Advancements in Trigeminal Neuralgia Diagnosis and Biomarkers
Bridging Imaging and Molecular Biomarkers in Trigeminal Neuralgia: Toward Precision Diagnosis and Prognostication in Neuropathic Pain
Matthew Abikenari, M.S¹, Bhav Jain, B.S¹, Risheng Xu, M.D, Ph.D², Christopher Jackson, M.D², Judy Huang, M.D², Chetan Bettegowda, M.D, Ph.D², Michael Lim, M.D¹
- Department of Neurosurgery, Stanford University School of Medicine, Stanford, California, USA
- Department of Neurosurgery, John Hopkins University School of Medicine, Baltimore, Maryland, USA
OPEN ACCESS
PUBLISHED: 31 May 2025
CITATION: Abikenari, H., et al., 2025.
Bridging Imaging and Molecular Biomarkers in Trigeminal Neuralgia: Toward Precision Diagnosis and Prognostication in Neuropathic Pain. Medical Research Archives, [online] 13(5).
https://doi.org/10.18103/mra.v1i3.6605
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v1i3.6605
ISSN 2375-1924
ABSTRACT
Despite advances in imaging and clinical assessment, current diagnostic paradigms lack precision needed to subcategorize disease or predict therapeutic response with accuracy. This review proposes precision neuropathic pain diagnostics through the combination of artificial intelligence-guided imaging and molecular biomarker identification, using the use-case of Trigeminal neuralgia (TN), a refractory craniofacial pain disorder characterized by paroxysmal, one-sided facial pain and variable treatment response. We use trigeminal neuralgia as a case example to explore how deep learning, advanced imaging, and molecular profiling can work together to improve the diagnosis and treatment of neuropathic pain.
First, the current paper reviews contemporary advances in deep learning for neuroimaging, particularly convolutional neural networks and U-Net architectures that have enabled automatic segmentation of the trigeminal nerve and paved the way for radiomic measurement of neurovascular compression. Such advances optimize the objectivity of surgery stratification and help to distinguish classical from idiopathic Trigeminal Neuralgia. In parallel, proteomic profiling of cerebrospinal fluid and plasma has revealed TN-specific molecular signatures, including upregulation of inflammatory, stress-related, and axonal damage markers that are different from those of similar disorders such as multiple sclerosis.
Both markers not only have implications for disease pathobiology but also for identifying new therapeutic targets. The current study proposes a multimodal data integration platform combining imaging and molecular phenotypes using machine learning and multiomics platforms. The integration permits mechanistic subtyping and predictive modeling of treatment response, with potential applications expanding to diabetic neuropathy and complex regional pain syndrome. Clinical deployment challenges, such as heterogeneity of data, ambiguity regarding regulation, and ethical risk, are addressed along with near-term solutions such as federated learning and interoperable biomarker registries.
At the intersection of neurosurgery, radiology, and computational science, Trigeminal Neuralgia offers a scalable model for precision pain medicine, transforming care through mechanism-based classification and patient-stratified interventions.
Keywords: Trigeminal Neuralgia, Neuropathic Pain, Precision Medicine, Deep Learning, Biomarkers, Multimodal Integration
Introduction
Trigeminal neuralgia (TN) is a disabling, excruciating craniofacial pain syndrome that is an episodic, electric shock-like pain paroxysmally occurring in the distribution of one or more divisions of the trigeminal nerve (CN V)¹². The pain is abrupt in onset and often precipitated by innocuous stimuli such as light touch, speaking, chewing, or even wind exposure³⁴. Such characteristic triggering features are found in as many as 79% of cases, are an important diagnostic feature, and are most often located in the perioral and nasal regions, corresponding to the preferential involvement of the V2 (maxillary) and V3 (mandibular) divisions⁵⁷.
Spontaneous episodes of pain may occur but are less common. Pain is distributed into the weblike pattern of the areas of the skin elsewhere referred to as the dermatome V1 [ophthalmic], V2 [maxillary], and V3 [mandibular], with V2 and V3 being the most common presentations. Cases of V1-only involvement are uncommon (around 4%) and should prompt consideration of other diagnoses, including trigeminal autonomic cephalalgias⁸¹⁰. Pain is usually unilateral, and the incidence of bilateral TN is low except for secondary causes such as multiple sclerosis. Around 50% of patients with TN experience background pain that is persistent, dull, or burning, interspersed with episodes of a band-like paroxysmal pain called TN with concomitant continuous pain¹¹¹². This phenotype is more common in younger age groups and women, and it is characterized by greater therapeutic demand and reduced responsiveness to standard treatments¹³¹⁵. However, such variability of clinical manifestations with both remitting and relapsing phases complicates the diagnostic process and calls for careful phenotyping.
