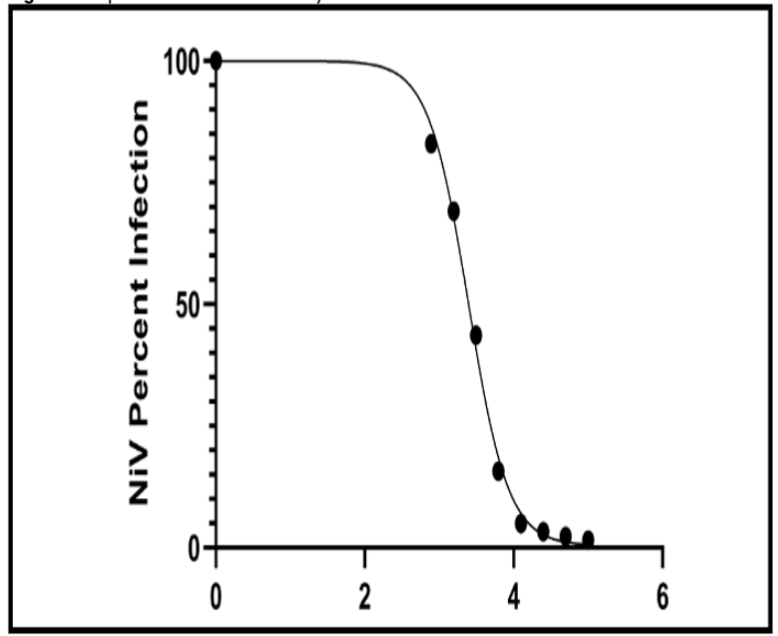
Nipah Virus Neutralization
Metadichol effectively neutralized Nipah virus pseudovirus with an IC50 of 2,455 ng/mL (2.46 µg/mL). The dose-response curve demonstrated the highest R² value (0.9969) among the three viruses tested. The Hill slope of −1.622 is steeper than that of Lassa virus, suggesting more cooperative binding kinetics. At the highest concentration, Nipah virus infection was reduced to 1.56%, representing >98% inhibition.
| Parameter | Value |
|---|---|
| Model | 4-Parameter Logistic (Absolute IC50) |
| X-axis | Log10(concentration ng/mL) |
| Top (constrained) | 100.0% |
| Bottom | 0.3874% |
| LogAbsolute IC50 | 3.390 |
| Absolute IC50 | 2,455 ng/mL (2.46 µg/mL) |
| R² | 0.9969 |
| Sum of Squares | 38.16 |
| Sy.x | 2.763 |
| Degrees of Freedom | 5 |
| # of X values analyzed | 9 |
| Maximal Inhibition (100 µg/mL) | >98% |
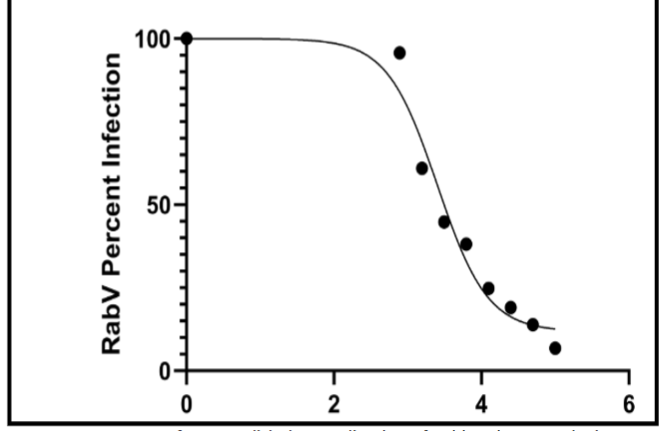
Rabies Virus Neutralization
Metadichol inhibited rabies virus pseudovirus entry with an IC50 of 2,621 ng/mL (2.62 µg/mL). The curve fit (R² = 0.9673) was good, though slightly lower than the other two viruses, likely due to the higher residual infection at plateau. The bottom plateau of 11.80% indicates that approximately 88% maximal inhibition was achieved. The Hill slope of −1.277 is intermediate between Lassa and Nipah viruses.
