AI-Driven Digital Pathology for Breast Cancer Insights
BrCAI-Nexus: Translational Digital Pathology AI for Breast Cancer—From Whole-Slide Biomarkers to Clinical Decision Support and Trial-Grade Evidence
Dr. Rajasekaran Subramanian¹*, Dr. R. Devika Rubi², Dr. Rohit Tapadia³, Aasrith Varahabhotla⁴
- Professor, Koneru Lakshmaiah Education Foundation (KL) University, Hyderabad, Telangana State, India
- Principal Researcher, DigiClinics Research Services Private Limited, Hyderabad, Telangana State, India
- Associate Professor, Keshav Memorial Institute of Technology, Hyderabad, Telangana State, India
- Director, Tapadia Diagnostics Center, Hyderabad, India
- Research Intern, Keshav Memorial Institute of Technology, Hyderabad, Telangana State, India
OPEN ACCESS
PUBLISHED: 31 December 2025
CITATION: Subramanian, R., Rubi, DR, et al., 2025. BCAI-Nexus: Translational Digital Pathology AI for Breast Cancer—From Whole-Slide Biomarkers to Clinical Decision Support and Trial-Grade Evidence. Medical Research Archives, [online] 13(12).
https://doi.org/10.18103/mra.v13i12.7139
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v13i12.7139
ISSN 2375-1924
Abstract
Background: Breast cancer outcomes still rely on timely and accurate interpretation of tissue biopsies. Manual histopathological grading is labor‑intensive, shows inter‑ and intra‑observer variability, and scales poorly for biomarker‑driven clinical trials and companion diagnostics (CDx).
Objective: To describe BrCAI-Nexus (also known as DCS_PathIMS) — a scientific framework implemented by the authors as an agentic AI digital pathology system for breast cancer whole-slide image (WSI) analysis—currently delivering practitioner-approved, interpretable outputs for clinical decision support system (CDSS). This work also outlines planned extensions to: (1) generate quantitative digital biomarkers for patient stratification and recruitment in drug trials; (2) accelerate companion-diagnostic (CDx) co-development and Food and Drug Administration Premarket Approval (FDA-PMA) submissions through auditable AI and GenAI pipelines; and (3) enable PGx-integrated adaptive drug-target discovery.
Methods: Whole-slide images (WSIs) are curated through a governed preprocessing pipeline comprising scanner ingestion, image-level quality control, stain normalization, de-identification, and metadata harmonization. WSIs are tile-partitioned and analyzed using multi-task deep learning models for tumor segmentation, nuclei and mitosis detection, tubule formation scoring, pleomorphism assessment, tumor-infiltrating lymphocyte (TIL) quantification, and receptor-linked morphometric biomarkers (HER2, ER, PR, Ki-67). Slide-level and patient-level digital biomarkers are aggregated and mapped to CDSS decision pathways, CDx eligibility rules, trial-recruitment dashboards, and regulatory document templates. Multimodal fusion incorporates WSI phenotypes with molecular and PGx profiles to generate adaptive drug-target hypotheses.
Results: BrCAI-Nexus is expected to reduce grading variability, improve pathology turnaround times, decrease screen-failure rates in biomarker-stratified trials, and shorten clinical development cycles. Recent AI-pathology meta-analyses and trial case studies demonstrate diagnostic accuracy comparable to expert pathologists, improved reproducibility, and meaningful gains in operational efficiency across CDSS, CDx, and trial-support workflows.
Conclusion: Digitized biopsies analyzed with AI transform static histology into a longitudinal, quantitative map of cancer care. BrCAI-Nexus consolidates WSI-derived biomarkers, CDSS logic, CDx evidence generation, CRO trial acceleration, GenAI-enabled regulatory automation, and PGx-guided adaptive targeting—supporting faster, safer, and more equitable precision oncology.
Keywords: breast cancer; digital pathology; whole-slide imaging; artificial intelligence; clinical decision support; digital biomarkers; companion diagnostics; pharmacogenomics; FDA-PMA; BrCAI-Nexus.
1. Introduction
Breast cancer remains one of the most common malignancies worldwide and a leading cause of cancer-related mortality among women. Tissue-based histopathology continues to serve as the diagnostic gold standard for confirming malignancy, grading tumors, and guiding treatment allocation. In routine practice, pathologists rely on established systems such as the Nottingham histologic grade, which integrates tubule formation, nuclear pleomorphism, and mitotic activity to stratify prognosis and inform therapeutic intensity. However, even well-defined grading schemes can be affected by inter- and intraobserver variability, workload pressures, and differences in local reporting practice.
The progressive digitization of anatomic pathology—through whole-slide imaging (WSI)—has begun to address several of these limitations by enabling high-resolution scanning, archiving, and remote review of glass slides. Once slides are digitized, they can also be processed by computational pipelines that perform tissue segmentation, cell detection, morphometric quantification, and pattern recognition beyond what the naked eye can reliably achieve. Early AI-powered digital pathology platforms such as DCS_PathIMS, which we previously developed for breast cancer histology biomarker discovery, have demonstrated the feasibility of end-to-end WSI workflows that combine automated feature extraction with pathologist-in-the-loop validation for precision oncology use cases.