Most patients with idiopathic or classical TN show no demonstrable deficit on neurological examination. Neurological examination is characteristically normal, but minor sensory abnormalities (including mild cases of hypoesthesia or hyperesthesia) may be present in 10–30% of cases¹⁶. The occurrence of severe or worsening sensory loss, however, might indicate secondary causes such as multiple sclerosis, neoplasms, and postherpetic neuropathy, leading to further diagnostic imaging and workup¹⁷¹⁸. Magnetic resonance imaging (MRI) of the brainstem is the cornerstone of diagnostic investigation in TN, which allows clinicians to appreciate the neurovascular contact, generally observed between the transverse portion of the trigeminal nerve at the root entry zone and the superior cerebellar artery¹⁹²⁰. High-resolution sequences (3D T2, 3D T1 with gadolinium, time-of-flight angiography) are useful in confirming classical TN, defined by morphological changes of the nerve, including distortion or atrophy, in concordance with the clinical pattern observed²¹²².
Importantly, neurovascular contact does not always lead to the presentation of TN symptoms, as contact is also seen in asymptomatic subjects²³. Therefore, the diagnosis is more confidently made when MRI findings and further imaging correspond to clinical laterality and pain distribution. Trigeminal reflex testing might offer ancillary diagnostic value when MRI is contraindicated or not available, particularly in differentiating TN from painful trigeminal neuropathy or demyelinating diseases. Yet, it is not specific enough to distinguish classical and idiopathic TN and should be reserved for atypical cases²⁴²⁵.
TN is characterized by overlapping symptomatology with other facial pain syndromes, including painful post-traumatic trigeminal neuropathy, glossopharyngeal neuralgia, persistent idiopathic facial pain, orofacial pain with dental or temporomandibular involvement²⁶. Hence, a comprehensive clinical evaluation is crucial. Overdiagnosis of TN, especially in the setting of neurosurgery, may occur due to the overinterpretation of imaging findings without sufficient clinical correlation²⁷²⁸. This underscores the necessity of an extensive workup, which needs to include meticulous patient history, precise drawing of the sensory field on examination, and exclusion of dental pathologies.
Despite increasing awareness, TN still presents a diagnostic dilemma, particularly for dental and
primary care physicians. Insufficiently validated screening tools and many inappropriate online patient resources highlight the requirement for improved diagnostic algorithms and patient resources²⁹³⁰. Early and accurate diagnosis is crucial since cases with both surgical and non-surgical indications vary dramatically according to the TN phenotype. Given that traditional clinical and radiological parameters are limited in terms of sensitivity and specificity, there is a developing interest in combining molecular biomarker research with advanced image acquisition techniques for improved diagnosis, prognostication, and treatment stratification of TN. New advances in nerve segmentation using deep learning machine learning techniques, together with cerebrospinal fluid and plasma-based proteomics, provide a roadmap for multimodal precision diagnostics³¹³³. Here we present TN as an exemplary use case, inviting further inspection of how artificial intelligence, high-resolution imaging, and molecular profiling might converge to address wider challenges of neuropathic pain syndromes.
Deep Learning in Neuropathic Pain Imaging: Current Landscape
Deep learning (DL) models have revolutionized the interpretation of medical images, notably in domains with the need for high-resolution anatomical precision, such as neuropathic pain syndromes. Methods based on convolutional neural networks (CNNs), U-Net architectures, as well as attention-based systems, have become essential for segmenting peripheral and central nervous system structures, detecting microstructural abnormalities, or extracting quantitative radiomic features that are related to the respective clinical phenotypes³²³⁶. In the setting of TN, DL has held promise in automating the segmentation of the trigeminal nerve from high-resolution T2-weighted and diffusion tensor imaging (DTI) volumes in the past, which require extensive manual and time-consuming training tasks³²³⁵³⁶.
Over the last year, U-Net-based networks have been used for the delineation of the Root Entry Zone (REZ) of the trigeminal nerve and nearby vascular loops and volume calculation of the neurovascular contact and asymmetry metrics³⁴. Such novel approaches may help in identifying objective biomarkers for the differential diagnosis of classical and idiopathic TN and of surgical indication. Besides TN, DL-based radiomics personalized the differential diagnosis of classical and idiopathic TN and of surgical indication. Besides TN, DL-based radiomics personalized the differential diagnosis of classical and idiopathic TN and of surgical indication. Besides TN, DL-based radiomics have also shown value in other neuropathic pain diseases. In sciatica, CNN-based classification models trained on lumbar MRI have been able to predict the compression of the nerve roots and the severity of the related pain³⁵. Additionally, in the field of diabetic neuropathy, DL models fusing MRI and nerve conduction features have increased the diagnostic accuracy and risk stratification³⁷³⁸. These findings highlight the promise of multimodal DL pipelines in capturing subtle structural and functional changes that may be missed by the human eye (Table 1).