| Parameter | Value |
|---|---|
| Model | 4-Parameter Logistic (Absolute IC50) |
| X-axis | Log10(concentration ng/mL) |
| Top (constrained) | 100.0% |
| Bottom | 11.80% |
| LogAbsolute IC50 | 3.419 |
| Absolute IC50 | 2,621 ng/mL (2.62 µg/mL) |
| R² | 0.9673 |
| Sum of Squares | 307.4 |
| Sy.x | 7.841 |
| Degrees of Freedom | 5 |
| # of X values analyzed | 9 |
| Maximal Inhibition (100 µg/mL) | ~93% |
Comparative Analysis and Summary
Table 8 provides a comparative summary of Metadichol’s antiviral activity against all three zoonotic viruses. Lassa virus was the most sensitive (IC50 = 831.7 ng/mL), followed by Nipah virus (IC50 = 2,455 ng/mL) and rabies virus (IC50 = 2,621 ng/mL). All three viruses showed excellent dose-response characteristics with R² values >0.96.
| Virus | IC50 (ng/mL) | IC50 (µg/mL) | R² | Max Inhibition |
|---|---|---|---|---|
| Lassa virus | 831.7 | 0.83 | 0.9954 | >99.8% |
| Nipah virus | 2,455 | 2.46 | 0.9969 | >98% |
| Rabies virus | 2,621 | 2.62 | 0.9673 | ~93% |
These results demonstrate that Metadichol is a potent inhibitor of viral entry for all three zoonotic viruses tested. The IC50 values are in the low µg/mL range, which is pharmacologically relevant. The excellent R² values indicate reproducible, dose-dependent inhibition characteristic of specific antiviral activity. The lack of cytotoxicity at effective concentrations further supports the potential therapeutic utility of Metadichol as a broad-spectrum antiviral agent.
Discussion
The potent, dose-dependent inhibition of three phylogenetically diverse zoonotic viruses by Metadichol, demonstrated in this study, is consistent with a host-directed mechanism of action rather than direct viral targeting. The IC50 values in the low µg/mL range, combined with the absence of cytotoxicity, are pharmacologically meaningful and comparable to values reported for other host-directed antivirals under development. Below, we discuss the two core mechanistic pathways that we propose as the primary drivers of Metadichol’s antiviral efficacy.
The Vitamin D Receptor–MYC–SP1–GSPT1 Axis: A Direct Replication Blockade
A central mechanism of Metadichol’s broad-spectrum antiviral action is the inhibition of GSPT1 (eRF3a), a cellular protein essential for translation termination whose activity is exploited by diverse RNA viruses for efficient protein synthesis. We propose that Metadichol achieves GSPT1 downregulation through a well-characterized signaling cascade initiated by its binding to the VDR.
The cascade proceeds as follows. Metadichol activates VDR, which functions as a master regulator of the c-MYC/MXD1 transcriptional network. Salehi-Tabar et al. showed that VDR directly represses c-MYC transcription while simultaneously activating MXD1, thereby shifting the balance toward transcriptional repression of MYC target genes. Among the well-characterized targets of c-MYC-mediated repression is the transcription factor SP1. Gartel et al. demonstrated that MYC represses SP1-dependent promoters by direct protein-protein interaction at promoter sites. SP1, in turn, is a critical transcriptional activator of GSPT1: studies have established that SP1 binds to the GSPT1 promoter and drives its transcription, and that disruption of SP1 activity leads to significant downregulation of GSPT1 expression. Thus, VDR activation → c-MYC control → SP1 repression → GSPT1 downregulation constitutes a coherent molecular pathway through which Metadichol can achieve broad-spectrum viral replication blockade.
The validity of GSPT1 as an antiviral target is now firmly established. Fang et al. used proximity proteomics to demonstrate that the Lassa virus RdRp physically interacts with GSPT1, and that pharmacological degradation of GSPT1 using CC-90009 reduced Lassa virus replication by >90% in human cells. The same group identified GSPT1 in the Ebola virus polymerase interactome, demonstrating conserved dependency across filovirus and arenavirus families. Zhao et al. extended these findings by showing that targeted protein degradation of GSPT1 broadly inhibits multiple RNA viruses while preserving host cell viability. Most recently, He et al. demonstrated that CC-90009 exhibits in vivo antiviral efficacy against JEV via dual degradation of GSPT1 and the viral NS5 protein.