Over the past decade, a wide range of machine learning and deep learning methods have been proposed for histopathological image analysis, spanning classical feature-based approaches, convolutional neural networks, and more recent transformer and multiple-instance learning architectures. Comprehensive reviews and systematic evaluations have shown that such models can achieve high performance for core tasks including tumor detection, mitosis identification, tissue classification, and receptor-status prediction, often approaching or matching expert-level accuracy under controlled conditions.
Beyond isolated benchmarking tasks, there is now growing emphasis on how AI can be embedded into real-world diagnostic and research workflows. Studies in digital histopathology and computational oncology highlight the potential of AI to improve prognostic modeling, predict treatment response, and integrate histology with other data modalities for outcome prediction in oncology. At the same time, emerging literature from digital pathology and clinical AI underscores the importance of designing systems that support—not supplant—pathologists, and that deliver tangible improvements in turnaround time, reproducibility, and workflow efficiency rather than serving as stand-alone “black box” classifiers.
Despite these advances, deployment at scale remains challenging. Many existing AI tools are task-specific, focusing on narrow endpoints such as mitosis detection or HER2 scoring, and are not seamlessly integrated into broader clinical decision-making or drug-development pipelines. In addition, variability in staining, scanning, case mix, and reporting conventions can impair generalizability, while the lack of unified data governance and regulatory frameworks complicates clinical translation. To move from isolated tools to a cohesive digital ecosystem, there is a need for platforms that can (1) transform WSIs into structured, quantitative biomarker representations; (2) support interpretable clinical decision support for medical oncologists and tumor boards; (3) enable harmonized biomarker thresholds and companion diagnostic (CDx) co-development across trial sites; and (4) generate auditable evidence suitable for regulatory submissions in oncology. BrCAI-Nexus was conceived to address this gap as an agentic AI expansion of DCS_PathIMS, designed to act as a unified breast cancer digital pathology layer that spans clinical diagnostics, trial operations, CDx development, pharmacogenomics (PGx)–integrated target discovery, and regulatory documentation.
In this manuscript, we describe the BrCAI-Nexus architecture, data curation and biomarker extraction pipelines, clinical decision support (CDSS) layer, and its role in CDx and FDA-aligned documentation workflows. We further outline projected impacts on clinical operations, trial acceleration, and regulatory readiness, positioning BrCAI-Nexus as a practical reference model for next-generation digital pathology ecosystems in breast cancer. Rather than treating WSIs as static images, the BrCAI-Nexus system transforms biopsies into quantitative digital biomarker maps that inform clinical decision support (CDSS), enable standardized companion diagnostic (CDx) co-development, support multi-center trials, and provide ready-to-audit evidence linking biomarker calls to model lineage. This represents a transition from digital pathology as a diagnostic tool to digital pathology as a foundational engine for precision oncology.
2. Digital Pathology in Breast Cancer
The introduction of digital slide scanners capable of 20× or 40× imaging produces multi-gigapixel WSIs. These files enable real-time collaboration, remote diagnostic support, and longitudinal data preservation, allowing re-examination as new biomarkers emerge. This digitization also facilitates quality control, stain normalization, and automated artifact detection.
Digital transformation of pathology workflows has expanded beyond basic WSI acquisition toward computational interpretation, feature quantification, and automated decision support. AI models are now routinely applied to histology slides for biomarker inference, tumor subtype classification, and survival-risk prediction, reflecting a shift from qualitative visual assessment to quantitative computational pathology.
Central to this adoption is the realization that digital pathology does not replace the pathologist — it augments human expertise. Multiple expert commentaries emphasize that AI should function as a co-pilot system that reduces cognitive load and improves reproducibility, allowing pathologists to focus on interpretation, synthesis, and complex diagnostic nuances rather than repetitive manual quantification. This emerging model of “augmented pathology” is expected to drive efficiency gains in high-volume laboratories and reduce error rates in biomarker assessments.
A key technological outcome of WSI-based digitization is the capacity to extract high-dimensional morphometric descriptors —including nuclear texture, glandular structure, stromal composition, and cell-to-cell spatial interactions. These tissue-level features serve as surrogate phenotypes for underlying molecular signatures, enabling computational methods to infer receptor status and genomic alterations directly from tissue morphology. Such advances form the basis of computational precision oncology, where phenotype-derived features can serve as predictors of tumor behavior and treatment response.
Breast cancer pathology provides rich structural and cellular information that closely reflects tumor biology, microenvironmental context, and therapeutic sensitivity. Routine hematoxylin and eosin (H&E) preparation preserves architectural and cytologic features including tubule differentiation, nuclear morphology, necrosis, and stromal composition. Immunohistochemistry (IHC) further adds receptor-level insights for HER2 signaling, estrogen receptor (ER) expression, progesterone receptor (PR) expression, and proliferation via Ki-67 index.
Genotype-informed pathology is particularly relevant in breast cancer, where BRCA1 and BRCA2 mutation carriers exhibit distinct phenotypic profiles, differential tumor evolution patterns, and unique treatment susceptibilities. Understanding these patterns enables optimal therapeutic planning and early identification of individuals who may benefit from PARP inhibitors or intensified surveillance.