However, various obstacles limit the extent of their clinical application. A significant limitation is the variability of imaging protocols within and across institutions, restricting the generalizability of the model. Small sample sizes and a lack of external validation are common problems, especially in rare diseases such as TN. Third, DL is not intuitively interpretable, which in turn reduces DL’s potential for adoption into regulation and trust among clinicians³⁹. These challenges may be partially addressed by combining multi-center learning frameworks and explainable AI techniques. Insistence on multicenter harmonization of imaging protocols, substantial external validation cohorts, and prospective evaluation as part of clinical decision pathways will be needed before exhaustive deployment into the medical field.
Table 1. Deep Learning Applications in Neuropathic Pain Imaging. Table summarizes key insights demonstrating the role of DL models in imaging-based diagnosis and stratification of neuropathic pain conditions, along with clinical benefits and current limitations.
| Neuropathic Pain Condition | Imaging Modality | DL Model Used | Clinical Value | Limitations |
|---|---|---|---|---|
| Trigeminal Neuralgia (TN) | High-res T2-weighted MRI, DTI | U-Net, CNNs | Automated segmentation of the trigeminal nerve; stratification of classical vs idiopathic TN | Small sample sizes, limited generalizability across centers, and interpretability concerns |
| Sciatica | Lumbar MRI | CNN-based classifiers | Prediction of nerve root compression and pain severity | Lack of standardization; model transferability issues |
| Diabetic Neuropathy | MRI + Nerve Conduction Studies | Fusion DL models (MRI + NCS) | Enhanced diagnostic accuracy and patient risk stratification | Integration challenges of multimodal inputs; need for large external datasets |
DL = Deep Learning; MRI = Magnetic Resonance Imaging; DTI = Diffusion Tensor Imaging; CNN = Convolutional Neural Network; NCS = Nerve Conduction Studies.
Molecular Biomarkers in Neuropathic Pain: Proteomics and Beyond
In tandem with imaging, molecular diagnostics for neuropathic pain have been progressing rapidly with the development of high-throughput proteomics, metabolomics, and transcriptomic technologies⁴⁰⁴¹. In TN, profiling of the proteome of cerebrospinal fluid (CSF) and serum has revealed different molecular signatures that distinguish TN from healthy controls and neuroinflammatory controls, such as individuals with multiple sclerosis (MS)⁴². Also, Lafte et al. used the proximity extension assay (PEA) technology to study 72 neurologically relevant proteins in TN, MS, and control individuals. Among the 19 TN patients analyzed, 15 exhibited upregulation of proteins linked to neuroinflammation (SFRP1), glucocorticoid signaling and stress induction (FKBP5), as well as cytoskeletal remodeling of cells (TBCB), reflecting mutual pathophysiologic factors involving immune activation, chronic stress, and axonal degeneration⁴². Interestingly, surgical treatment by microvascular decompression tended to normalize several of these protein levels, supporting their disease association rather than reflecting downstream effects of chronic pain behavior.
Furthermore, the very limited overlap of trigeminal neuralgia and multiple sclerosis proteomic signatures calls into question the conventional wisdom of common inflammatory etiologies, favoring the concept of biologic heterogeneity. In addition to TN, other neuropathic pain syndromes have shown proteomic and metabolic abnormalities in both CSF and plasma. In complex regional pain syndrome (CRPS), there is increased pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6, and components of the complement cascade such as C3a, C5a, and C1q. In diabetic neuropathy, however, there is upregulation of the markers of oxidative stress (8-isoprostane, MDA), mitochondrial damage (cytochrome c, ATP synthase subunits), and deranged lipid metabolism (ceramides, acylcarnitines)⁴³⁴⁵.
These observations support a model in which distinct molecular cascades underlie divergent neuropathic pain phenotypes, ultimately converging on shared downstream effectors such as neuronal sensitization and glial activation. Therapeutic targets aligned with these mechanisms have been increasingly identified through molecular profiling and differential gene expression studies⁴⁶. As an inhibitor of Wnt signaling and microglia–astrocyte crosstalk, SFRP1 may contribute to sustaining neuroinflammation in TN and might serve as a new therapeutic target⁴⁷. FKBP5, previously associated with glucocorticoid resistance and stress-induced synaptic remodeling, has received much attention in both psychiatric and neurodegenerative disorders, and it may be similarly important, and under study, in chronic pain⁴⁸. Furthermore, changes in inflammation-related molecules such as GRO-alpha and EGF have been shown to accompany symptom remission in late-life depression, further recapitulating the broader relevance of immune changes in treatment response across neurologic and neuropsychiatric conditions and neuropathic pain syndromes⁴⁹. Such findings may be used to inform biomarker-based clinical trials, in which molecular genotyping guides the selection of patients, monitoring of response to treatment, and development of novel pain management therapeutics (Figure 1).
Figure 1. Precision Diagnostics in Trigeminal Neuralgia: Multimodal Integration of Imaging, Biomarkers, and Deep Learning. This figure illustrates a diagnostic pathway for accuracy in trigeminal neuralgia (TN) involving high-resolution imaging, deep learning analysis, and molecular biomarker profiling. Convolutional neural networks and U-Net segmentation improve automated REZ compression detection and support TN subtype classification. Simultaneously, proteomic analysis of cerebrospinal fluid and serum detects disease-specific biomarkers associated with neuroinflammation, stress signaling, and axonal injury. Machine learning integrates radiomic and molecular data to develop disease phenotypes that inform surgical candidacy and therapeutic response prediction. Figure created using BioRender.com.