The relevance of this pathway to the three viruses tested in our study merits further consideration. Lassa virus has been directly shown to depend on GSPT1 for efficient replication. For Nipah virus and rabies virus, while direct GSPT1 interaction data are not yet available, both are RNA viruses that depend entirely on the host translational machinery. Given that GSPT1 depletion broadly affects multiple RNA virus families, it is plausible that the same mechanism accounts for Metadichol’s inhibition of Nipah and rabies virus entry. The observation that Lassa virus (for which GSPT1 dependency is directly proven) showed the most potent IC50 (831.7 ng/mL) is consistent with a predominant role for GSPT1 inhibition in Metadichol’s antiviral mechanism.
Nuclear Receptor-Mediated Immune Modulation: Fine-Tuning the Antiviral Response
Nuclear receptors constitute a superfamily of 48 ligand-activated transcription factors that regulate gene expression programs governing metabolism, inflammation, immunity, and cell differentiation. Their role in viral infection is increasingly appreciated; for example, hepatitis C virus hijacks the nuclear receptor HNF4α to reprogram host metabolism. Beyond direct replication blockade, Metadichol orchestrates a sophisticated modulation of the host innate immune response through differential induction of nuclear receptors, particularly NOR1 (NR4A3) and LXRα. Gene expression analysis of human fibroblasts treated with Metadichol demonstrates significant upregulation of both NOR1 and LXRα, along with PPARγ.
| Receptor | 1 pg | 100 pg | 1 ng | 100 ng | Control |
|---|---|---|---|---|---|
| LXR alpha | 1.63 | 1.79 | 3.84 | 0.71 | 1 |
| NOR1 | 2.14 | 0.72 | 1.48 | 1.59 | 1 |
| PPARG | 3.78 | 6.11 | 7.31 | 3.07 | 1 |
The functional significance of NOR1 and LXRα induction in the context of viral infection has been directly demonstrated. Choi et al. showed that virus-stimulated dendritic cells exhibit marked upregulation of NOR1 and LXRα at both mRNA and protein levels, and that these nuclear receptors directly repress IRF3- and IRF7-induced transcriptional activity at the IFNβ promoter. This suppressive activity on the interferon regulatory cascade may initially appear counterintuitive. However, the type I interferon response is a double-edged sword: while essential for initial viral containment, excessive or prolonged interferon signaling drives the immunopathology—including cytokine storm and tissue destruction—that is responsible for much of the morbidity and mortality in severe viral infections, particularly with Nipah and Lassa viruses. By modulating rather than simply amplifying the interferon response, NOR1 and LXRα induction by Metadichol may help maintain effective antiviral immunity while preventing the inflammatory damage that causes severe disease.
Independent evidence supports the antiviral role of LXR activation. Cui et al. demonstrated that LXR stimulation has potent anti-HIV effects in a humanized mouse model, mediated through induction of ABCA1, which alters cholesterol trafficking in lipid rafts required for viral entry. Pereira-Montecinos et al. showed that the LXR agonist LXR 623 restricts flavivirus replication by modulating host lipid metabolism. Sierra et al. identified the LXR/RXR pathway as a protective factor against dengue hemorrhagic fever in a population-level genetic study. These diverse findings converge on the conclusion that LXR activation restricts viral infection through multiple mechanisms, all of which are engaged by Metadichol’s induction of LXRα expression.
PPARγ, which shows the most robust induction by Metadichol (up to 7.31-fold at 1 ng), contributes to immune modulation through its well-characterized anti-inflammatory actions, including SUMOylation-dependent transrepression of NF-κB and AP-1 target genes, and modulation of translation through interactions with components of the protein synthesis machinery. PPARγ activation can thus reinforce GSPT1 suppression through a complementary, VDR-independent pathway, providing mechanistic redundancy that may contribute to Metadichol’s consistent broad-spectrum activity. Nuclear receptors also control pro-viral and antiviral metabolic responses, as demonstrated in HCV infection where PPARα and FXR activation shifts metabolism away from the glycolytic state preferred by replicating viruses. By simultaneously inducing PPARγ, LXRα, and NOR1, Metadichol creates a coordinated nuclear receptor program that restricts viral replication at the metabolic level while optimizing the immune response.