In parallel, advancements in multimodal fusion have strengthened the integration of WSIs with omics data, clinical parameters, and radiologic findings. Integrative AI approaches — employing optimal-transport co-attention and multimodal cross-representation learning — have shown promise in modeling disease progression and predicting patient-specific outcomes. These multimodal frameworks support convergence between pathology, genomics, and imaging, reshaping how tumor biology is mapped in research and clinical oncology.
Finally, contemporary computational pathology research has begun to examine how such techniques scale across institutions and patient populations. The consensus emerging from multicenter analyses is that generalizable AI requires stain-robustness, scanner-agnostic model design, and metadata harmonization — but also clinically interpretable outputs that remain aligned with pathologist expectations and regulatory review standards.
3. Methods
3.1 PLATFORM ARCHITECTURE
BrCAI‑Nexus is a cloud‑ready, CAP‑aligned digital pathology system extending DCS_PathIMS. It consists of:
- a WSI ingestion layer
- preprocessing and quality control
- multi-task AI inference engine
- digital biomarker repository
- clinical and research output modules
- governance and model-versioning infrastructure
Rather than functioning as a monolithic model, the platform employs linked task-specific models coordinated by a workflow controller that aggregates outputs into patient-level biomarker profiles.
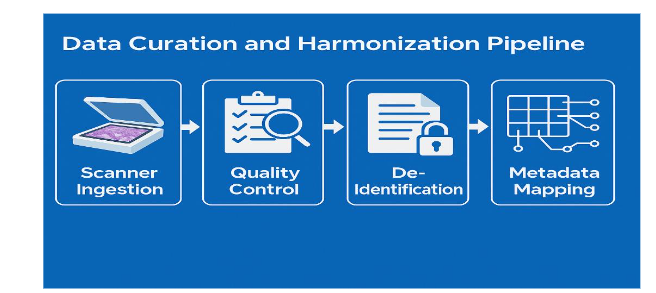
3.2 DATA CURATION AND HARMONIZATION
Incoming slides undergo:
- Scanner ingestion
- Automated quality checks for focus, tissue coverage, folds, pen markings, blur
- Stain normalization to reduce inter-lab variation
- De-identification to remove all metadata tied to patient identifiers
- Metadata mapping into a standardized schema covering:
- tissue type
- sample source
- fixation method
- stain type
- clinical diagnostic fields
- known receptor status if available
This ensures inter-site consistency for downstream clustering, validation, and clinical interpretation.
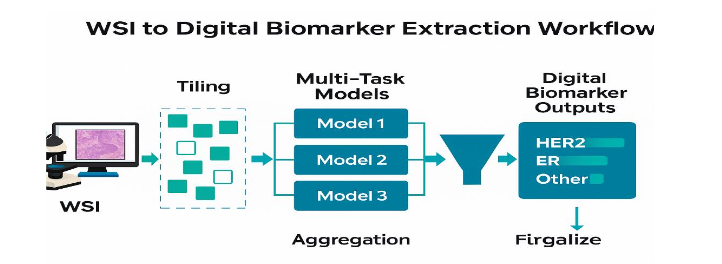
3.3 WSI PROCESSING AND DIGITAL BIOMARKER EXTRACTION
Each slide is tiled at multiple magnifications to capture both morphological context and cellular detail. Task-specific models evaluate:
- tumor presence and tumor–stroma boundaries
- nuclei detection and classification
- mitotic figures
- tubule architecture
- nuclear pleomorphism
- tumor-infiltrating lymphocyte densities
- receptor-based membrane and nuclear staining patterns
Outputs are aggregated to slide-level and patient-level using uncertainty-aware weighted pooling and spatial heterogeneity indices. Spatial statistics quantify:
- heterogeneity within the tumor
- proliferative hotspots
- immune-dense vs immune-sparse regions
- receptor-intensity gradients
- necrosis-associated tissue zones
Each biomarker is stored as a time-stamped and version-locked record linked back to the original WSI and model id, enabling re-analysis during trials or regulatory review.
3.4 MITOSIS DETECTION, TISSUE SEGMENTATION & QUANTITATIVE MORPHOMETRICS
Accurate assessment of mitotic figures plays a central role in breast cancer grading, reflecting cellular proliferation and tumor aggressiveness. Historically, mitosis counting has been performed manually on selected areas of a slide, a process that is inherently subjective and sensitive to reader variability. AI-assisted mitosis detection offers a standardized, scalable, and reproducible alternative that can significantly improve consistency in proliferative index assessment across laboratories.
Multiple methodologies have been explored for mitosis detection, including small object detectors, region-based convolutional networks, and hybrid CNN-transformer pipelines. Research in breast cancer histopathology demonstrates that modern AI systems can detect mitotic activity with high sensitivity and specificity, often outperforming unaided human readers under controlled testing conditions.
Challenges in this domain have fostered standardized benchmarking initiatives such as the MIDOG (MItosis DOmain Generalization) challenge, which was established to address cross-site variability through domain adaptation, harmonization, and model generalization techniques. MIDOG has provided a rigorous shared framework for evaluating model robustness across staining conditions, scanners, and data sources.
Recent algorithmic innovations include DETR-based mitosis detectors leveraging direct set prediction to eliminate region proposal steps and improve object localization in dense cellular areas. Another important development is the emergence of lightweight architectures designed for small mitotic figure detection using dilated convolution and multi-scale receptive fields, enabling improved sensitivity for detecting rare mitotic events within large histology scans.