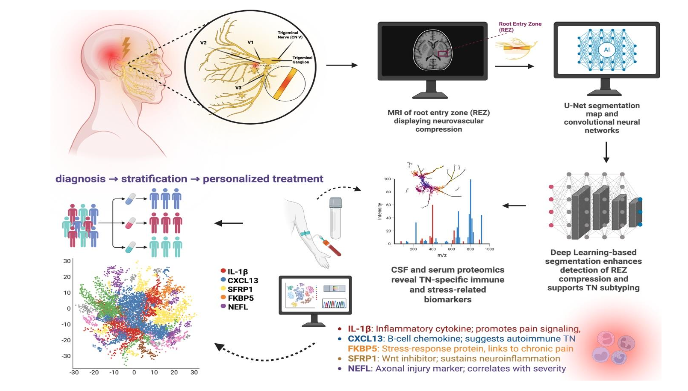
Table 1. Deep Learning Applications in Neuropathic Pain Imaging. Table summarizes key insights demonstrating the role of DL models in imaging-based diagnosis and prognostication of neuropathic pain conditions.
Abbreviations – IL-1β: Interleukin-1 beta (inflammatory cytokine); CXCL13: C-X-C motif chemokine ligand 13 (B-cell chemokine); FKBP5: FK506-binding protein 5 (stress-response regulator); SFRP1: Secreted frizzled-related protein 1 (Wnt signaling inhibitor); NEFL: Neurofilament light chain (axonal damage marker); REZ: Root Entry Zone; CSF: Cerebrospinal Fluid; TN: Trigeminal Neuralgia; AI: Artificial Intelligence.
Integrating Imaging and Molecular Biomarkers: Toward Precision Neuropathic Pain Care
The whole picture of neuropathic pain demands an integration of structural, functional, and molecular dimensions. High-resolution imaging techniques, such as diffusion tensor imaging (DTI), magnetic resonance neurography (MRN), and high-resolution 3D MRI, enable the characterization of trigeminal nerve compression, demyelination, and central white matter changes²⁶. Concurrently, proteomic and transcriptomic studies have identified molecular derangements in the neurovascular interface of TN patients undergoing microvascular decompression (MVD), such as upregulation of axonal injury markers (NEFL), endothelial signaling proteins (vWF, ICAM1), and inflammatory mediators such as CXCL13 and IL-1β⁵⁰.
This multimodal approach allows for the generation of disease phenotypes that link radiographic abnormalities with molecular states (Table 2). For instance, a patient with loss of DTI-derived fractional anisotropy in the trigeminal nerve root entry zone with an increase in markers of axonal degeneration in CSF or peripheral blood would be a mechanistic subtype that would be optimally treated by decompressive rather than ablative procedures, such as microvascular decompressions (MVD) through retrosigmoid craniotomy⁵¹.
Table 2. Key Molecular Biomarkers in Trigeminal Neuralgia.
Biomarkers were selected based on their relevance to neuroinflammation, immune dysregulation, stress signaling, and axonal injury in trigeminal neuralgia. These candidates may support subtype stratification and guide biomarker-informed treatment strategies.
| Biomarker | Pathophysiologic Role | Clinical Implications |
|---|---|---|
| IL-1β | Pro-inflammatory cytokine; contributes to glial activation | Potential marker of inflammatory subtype; candidate for immunomodulatory treatment |
| CXCL13 | B cell chemokine; involved in meningeal inflammation | Suggests autoimmune-like features; differentiator from idiopathic TN |
| FKBP5 | Stress response mediator; linked to glucocorticoid signaling | May serve as a stress-related biomarker; possible psychiatric overlap |
| SFRP1 | Wnt pathway inhibitor; modulates microglial/astrocyte interactions | Therapeutic target for sustaining neuroinflammation |
| TBCB | Cytoskeletal regulation; implicated in axonal degeneration | Supports the axonal degeneration mechanism in TN |
| NEFL | Marker of axonal injury; reflects white matter damage | Predictor of surgical response; correlates with imaging changes |
IL-1β = Interleukin-1 beta; CXCL13 = C-X-C motif chemokine ligand 13; FKBP5 = FK506-binding protein 5; SFRP1 = Secreted frizzled-related protein 1; TBCB = Tubulin-folding cofactor B; NEFL = Neurofilament light chain.