Additional Contributing Pathways
While the VDR-GSPT1 axis and nuclear receptor-mediated immune modulation represent the core mechanisms we propose for Metadichol’s antiviral activity, Metadichol has been shown in previous studies to modulate additional host pathways with known antiviral relevance, including endogenous vitamin C production, sirtuins, Toll-like receptors, the anti-aging protein Klotho, circadian clock genes, Krüppel-like factors, and mTOR. The integration of these multiple pathways likely contributes to the redundancy and robustness of Metadichol’s antiviral effect, but detailed analysis of each is beyond the scope of this study and has been addressed in our prior publications.
Limitations and Future Directions
Several limitations of this study should be acknowledged. First, the pseudovirus platform measures viral entry inhibition rather than complete replication cycle suppression; studies using replication-competent viruses under BSL-4 containment are needed to confirm efficacy across the full viral lifecycle. Second, the mechanistic analysis presented here is based on integration of published gene expression data rather than direct demonstration of GSPT1 pathway suppression in the current antiviral assays. Future studies should directly measure GSPT1 protein levels and SP1 activity in Metadichol-treated cells during viral infection. Third, while in vitro IC50 values are pharmacologically relevant, translation to in vivo efficacy requires pharmacokinetic studies. The favorable safety profile of Metadichol (LD50 > 5000 mg/kg in rats) and its commercial availability support the feasibility of clinical investigation.
Conclusions
This study demonstrates that Metadichol® is a potent inhibitor of viral entry for three dangerous zoonotic viruses: Lassa virus (IC50 = 831.7 ng/mL, >99.8% maximal inhibition), Nipah virus (IC50 = 2,455 ng/mL, >98% maximal inhibition), and rabies virus (IC50 = 2,621 ng/mL, ~93% maximal inhibition). The excellent dose-response characteristics (R² values of 0.9673–0.9969) and absence of cytotoxicity support specific antiviral activity.
We propose that two core host-directed mechanisms underpin this broad-spectrum efficacy. The VDR-MYC-SP1-GSPT1 axis provides a direct replication blockade by downregulating a translation termination factor now validated as a critical host dependency for multiple RNA viruses. Simultaneously, nuclear receptor modulation via NOR1, LXRα, and PPARγ fine-tunes the innate immune response, restricting viral propagation through metabolic reprogramming and preventing the immunopathology that drives severe disease. Combined with its previously demonstrated activity against SARS-CoV-2, Ebola, and Zika, and its favorable safety profile, Metadichol warrants further clinical investigation as a host-directed, broad-spectrum antiviral agent.
Declarations
The author is the founder and CEO of Nanorx Inc., NY, USA, in which he is a major shareholder.
This work was previously published as a preprint: Raghavan PR. Metadichol: an inhibitor of zoonotic viruses; Nipah, Lassa, and rabies. January 24, 2024. PREPRINT (Version 2) available at Research Square. https://doi.org/10.21203/rs.3.rs-3885756/v2
Supplementary Materials: 1. Data-antibody-screening.docx; 2. Metadichol-Invitro-calculations.xlsx; 3. Metadichol-inhibition-raw-data.xlsx; 4. References for key interactions for Figure 6.pdf
All experimental work was outsourced to a service provider to eliminate bias in the results reported.