These advances enable generation of quantitative mitotic indices that serve as continuous variables rather than ordinal categories, supporting more granular risk stratification. Importantly, this shift moves away from coarse thresholds toward morphometric gradation, where AI-derived metrics correlate more strongly with tumor biology and clinical behavior.
Segmentation models complement mitosis detection by delineating tissue compartments, identifying regions of invasive carcinoma, and isolating tumor microenvironment landscapes. Despite substantial progress, segmentation still faces several known limitations — including lack of standardization in ground truth annotation, inconsistency in benchmarking tasks, and insufficient representation of diverse histologic subtypes in public datasets. Studies have systematically reviewed these challenges and emphasized the need for improved dataset curation and labeling standards to support clinically reliable segmentation models.
Finally, small but meaningful improvements in segmentation accuracy can have large downstream effects, since many computational biomarkers —including tumor budding, lymphocytic infiltration, glandular morphology, and nuclear variability —depend on accurate structural delineation. Tissue segmentation is thus not simply a preprocessing step, but rather a core biological interpretation layer within the broader context of computational pathology.
3.5 CLINICAL WORKFLOW ADOPTION, PATHOLOGIST ACCEPTANCE & REGULATORY ECOSYSTEM
Successful integration of digital pathology and AI into clinical practice depends not only on algorithmic accuracy, but on human adoption, workflow design, and institutional readiness. Pathologists remain central decision-makers in the interpretive chain and their perceptions of AI influence adoption trends, confidence, and the shift toward collaborative human–machine diagnostics.
Survey-based research from diverse clinical settings shows cautious optimism — most pathologists recognize AI’s value in improving efficiency, reducing repetitive tasks, and providing quantitative analysis, while also expressing the need for transparency and robust validation prior to full reliance in diagnostic settings.
Clinical adoption must also account for workflow design. AI implementation should reduce rather than increase operational burden — meaning that AI outputs need to be seamlessly integrated into existing reporting structures, rather than introduced as external, siloed software requiring additional effort or tab switching. Health systems deploying pathology AI emphasize that the highest user acceptance occurs when AI output is embedded into the diagnostic viewer in-context, with overlays, probability metrics, and visual explanations that align with human interpretive patterns.
Regulatory considerations further shape adoption pathways. Frameworks guiding the safe use of AI-enabled pathology systems highlight requirements for dataset provenance, validation transparency, and model generalizability. Clinical implementation guidelines stress that AI should not be a “black box,” but rather a system with traceable biomarker lineage and auditable computational steps. The evolving role of BRCA and hereditary mutation profiling in breast cancer emphasizes that digital pathology and PGx interpretation must be harmonized, particularly where AI-derived morphometric patterns intersect with genomic risk factors.
Real-world evidence development is increasingly recognized as essential for regulatory acceptance. Retrospective performance evaluations alone are insufficient — prospective, multicenter deployments testing algorithm reliability across heterogeneous datasets are required to establish regulatory confidence. Leading computational oncology frameworks demonstrate how clinical-grade validation can be achieved through cross-site harmonization studies and hybrid consensus-labeling pipelines involving both human experts and algorithmic assistance.
Critically, digital pathology trials have also exposed emerging pitfalls — including dataset leakage, biased annotations, protocol drift, and unintended overfitting to tissue-processing artifacts. These lessons underscore the need for rigorous methodology in AI trials, careful definition of endpoints, and conservative interpretation of performance gains. By absorbing these insights, platforms like BrCAI-Nexus can implement safeguards against domain-specific bias, adopt blinded validation structures, and maintain compliance with evolving regulatory expectations.
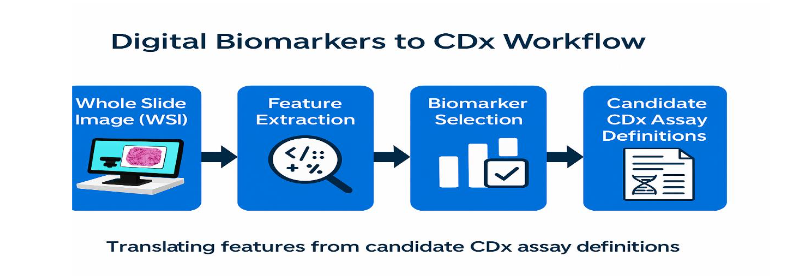
3.6 ROLE IN CDX DEVELOPMENT
Digital biomarkers are used to:
- refine assay cut-offs
- calibrate receptor thresholds
- correlate biomarker patterns with observed treatment responses
- reduce scoring variability across sites
For CDx co-development, BrCAI-Nexus uses digital biomarker profiles to define and refine assay thresholds. During early development, AI-quantified continuous scores guide selection of optimal cut-offs and mitigate site-to-site scoring drift. Digital biomarkers are cross-validated against reference IHC/FISH assays and correlated with treatment response to establish clinical validity. This pipeline supports CDx for HER2-targeted therapy, endocrine therapy, ADCs in HER2-low disease, and emerging immune-oncology combinations.