Findings in TN may be generalizable to other neuropathic pain syndromes, such as diabetic peripheral neuropathy and complex regional pain syndrome (CRPS), that share molecular signatures, such as axonal degeneration, neuroinflammation, and aberrant glial signaling⁴³⁴⁵⁵². Radiomic features from high-resolution MR neurography or spinal imaging may unmask conserved structural patterns (such as dorsal root ganglia hypertrophy, white matter tract reorganization) among them. Moreover, proteomic markers identified in TN, such as S100B, PRDX1, and GFAP, may be universal biomarkers of neuropathic pain severity and therapeutic response. Hence, the application of integrative machine learning methods to these datasets allows for elucidation of common pathogenic mechanisms, prediction of therapeutic response, and the development of unified biomarker panels for application across a spectrum of pain etiologies.
Graph-based and deep learning artificial intelligence (AI) models enable the integration of heterogeneous datasets spanning imaging, proteomics, genomics, and clinical variables. Autoencoders, CNNs, and attention-based transformers represent some of the models that can learn complex interactions between radiomic and molecular features, enabling exact stratification of patient subtypes⁵³⁵⁵. Multi-omics platforms also strengthen such models by adding transcriptomic, proteomic, and epigenomic data layers, resulting in unprecedented granularity in the description of neuropathic pain biology⁵⁶. In TN, for example, the fusion of pre-operative MRI radiomics and label-free quantification (LFQ) proteomics of CSF or blood samples was able to predict MVD responders from non-responders⁵⁰. Such models can also be used to guide treatment in CRPS or diabetic neuropathy by identifying those patients most likely to respond to immune-modulating therapies or neuromodulation.
Clinical Translation: Challenges and Opportunities
Despite the potential of multimodal data fusion, clinical translation is held back by formidable barriers. These include a lack of standardization across imaging modalities, limited interoperability of electronic health records, batch effects of proteomic assays, and a shortage of validated thresholds for interpretation of radiomic or molecular biomarkers. Uncertainty regarding the regulation of AI-based diagnostic devices also prevents full implementation of such techniques⁵⁷.
To address these issues, federated learning is a potential solution. This distributed machine learning framework allows institutions to collaboratively train models across locally retained data without sharing sensitive patient information⁵⁸⁵⁹. Such strategies are particularly relevant in neuropathic pain research, in which cohort sizes at an individual center are generally small and heterogeneous. Federated learning can aid model generalizability while preserving privacy and embracing institutional variability in imaging and proteomic protocols. As AI and biomarker-guided decision tools become more widely adopted in clinical practice, it is essential to address ethical concerns in advance. Such concerns encompass potential algorithmic prejudice, underrepresentation of minority groups in training datasets, and low-specificity biomarker signatures leading to medical overdiagnosis. Clear model explainability and oversight frameworks are critical to the fair deployment of such technologies⁵⁷⁶⁰⁶¹.
A sequential approach to using these tools in the clinic could be:
(1) establishing the validity of combined imaging and molecular markers in real patients;
(2) using these markers to guide treatment choices such as MVD surgery vs. radiation for trigeminal neuralgia, or immunotherapy vs. nerve ablation for complex regional pain syndrome; and
(3) building shared databases that integrate imaging, laboratory results, and outcomes. This strategy could begin with patients who have severe, treatment-resistant, and medication-refractory neuropathic pain, where no alternative managements currently exist.
Future Directions in Neuropathic Pain Research
Achieving precision in pain medicine will require next-generation, large-scale trials that integrate imaging, proteomics, single-cell transcriptomics, and clinical outcomes across diverse patient cohorts. Improved single-cell proteomics and spatial transcriptomics will enable cell-type-specific mapping of neuropathic pain drivers in pain-affected nerves, DRG, and spinal cord segments⁶². Concurrently, radiogenomic models that link genotype, molecular phenotype, and imaging biomarkers are poised to enhance subtype-specific diagnosis and guide individualized treatment. Within TN specifically, they may elucidate unresolved biological heterogeneity, such as why some patients manifest exclusively paroxysmal pain and others develop continuous pain, or why just a proportion of patients respond therapeutically in a stable fashion to surgical decompression. Radiogenomic frameworks incorporating MRI-derived neurovascular contact patterns with transcriptomic and proteomic data could potentially delineate mechanistic TN subtypes to inform patient-stratified therapy.
More broadly, these radiogenomic approaches will be capable of enhancing subtype-specific diagnosis across neuropathic pain syndromes and identifying predictive biomarkers of treatment response⁶²⁶³. Large-scale initiatives such as NIH’s Bridge2AI and Pain Consortium⁶⁴ or the European IMI-PainCare platforms can form the foundation for data-sharing consortia, model training, and cross-validation across diverse institutions. Open-source, privacy-preserving repositories will be necessary to allow for equitable global participation and ensure TN and other focal neuropathic syndromes are represented.