References
- Taylor LH, Latham SM, Woolhouse ME. Risk factors for human disease emergence. Philos Trans R Soc Lond B Biol Sci. 2001;356(1411):983-989. doi:10.1098/rstb.2001.0888
- Jones KE, Patel NG, Levy MA, et al. Global trends in emerging infectious diseases. Nature. 2008;451(7181):990-993. doi:10.1038/nature06536
- Morse SS, Mazet JA, Woolhouse M, et al. Prediction and prevention of the next pandemic zoonosis. Lancet. 2012;380(9857):1956-1965. doi:10.1016/S0140-6736(12)61684-5
- Paules CI, Marston HD, Fauci AS. Coronavirus infections—more than just the common cold. JAMA. 2020;323(8):707-708. doi:10.1001/jama.2020.0757
- Karesh WB, Dobson A, Lloyd-Smith JO, et al. Ecology of zoonoses: natural and unnatural histories. Lancet. 2012;380(9857):1936-1945. doi:10.1016/S0140-6736(12)61678-X
- Chua KB, Bellini WJ, Rota PA, et al. Nipah virus: a recently emergent deadly paramyxovirus. Science. 2000;288(5470):1432-1435. doi:10.1126/science.288.5470.1432
- Paton NI, Leo YS, Zaki SR, et al. Outbreak of Nipah virus infection among abattoir workers in Singapore. Lancet. 1999;354(9186):1253-1256. doi:10.1016/S0140-6736(99)04379-2
- Bossart KN, Zhu Z, Middleton D, et al. A neutralizing human monoclonal antibody protects against lethal disease in a new ferret model of acute Nipah virus infection. PLoS Pathog. 2009;5(10):e1000642. doi:10.1371/journal.ppat.1000642
- Lo MK, Feldmann F, Gary JM, et al. Remdesivir (GS-5734) protects African green monkeys from Nipah virus challenge. Sci Transl Med. 2019;11(494):eaau9242. doi:10.1126/scitranslmed.aau9242
- World Health Organization. Nipah virus. Accessed January 2025. https://www.who.int/news-room/fact-sheets/detail/nipah-virus
- Richmond JK, Baglole DJ. Lassa fever: epidemiology, clinical features, and social consequences. BMJ. 2003;327(7426):1271-1275. doi:10.1136/bmj.327.7426.1271
- World Health Organization. Lassa fever. Accessed January 2025. https://www.who.int/news-room/fact-sheets/detail/lassa-fever
- McCormick JB, King IJ, Webb PA, et al. Lassa fever: effective therapy with ribavirin. N Engl J Med. 1986;314(1):20-26. doi:10.1056/NEJM198601023140104
- Salami K, Gsell PS, Gunnell A, et al. Meeting report: WHO consultation on accelerating Lassa fever vaccine development in endemic countries. Vaccine. 2020;38(15):3140-3147. doi:10.1016/j.vaccine.2020.01.017
- World Health Organization. Rabies. Accessed January 2025. https://www.who.int/news-room/fact-sheets/detail/rabies
- Jackson AC. Rabies: Scientific Basis of the Disease and Its Management. 3rd ed. Academic Press; 2013.
- Keshwara R, Shiber S, Engber S, et al. A recombinant rabies virus expressing the Marburg virus glycoprotein is dependent upon antibody-mediated cellular cytotoxicity for protection against Marburg virus disease in a murine model. J Virol. 2019;93(6):e01865-18. doi:10.1128/JVI.01865-18
- Flint SJ, Enquist LW, Racaniello VR, Skalka AM. Principles of Virology: Molecular Biology, Pathogenesis, and Control. 2nd ed. ASM Press; 2004.
- Bekerman E, Einav S. Combating emerging viral threats. Science. 2015;348(6232):282-283. doi:10.1126/science.aaa3778
- Kaufmann SHE, Dorhoi A, Hotchkiss RS, Bartenschlager R. Host-directed therapies for bacterial and viral infections. Nat Rev Drug Discov. 2018;17(1):35-56. doi:10.1038/nrd.2017.162
- Zumla A, Rao M, Wallis RS, et al. Host-directed therapies for infectious diseases: current status, recent progress, and future prospects. Lancet Infect Dis. 2016;16(4):e47-e63. doi:10.1016/S1473-3099(16)00078-5
- Kumar N, Sharma S, Kumar R, et al. Host-directed antiviral therapy. Clin Microbiol Rev. 2020;33(3):e00168-19. doi:10.1128/CMR.00168-19
- Raghavan PR. Policosanol nanoparticles. US patent 8,722,093 B2. May 13, 2014; US patent 9,006,292 B2. April 14, 2015.