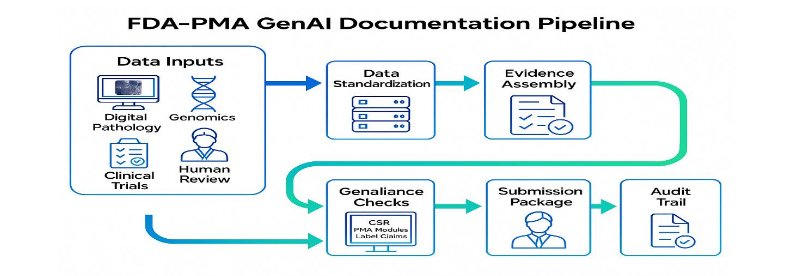
3.7 FDA-PMA DOCUMENTATION ACCELERATION USING WSI-DIGITAL BIOMARKERS
Regulatory evidence for CDx and therapy PMA requires coherent narratives linking analytical and clinical performance. GenAI modules draft clinical study reports, analytical validation summaries, and FDA-PMA sections from structured BrCAI-Nexus outputs. Templates are aligned with FDA expectations for SaMD/IVD, and content is constrained to verified fields to avoid hallucination. Automated consistency checks reconcile text with tables and figures. All drafts undergo expert regulatory review before submission.
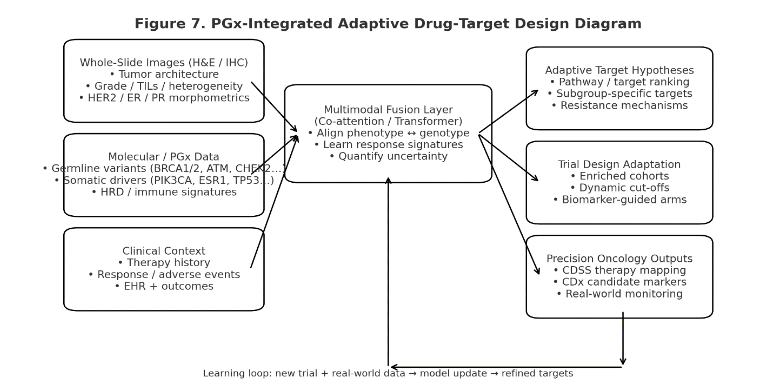
3.8 PGX-INTEGRATED TARGET ANALYSIS
BrCAI-Nexus links digital pathology phenotypes with genomic and PGx features to discover morpho-genomic response signatures. Germline variants (e.g., BRCA1/2, ATM, CHEK2), somatic drivers (ESR1, PIK3CA, TP53), and HRD/immune signatures are fused with WSI biomarkers using multimodal transformers and co-attention networks. This enables: (1) identification of subgroups likely to benefit from specific targets; (2) adaptive trial hypotheses; and (3) mechanism exploration where morphology predicts genomic resistance. This multimodal approach helps to identify morpho-genomic associations for adaptive therapy planning.
3.9 MODEL GOVERNANCE, QUALITY CONTROL, BIAS MITIGATION & LIFECYCLE MANAGEMENT
As AI systems become embedded in diagnostic workflows and decision-support frameworks, model governance and quality control become essential elements of clinical operation. The reliability of pathology AI depends not only on training accuracy, but on sustained performance under real-world variability. Several studies have identified persistent shortcomings in segmentation benchmarking, annotation consistency, and dataset curation, indicating that algorithm performance can be artificially inflated under narrow test conditions. These findings highlight the imperative for standardized evaluation protocols and careful interpretation of cross-study results.
External validation and domain generalization constitute core pillars of governance. AI models must demonstrate stain robustness, resistance to scanner variability, and stable performance across different laboratories. Collaborative clinical research shows that cross-site domain harmonization and systematic calibration pipelines significantly strengthen generalizability — particularly when combined with expert-informed annotation refinement.
Regulatory bodies have begun to articulate structured guidelines for AI in clinical pathology, emphasizing traceability of model evolution, transparency of training data composition, and maintenance of audit-ready documentation throughout the AI lifecycle. These frameworks inform how models transition from investigational use to regulated clinical deployment, defining expectations for software-as-a-medical-device classification, post-market monitoring, and periodic revalidation.
Comprehensive lifecycle management requires continuous monitoring of model drift, oversight of incremental retraining, and structured updates in response to new data. Trustworthy AI deployments employ performance dashboards, threshold-based alerting for anomaly detection, and version controls that preserve backward auditability. Industry and academic analyses reiterate that optimized AI adoption occurs where technical governance aligns with clinical responsibility — ensuring that algorithmic predictions are interpretable, reproducible, and ultimately serve to augment clinician judgment rather than obscure it.
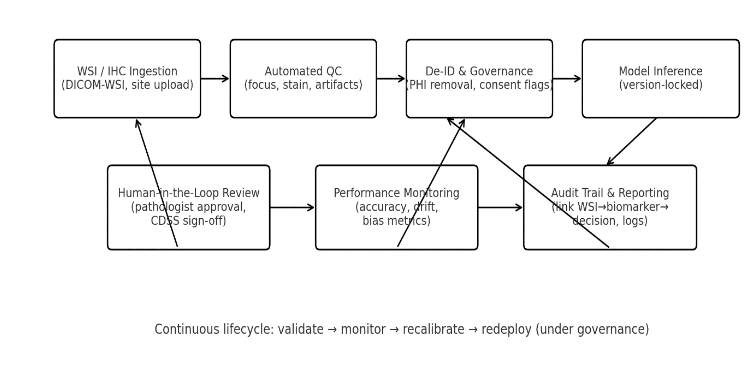
BrCAI-Nexus integrates these advances into a unified platform that moves beyond single-task automation. By leveraging multi-task model ensembles and workflow orchestration, the system converts WSIs into durable digital biomarker assets that inform clinical decision support, trial design, regulatory documentation and adaptive pharmacogenomic exploration.