Conclusion
The integration of high-resolution imaging with molecular and proteomic biomarkers offers an unprecedented chance to transform the treatment of neuropathic pain patients. By bridging molecular phenotypes to structural change, clinicians can move beyond symptom-based diagnosis to mechanism-based precision medicine. Instantiation of such precision-based prognostication of neuropathic pain can only be achieved through a combined team effort of neurology, neurosurgery, radiology, bioinformatics, and ethics in defining, validating, and implementing multimodal approaches for guiding diagnosis, predicting treatment effects, and personalizing care trajectories. By integrating high-level machine learning, inter-professional collaboration, and standardization of diagnostic data (e.g., MRI sequences), the practice of neuropathic pain medicine can provide substantial improvements in diagnostic precision, therapeutic efficacy, and long-term patient outcomes, with the hope of ultimately setting a new standard for personalized neurological care.
Conflict of Interest Disclosure:
No pertinent conflicts of interest relevant to this manuscript.
Chetan Bettegowda is a consultant for Bionaut Labs, Haystack Oncology and Privo Technologies. CB is a co-founder of Belay Diagnostics and OrisDx. Michael Lim (Funding from Arbor Pharmaceuticals, Accuray, BMS, Novartis; Consultant: BMS, Merck, SQZ Biotechnologies, Tocagen, VBI; Patents: Combining Focused Radiation and Immunotherapy, Combining Local Chemotherapy and Immunotherapy; Shareholder: Egret Therapeutics).
Financial Disclosure:
None.
Funding Statement:
None.
Author Contributions:
Conception and Design: ML, CB, JH, CJ, RX
Writing: M.A.
Writing and Editing: MA, BJ
Supervision: ML
All authors read and approved the final manuscript.
References
1. Cruccu, Giorgio, Giulia Di Stefano, and Andrea Truini. “Trigeminal neuralgia.” New England Journal of Medicine 383.8 (2020): 754-762.
2. Zakrzewska, Joanna M. “Diagnosis and differential diagnosis of trigeminal neuralgia.” The Clinical journal of pain 18.1 (2002): 14-21.
3. Cruccu, Giorgio, et al. “Trigeminal neuralgia: new classification and diagnostic grading for practice and research.” Neurology 87.2 (2016): 220-228.
4. Bendtsen, Lars, et al. “Advances in diagnosis, classification, pathophysiology, and management of trigeminal neuralgia.” The Lancet Neurology 19.9 (2020): 784-796.
5. Di Stefano, G., Maarbjerg, S., Nurmikko, T., Truini, A. & Cruccu, G. Triggering trigeminal neuralgia. Cephalalgia 38, 1049–1056 (2018).
6. Lambru, Giorgio, Joanna Zakrzewska, and Manjit Matharu. “Trigeminal neuralgia: a practical guide.” Practical neurology 21.5 (2021): 392-402.
7. Headache classification committee of the International Headache Society (IHS) the International Classification of Headache Disorders, 3rd edition. Cephalalgia 38, 1–211 (2018).
8. Maarbjerg, S. & Benoliel, R. The changing face of trigeminal neuralgia-a narrative review. Headache 61, 817–837 (2021).
9. Araya, Erika I., et al. “Trigeminal neuralgia: basic and clinical aspects.” Current neuropharmacology 18.2 (2020): 109-119.
10. Goadsby, Peter J. “Trigeminal autonomic cephalalgias.” CONTINUUM: Lifelong Learning in Neurology 18.4 (2012): 883-895.
11. Obermann, Mark, and Zaza Katsarava. “Update on trigeminal neuralgia.” Expert review of neurotherapeutics 9.3 (2009): 323-329.
12. Maarbjerg, S., Gozalov, A., Olesen, J. & Bendtsen, L. Concomitant persistent pain in classical trigeminal neuralgia — evidence for different subtypes. Headache 54, 1173–1183 (2014)
13. Latorre, G., et al. “Diagnosis and treatment of trigeminal neuralgia: Consensus statement from the Spanish Society of Neurology’s Headache Study Group.” Neurología (English Edition) (2023).
14. Khawaja, Shehryar Nasir, and Steven J. Scrivani. “Trigeminal neuralgia.” Dental Clinics 67.1 (2023): 99-115.
15. Hung, Peter Shih-Ping, et al. “Sex differences in patient journeys to diagnosis, referral, and surgical treatment of trigeminal neuralgia: implications for equitable care.” Journal of Neurosurgery 139.2 (2022): 463-471.
16. Maarbjerg, S., Gozalov, A., Olesen, J. & Bendtsen, L. Trigeminal neuralgia — a prospective systematic study of clinical characteristics in 158 patients. Headache 54, 1574–1582 (2014).
17. Allam, Anthony K., et al. “Trigeminal neuralgia: diagnosis and treatment.” Neurologic Clinics 41.1 (2023): 107-121.
18. Andersen, A. S. S. et al. Microvascular decompression in trigeminal neuralgia — a prospective study of 115 patients. J. Headache Pain 23, 145 (2022).
19. Bendtsen, L. et al. European Academy of Neurology guideline on trigeminal neuralgia. Eur. J. Neurol. 26, 831–849 (2019).