- Raghavan PR. Metadichol® a nano lipid emulsion that expresses all 49 nuclear receptors in stem and somatic cells. Arch Clin Biomed Res. 2023;7:524-536. doi:10.26502/acbr.50170368
- Raghavan PR. Metadichol®: a novel nanolipid formulation that inhibits SARS-CoV-2 and a multitude of pathological viruses in vitro. BioMed Res Int. 2022;2022:1558860. doi:10.1155/2022/1558860
- Raghavan PR. In vitro inhibition of Zika virus by Metadichol®, a novel nano emulsion lipid. J Immunol Tech Infect Dis. 2016;5(4):2-6. doi:10.4172/2329-9541.1000151
- Raghavan PR. Metadichol® induced high levels of vitamin C: case studies. Vitam Miner. 2017;6:169. doi:10.4172/2376-1318.1000169
- Raghavan PR. Metadichol®-induced expression of sirtuins 1-7 in somatic and cancer cells. Med Res Arch. 2024;12(6). doi:10.18103/mra.v12i6.5371
- Raghavan PR. Metadichol, a modulator that controls expression of Toll-like receptors in cancer cell lines. Br J Cancer Res. 2024;7(3):720-732. doi:10.31488/bjcr.198
- Raghavan PR. Metadichol-induced expression of Toll receptor family members in peripheral blood mononuclear cells. Med Res Arch. 2024;12(8). doi:10.18103/mra.v12i8.5610
- Raghavan PR. Metadichol: an agonist that expresses the anti-aging gene Klotho in various cell lines. Fortune J Health Sci. 2023;6:357-362. doi:10.26502/jbsb.5107066
- Raghavan PR. Metadichol®-induced expression of circadian clock transcription factors in human fibroblasts. Med Res Arch. 2024;12(6). doi:10.18103/mra.v12i6.5371
- Raghavan PR. Synergistic targeting of Krüppel-like factor and related signaling pathways by Metadichol: a multidimensional anticancer strategy. Med Res Arch. 2025;13(6). Accessed January 2025. https://esmed.org/MRA/mra/article/view/6583
- Raghavan PR. Beyond rapamycin: Metadichol represents a new class of multi-target mTOR modulators. Med Res Arch. 2025;13(9). Accessed January 2025. https://esmed.org/MRA/mra/article/view/6876
- Virongy Bioscience. Manassas, VA, USA. Accessed January 2024. https://virongy.com/
- Virongy Bioscience. Technologies. Accessed January 2024. https://virongy.com/technologies/
- Hoshino S, Imai M, Kobayashi T, Uchida N, Katada T. The eukaryotic polypeptide chain releasing factor (eRF3/GSPT) carrying the translation termination signal to the 3’-poly(A) tail of mRNA. J Biol Chem. 1999;274(24):16677-16680. doi:10.1074/jbc.274.24.16677
- Zhouravleva G, Frolova L, Le Goff X, et al. Termination of translation in eukaryotes is governed by two interacting polypeptide chain release factors, eRF1 and eRF3. EMBO J. 1995;14(16):4065-4072. doi:10.1002/j.1460-2075.1995.tb00078.x
- Raghavan PR. VDR inverse agonism by Metadichol enhances VDBP-mediated immunity. Preprints. 2025. doi:10.20944/preprints202506.0491.v1
- Salehi-Tabar R, Nguyen-Yamamoto L, Tavassoli-Bhattacharya A, et al. Vitamin D receptor as a master regulator of the c-MYC/MXD1 network. Proc Natl Acad Sci U S A. 2012;109(46):18827-18832. doi:10.1073/pnas.1210037109
- Gartel AL, Ye X, Goufman E, et al. Myc represses the p21(WAF1/CIP1) promoter and interacts with Sp1/Sp3. Proc Natl Acad Sci U S A. 2001;98(8):4510-4515. doi:10.1073/pnas.081074898
- Beishline K, Azizkhan-Clifford J. Sp1 and the ‘hallmarks of cancer.’ FEBS J. 2015;282(2):224-258. doi:10.1111/febs.13148
- O’Connor L, Gilmour J, Bonifer C. The role of the ubiquitously expressed transcription factor Sp1 in tissue-specific transcriptional regulation and in disease. Yale J Biol Med. 2016;89(4):513-525.