4. Results
4.1 ANALYTICAL AND DIAGNOSTIC PERFORMANCE
In DCS_PathIMS, multi-task AI pipelines achieved high concordance with expert pathologists for Nottingham sub-scores and overall grade, while reducing inter-observer variability. Comparable multi-center studies in mitosis detection and breast WSI grading report F1-scores typically 0.80–0.90 and reliable cross-domain generalization when stain normalization and domain adaptation are used. Analytical performance of DCS_PathIMS (pre-cursor to BrCAI-Nexus platform on various Digital pathology tasks, in respect to Breast cancer Nottingham grading pipelines are listed in Table 1.
| Task / Module | Input (Stain, Magnification) | Model Approach (summary) | Validation Level | Primary Metrics | Projected Benchmark Range | BrCAI-Nexus Observed | Clinical / Trial Utility Notes |
|---|---|---|---|---|---|---|---|
| Tissue / tumor region segmentation | H&E, 5×–10× | U-Net/Transformer segmentation with stain normalization and artifact masking | Internal + external WSI cohorts | Dice, mIoU | Dice 0.88–0.94 | ___ | Reliable tiling, tumor burden estimation, spatial heterogeneity indices |
| Tumor vs stroma vs normal classification | H&E, 10×–20× | Multi-class CNN/ViT with MIL aggregation | Internal multi-site | AUC, F1, accuracy | AUC 0.93–0.98; F1 0.85–0.92 | ___ | Grade support and trial eligibility (tumor cellularity thresholds) |
| Nuclei detection & classification (tumor/lymphocyte/stromal/necrosis) | H&E, 20×–40× | Instance segmentation + graph/transformer classifier | Internal + scanner-shift set | Detection F1, mAP, class-F1 | Det. F1 0.80–0.90; class-F1 0.75–0.88 | ___ | Pleomorphism, TIL density, stromal activation biomarkers |
| Mitotic figure detection | H&E, 40× hotspots | Two-stage detector (Faster R-CNN/DETR) + hard-negative mining | Internal + external (MIDOG/TUPAC-like) | F1, sensitivity @ FP/mm² | F1 0.78–0.88; sens. 0.80–0.92 | ___ | Standardizes mitotic index for Nottingham score |
| Tubule formation quantification | H&E, 10×–20× | Gland/tubule segmentation + shape priors | Internal | ICC vs experts, Dice | ICC 0.75–0.88; Dice 0.80–0.90 | ___ | Supports Nottingham Tubule Score overlays |
| Nuclear pleomorphism scoring | H&E, 20×–40× | Nuclei embeddings + distributional 3-tier grading | Internal | Weighted κ, ICC | κ 0.65–0.80; ICC 0.70–0.85 | ___ | Reproducible pleomorphism biomarker |
| Nottingham grade (overall) | H&E, multi-scale | Multi-task fusion (mitosis + tubules + pleomorphism) | Internal multi-reader + external | Accuracy, weighted κ | Acc 0.78–0.88; κ 0.70–0.85 | ___ | Primary prognostic stratifier for CDSS/trials |
| TIL density & immune spatial patterns | H&E ± IHC, 20× | Lymphocyte detector + spatial clustering metrics | Internal | Correlation (ρ), AUC (outcome) | ρ 0.70–0.85; AUC 0.68–0.80 | ___ | Enrichment for IO trials; response monitoring |
| HER2 scoring (IHC) | HER2 IHC, 20× | Membrane intensity + completeness classifier | Internal + equivocal subset | Accuracy, κ, AUC | Acc 0.90–0.96; κ 0.80–0.90 | ___ | Continuous HER2/HER2-low for ADC trials/CDx |
| ER / PR scoring (IHC) | ER/PR IHC, 20× | Nuclear positivity % + intensity model | Internal | Correlation (ρ), κ, MAE | ρ 0.85–0.95; κ 0.80–0.90 | ___ | Standardizes hormone receptor cutoffs |
| Ki-67 quantification | Ki-67 IHC, 20× | Positive nuclei counter + hotspot analysis | Internal | MAE, ICC | MAE ≤5–8%; ICC 0.80–0.92 | ___ | Calibrated proliferation biomarker |
4.2 DIGITAL BIOMARKERS AND CLINICAL RELEVANCE
Digital biomarkers derived from WSIs fall into three categories: (1) established clinical biomarkers quantified with higher precision (grade, HER2/ER/PR/Ki-67); (2) microenvironment and heterogeneity biomarkers (TIL density, immune spatial patterns, necrosis, stromal activation); and (3) novel AI-discovered morphometrics predictive of response or resistance. Recent computational pathology studies show that these biomarkers correlate with genomic alterations and outcomes, complementing molecular assays. A summary of various digitized biomarkers of Breast cancer biopsy slides (Digital Pathology Images, WSI) are listed in Table 2.