20. Di Stefano, G., Maarbjerg, S. & Truini, A. Trigeminal neuralgia secondary to multiple sclerosis: from the clinical picture to the treatment options. J. Headache Pain 20, 20 (2019).
21. Bora, Nikita, et al. “A systematic review of the role of magnetic resonance imaging in the diagnosis and detection of neurovascular conflict in patients with trigeminal neuralgia.” Cureus 15.9 (2023).
22. Zeng, Chen, et al. “Recent advances of magnetic resonance neuroimaging in trigeminal neuralgia.” Current pain and headache reports 25 (2021): 1-7.
23. Obermann, M. et al. Impaired trigeminal nociceptive processing in patients with trigeminal neuralgia. Neurology 69, 835–841 (2007).
24. Jani, R. H. et al. Trigeminal nerve compression without trigeminal neuralgia: intraoperative vs imaging evidence. Neurosurgery 84, 60–65 (2019).
25. Sandell T, Eide PK. Effect of microvascular decompression in trigeminal neuralgia patients with or without constant pain. Neurosurgery. 2008; 63(1):93-99.
26. DeSouza DD, Hodaie M, Davis KD. Abnormal trigeminal nerve microstructure and brain white matter in idiopathic trigeminal neuralgia. Pain. 2014; 155(1):37-44.
27. Eller JL, Raslan AM, Burchiel KJ. Trigeminal neuralgia: definition and classification. Neurosurg Focus. 2005;18(5):E3.
28. Slettebø, Haldor. “Is this really trigeminal neuralgia? Diagnostic re-evaluation of patients referred for neurosurgery.” Scandinavian Journal of Pain 21.4 (2021): 788-793.
29. Kolakowski, Lukasz, et al. “Interdisciplinary strategies for diagnosis and treatment of trigeminal neuralgia.” Swiss Medical Weekly 154.7 (2024): 3460.
30. Zakrzewska JM, Wu J, Mon-Williams M, et al. Evaluating the impact of orofacial pain on quality of life using a cross-sectional survey in primary care. Pain. 2013;154(10):2240-2247.
31. Witkin, J. M., Pandey, K. P. & Smith, J. L. Clinical investigations of compounds targeting metabotropic glutamate receptors. Pharmacol. Biochem. Behav. 219, 173446 (2022).
32. Halbert-Elliott, Kyra M., et al. “Deep learning–based segmentation of the trigeminal nerve and surrounding vasculature in trigeminal neuralgia.” Journal of Neurosurgery 1.aop (2025): 1-9.
33. Doshi TL, Nixdorf DR, Campbell CM, Raja SN. Biomarkers in Temporomandibular Disorder and Trigeminal Neuralgia: A Conceptual Framework for Understanding Chronic Pain. Can J Pain. 2020;4(1): 1-18. https://doi:10.1080/24740527.2019.1709163
34. Wu C, Chen Y, Liu J, et al. Automated segmentation of trigeminal nerve using U-net in MR images. Comput Methods Programs Biomed. 2021;200:105854.
35. Zhang H, Bai Y, Zhu J, et al. Deep learning based lumbar spinal nerve root classification using MRI. J Magn Reson Imaging. 2020;52(3):874-883.
36. Hwang, Jun Ho, et al. “Deep Learning Super-Resolution Technique Based on Magnetic Resonance Imaging for Application of Image-Guided Diagnosis and Surgery of Trigeminal Neuralgia.” Life 14.3 (2024): 355.
37. Mienye ID, Swart TG, Obaido G, Jordan M, Ilono P. Deep Convolutional Neural Networks in Medical Image Analysis: A Review. Information. 2025; 16(3):195.
https://doi.org/10.3390/info16030195
38. Jamalabadi H, Kaur N, Navab N, et al. Deep learning for diagnosis and risk stratification of diabetic neuropathy using multimodal data. Sci Rep. 2022;12(1):7894.
39. Tran AT, Zeevi T, Payabvash S. Strategies to Improve the Robustness and Generalizability of Deep Learning Segmentation and Classification in Neuroimaging. BioMedInformatics. 2025; 5(2):20. https://doi.org/10.3390/biomedinformatics5020020
40. Bielewicz J, Kamieniak M, Szymoniuk M, Litak J, Czyżewski W, Kamieniak P. Diagnosis and Management of Neuropathic Pain in Spine Diseases. Journal of Clinical Medicine. 2023; 12(4):1380. https://doi.org/10.3390/jcm12041380
41. Xu, Dongyuan, et al. “Elucidating molecular lipid perturbations in trigeminal neuralgia using cerebrospinal fluid lipidomics.” Scientific Reports 15.1 (2025): 11777
42. Lafta R, van Munster C, Grootenboer H, et al. Proteomic analysis of trigeminal neuralgia patients reveals distinct molecular signatures and therapeutic implications. J Neuroinflammation. 2023;20(1):55.
43. Goebel A, Baranowski A, Maurer K, et al. Elevated serum cytokines in patients with complex regional pain syndrome. Pain. 2005;115(1-2):243-249.