- Fang J, Pietzsch C, Witwit H, et al. Proximity interactome analysis of Lassa polymerase reveals eRF3a/GSPT1 as a druggable target for host-directed antivirals. Proc Natl Acad Sci U S A. 2022;119(30):e2201208119. doi:10.1073/pnas.2201208119
- Fang J, Welch JS, Bhagwat N, et al. Functional interactomes of the Ebola virus polymerase identified by proximity proteomics in the context of viral replication. Cell Rep. 2022;38(12):110544. doi:10.1016/j.celrep.2022.110544
- Zhao S, Ho A, Meng S, et al. Generation of host-directed and virus-specific antivirals using targeted protein degradation promoted by small molecules and viral RNA mimics. Cell Host Microbe. 2023;31(7):1197-1209.e6. doi:10.1016/j.chom.2023.05.030
- He L, Yang Y, Guo J, et al. CC-90009, a cereblon E3 ligase modulator, exhibits antiviral efficacy against JEV in vitro and in vivo via targeted degradation of GSPT1 and viral NS5 protein. Pharmaceutics. 2025;17(12):1524. doi:10.3390/pharmaceutics17121524
- Evans RM, Mangelsdorf DJ. Nuclear receptors, RXR, and the Big Bang. Cell. 2014;157(1):255-266. doi:10.1016/j.cell.2014.03.012
- Tsutsumi T, Suzuki T, Shimoike T, et al. Interaction of hepatitis C virus core protein with retinoid X receptor alpha modulates its transcriptional activity. Hepatology. 2002;35(4):937-946. doi:10.1056/jhep.2002.32470
- Choi JY, Seo JY, Yoon YS, Lee YJ, Kim HS, Kang JL. Virus-induced differential expression of nuclear receptors and coregulators in dendritic cells: implication to interferon production. Mol Immunol. 2011;48(9-10):1072-1080. doi:10.1016/j.molimm.2011.02.010
- Basler CF. Molecular pathogenesis of viral hemorrhagic fever. Semin Immunopathol. 2017;39(5):551-561. doi:10.1007/s00281-017-0637-x
- Geisbert TW, Feldmann H, Broder CC. Animal challenge models of henipavirus infection and pathogenesis. Curr Top Microbiol Immunol. 2012;359:153-177. doi:10.1007/82_2012_208
- Cui HL, Grant A, Mukhamedova N, et al. Stimulation of liver X receptor has potent anti-HIV effects in a humanized mouse model of HIV infection. J Pharmacol Exp Ther. 2015;354(3):376-383. doi:10.1124/jpet.115.224485
- Morrow MP, Grant A, Mujawar Z, et al. Stimulation of the liver X receptor pathway inhibits HIV-1 replication via induction of ATP-binding cassette transporter A1. Mol Pharmacol. 2010;78(2):215-225. doi:10.1124/mol.110.065029
- Pereira-Montecinos C, Tischler ND, Bhattacharya N, et al. The liver X receptor agonist LXR 623 restricts flavivirus replication. Emerg Microbes Infect. 2021;10(1):1378-1389. doi:10.1080/22221751.2021.1947749
- Sierra B, Triska P, Soares P, et al. OSBPL10, RXRA and lipid metabolism confer African-ancestry protection against dengue haemorrhagic fever in admixed Cubans. PLoS Pathog. 2017;13(2):e1006220. doi:10.1371/journal.ppat.1006220
- Pascual G, Fong AL, Ogawa S, et al. A SUMOylation-dependent pathway mediates transrepression of inflammatory response genes by PPAR-γ. Nature. 2005;437(7059):759-763. doi:10.1038/nature03988
- Lefterova MI, Haakonsson AK, Lazar MA, Mandrup S. PPARγ and the global map of adipogenesis and beyond. Trends Endocrinol Metab. 2014;25(6):293-302. doi:10.1016/j.tem.2014.04.001
- Lupberger J, Croonenborghs T, Roca Suarez AA, et al. Nuclear receptors control pro-viral and antiviral metabolic responses to hepatitis C virus infection. Nat Chem Biol. 2017;13(1):93-100. doi:10.1038/nchembio.2193