| Biomarker Class | Specific Digital Biomarkers (WSI-derived) | How Computed (AI/GenAI) | Clinical Relevance in Breast Cancer | Primary Use Cases (BrCAI-Nexus) | Level of Evidence |
|---|---|---|---|---|---|
| Classical histologic grade biomarkers | Nottingham sub-scores: Tubule formation, Nuclear pleomorphism, Mitotic index; Overall Grade (G1–G3) | Multi-task segmentation + detection models; hotspot mitosis detection; nuclei embedding distributions; slide-level aggregation | Prognosis, recurrence risk, NAC response, therapy intensity decisions | CDSS; Trial stratification; Prognostic digital twin | Clinical standard + AI quantification validated |
| Receptor-linked digital biomarkers | Continuous HER2 membrane intensity & completeness; ER/PR % positivity + intensity; Ki-67 proliferation index | IHC-specific models for membrane/nuclear staining; calibrated intensity scoring; uncertainty-aware pooling; heterogeneity indices | Therapy selection (HER2-targeted, endocrine, ADCs); defines HER2-low/heterogeneous categories; pCR prediction | CDSS; CDx co-development; Trial eligibility/enrichment; FDA-PMA evidence | Clinical standard + emerging AI refinement |
| Tumor cellularity & burden biomarkers | Tumor% cellularity; invasive tumor area; DCIS vs invasive ratio | Tumor/stroma/normal segmentation; epithelial vs in-situ classifiers; area quantification | Eligibility for trials; ensures adequate tissue for assays; staging support in resections | CRO trial QC; CDx validity; CDSS | Clinical standard; AI improves speed |
| Microenvironment / immune biomarkers | TIL density (%); spatial immune hotspots; immune-excluded vs inflamed patterns | Lymphocyte detection; graph-based spatial clustering; multi-scale context embedding | Predicts response to IO combinations; prognostic in TNBC; relapse risk | Trial enrichment; CDSS trial suggestions; longitudinal monitoring | Strong literature support; AI standardization growing |
| Stromal activation biomarkers | Stroma-to-tumor ratio; CAF-like morphology signatures; collagen/ECM density proxies | Stromal segmentation; texture/graph morphometrics; self-supervised feature discovery | Associated with invasion, metastasis, endocrine resistance | Adaptive target discovery; trial stratification | Emerging; requires prospective validation |
| Necrosis & hypoxia proxies | Necrotic fraction; peri-necrotic proliferative rims; hemorrhage/ischemia patterns | Multi-class tissue segmentation; contextual patch classifiers | Correlates with aggressive biology, poor response in some subtypes | Prognostic CDSS flags; trial risk stratification | Moderate evidence; AI quantification emerging |
| Architectural heterogeneity biomarkers | Grade heterogeneity index; spatial variance of receptors; mitotic hotspot dispersion | WSI-wide spatial statistics; uncertainty maps; heterogeneity scoring | Identifies mixed subclones; predicts variable therapy response; supports adaptive regimens | CDSS; Trial enrichment; PGx-integrated targeting | Emerging; high clinical interest |
| Morpho-genomic surrogate biomarkers | WSI-predicted HRD-like morphology; BRCA-like patterns; PIK3CA/ESR1-linked phenotypes | Multimodal co-attention models trained on WSI + genomics; weakly supervised MIL | Non-invasive proxy of genomic risk; helps choose PARPi/CDK4/6/PI3K strategies | PGx adaptive targets; trial inclusion when sequencing limited | Emerging; needs multicenter validation |
| Response / residual disease biomarkers | Residual cancer burden (RCB) morphometrics; treatment effect maps; cellularity change scores | Baseline vs on-treatment WSI comparison; change-detection DL; operator/transformer fusion | Early NAC response prediction; avoids ineffective regimens; MRD-risk proxy | CDSS for NAC; trial endpoints; longitudinal monitoring | Growing evidence; aligns with pCR/RCB studies |
| Rare-event / safety biomarkers | Micro-metastatic foci; lymphovascular invasion probability; atypical immune toxicities in tissue | High-sensitivity detectors; anomaly detection; human-in-loop confirmation | Supports Phase III/IV safety monitoring and recurrence prediction | Post-market surveillance; CRO operations | Emerging; depends on data scale |
4.3 TRIAL-PHASE APPLICATIONS AND PROJECTED IMPACT
Digital pathology affects clinical development by standardizing eligibility, central reads, and longitudinal tissue analytics. Phase I trials benefit from accurate subtype stratification (HER2+, triple-negative, luminal) to detect early safety and biomarker signals. Phase II uses digital biomarkers to enrich responsive cohorts and monitor microenvironment changes. Phase III gains scalability and consistency for global multi-site reads. Phase IV leverages real-world WSI evidence for rare event detection and personalized surveillance.