44. Callaghan BC, Price RS, Feldman EL. Distal symmetric polyneuropathy: a review. JAMA. 2015; 314(20):2172-2181.
45. Shi X, Guo TZ, Li WW, et al. C5a complement and cytokine signaling mediate the pronociceptive effects of complex regional pain syndrome patient IgM in fracture mice. Pain. 2021;162(5):1400-1415. https://doi:10.1097/j.pain.0000000000002150
46. Wang, H., et al. “Chronic neuropathic pain is accompanied by global changes in gene expression and shares pathobiology with neurodegenerative diseases.” Neuroscience 114.3 (2002): 529-546.
47. Rueda‐Carrasco, Javier, et al. “SFRP1 modulates astrocyte‐to‐microglia crosstalk in acute and chronic neuroinflammation.” EMBO reports 22.11 (2021): e51696
48. Maiarù M, Tochiki KK, Cox MB, et al. The stress regulator FKBP51 drives chronic pain by modulating spinal glucocorticoid signaling. Sci Transl Med. 2016;8(325):325ra19. https://doi:10.1126/scitranslmed.aab3376
49. Siddarth, Prabha, et al. “Inflammatory markers of geriatric depression response to Tai Chi or health education adjunct interventions.” The American Journal of Geriatric Psychiatry 31.1 (2023): 22-32.
50. Guo Y, Li H, Hu L, et al. Proteomic profiling of cerebrospinal fluid in trigeminal neuralgia patients reveals axonal damage and neuroinflammation. J Neuroinflammation. 2022;19(1):38. https://doi:10.1186/s12974-022-02313-5
51. Chen ST, Yang JT, Weng HH, Wang HL, Yeh MY, Tsai YH. Diffusion tensor imaging for assessment of microstructural changes associate with treatment outcome at one-year after radiofrequency Rhizotomy in trigeminal neuralgia. BMC Neurol. 2019;19(1):62. Published 2019 Apr 12. https://doi:10.1186/s12883-019-1295-5
52. Uceyler N, Zeller D, Kahn AK, et al. Small fibre pathology in patients with fibromyalgia syndrome. Brain. 2013;136(Pt 6):1857-1867. https://doi:10.1093/brain/awt053
53. Miotto R, Wang F, Wang S, Jiang X, Dudley JT. Deep learning for healthcare: review, opportunities and challenges. Brief Bioinform. 2018;19(6):1236-1246. https://doi:10.1093/bib/bbx044
54. Ashina, S., Robertson, C.E., Srikiatkhachorn, A. et al. Trigeminal neuralgia. Nat Rev Dis Primers 10, 39 (2024). https://doi.org/10.1038/s41572-024-00523-z
55. Wu, X., Zhang, S., Zhang, Z. et al. Biologically interpretable multi-task deep learning pipeline predicts molecular alterations, grade, and prognosis in glioma patients. npj Precis. Onc. 8, 181 (2024). https://doi.org/10.1038/s41698-024-00670-2
56. Hasin Y, Seldin M, Lusis A. Multi-omics approaches to disease. Genome Biol. 2017;18(1):83. https://doi:10.1186/s13059-017-1215-1
57. Topol EJ. High-performance medicine: the convergence of human and artificial intelligence. Nat Med. 2019;25(1):44-56. https://doi:10.1038/s41591-018-0300-7
58. Guan H, Yap PT, Bozoki A, Liu M. Federated learning for medical image analysis: A survey. Pattern Recognit. 2024;151:110424. https://doi:10.1016/j.p atcog.2024.110424
59. Dayan, I., Roth, H.R., Zhong, A. et al. Federated learning for predicting clinical outcomes in patients with COVID-19. Nat Med 27, 1735–1743 (2021). https://doi.org/10.1038/s41591-021-01506-3
60. Hagedorn, Jonathan M., et al. “How well do current laboratory biomarkers inform clinical decision-making in chronic pain management?.” Journal of Pain Research (2021): 3695-3710.
61. Rieke N, Hancox J, Li W, et al. The future of digital health with federated learning. NPJ Digit Med. 2020;3:119. https://doi:10.1038/s41746-020-00323-1
62. Clark SJ, Arneson D, Darnell G, et al. Single-cell epigenomics: powerful new methods for understanding gene regulation and cell identity. Genome Biol. 2022;23(1):188. https://doi:10.1186/s13059-022-02742-y
63. Diaz MM, Caylor J, Strigo I, et al. Toward Composite Pain Biomarkers of Neuropathic Pain-Focus on Peripheral Neuropathic Pain. Front Pain Res (Lausanne). 2022;3:869215. Published 2022 May 11. https://doi:10.3389/fpain.2022.869215
64. National Institutes of Health. NIH Bridge2AI Program. https://commonfund.nih.gov/bridge2ai
65. Innovative Medicines Initiative. IMI-PainCare Project. https://www.imi-paincare.eu