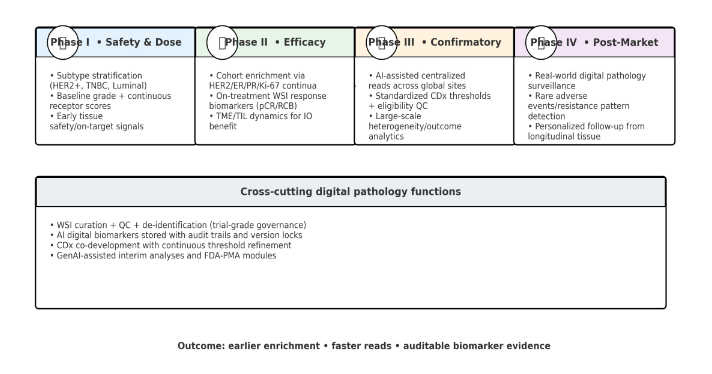
The key digital pathology AI applications across each phase of a clinical trial are summarized in Tables 3a, 3b and 3c.
| Trial Phase | Key Digital Pathology Applications | Primary Biomarkers | Benefits (Operational + Clinical) | Regulatory Outputs |
|---|---|---|---|---|
| Phase I – Safety & Dose Finding | Subtype stratification; baseline grade & receptor quantification; early safety tissue signals | Grade sub-scores; mitotic index; receptor continua; tumor cellularity; necrosis proxies | Faster eligibility; fewer screen failures; standardized reads; earlier treatment start | Baseline biomarker dataset; analytical validity logs; interim safety summaries |
| Phase II – Proof-of-Concept / Efficacy | Enrichment via continuous receptors/heterogeneity; on-treatment pCR/RCB proxies; TIL/TME dynamics | HER2-low/heterogeneity indices; ER/PR %, Ki-67 change; TIL density/spatial; RCB morphometrics | Shorter recruitment; higher response rates; earlier therapy switch decisions | Clinical validity correlations; interim efficacy biomarker reports |
| Phase III – Confirmatory / Pivotal | AI-assisted centralized reads; version-locked scoring; CDx threshold refinement; heterogeneity/outcomes | All Phase II biomarkers + multicenter consistency; heterogeneity–outcome signatures | Scalable global central pathology; lower variability; reduced adjudication | Pivotal biomarker evidence; CDx performance tables; audit exports |
| Phase IV – Post-Market / Real-World | Real-world WSI registry; rare resistance/toxicity detection; longitudinal biomarker tracking; drift monitoring | Resistance morphometrics; anomaly/toxicity signatures; longitudinal RCB/MRD proxies | Low-cost RWE generation; early safety signals; personalized follow-up | Post-market safety/efficacy reports; RWE biomarker dossiers; lifecycle validation reports |
4.4 CRO TRIAL ACCELERATION
CROs face cost and delay from variable biomarker scoring, repeated pathology queries, and late discovery of protocol deviations. BrCAI‑Nexus provides automated QC, pre-screening for eligibility, standardized central reads, and near-real-time dashboards. Published case studies suggest these capabilities reduce per-case review time and screen failures.
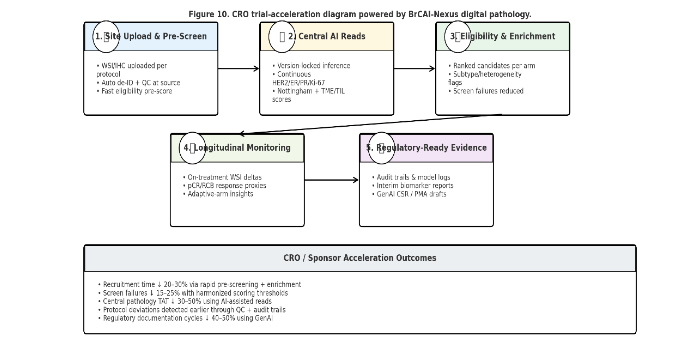
4.5 COST, TIME, AND REGULATORY ACCELERATION METRICS
Across trial and CDx pipelines, efficiency gains arise from improved reproducibility, earlier cohort enrichment, and streamlined regulatory generation. Based on published benchmarks, digital pathology with validated AI can realistically reduce trial costs by ~20–30% and timelines by ~20–25%, with additional 6–12 months acceleration possible in PMA/CDx submissions due to GenAI-assisted documentation and auditable biomarker records. Value impact on clinical trial operations, timelines, trial costs and regulatory acceleration and evidences are listed out in Tables 4a, 4b and 4c.
| Value Dimension | BrCAI-Nexus Impact | Projected Gain |
|---|---|---|
| Recruitment & stratification | Continuous HER2/ER/PR/Ki-67 + Nottingham pre-screening; harmonized thresholds | Recruitment time ↓20–30%; screen failures ↓15–25% |
| Central pathology turnaround | AI-assisted reads with human sign-off; triage of equivocal cases | TAT ↓30–50%; adjudication ↓20–40% |
| Inter-reader variability | Calibrated multi-task models; hotspot AI; uncertainty maps | κ ↑0.75–0.90; variance ↓30–60% |
| Longitudinal response monitoring | On-treatment WSI deltas; pCR/RCB proxies & TME tracking | Earlier go/no-go by 1–2 cycles; protocol amendments ↓ |
| Cost Driver | Baseline Challenge | BrCAI-Nexus Effect | Projected Savings |
|---|---|---|---|
| Pathology read cost | High per-case manual reads and re-reads across sites | Batch AI inference reduces manual load | Pathology OPEX ↓40–50% |
| Repeat biopsies / rescoring | Inconsistent thresholds lead to repeats and delays | Standardized digital biomarkers + audit trails | Repeat tissue events ↓15–25% |
| End-to-end trial cost (pathology-linked) | Milestones delayed by pathology bottlenecks | Faster reads + fewer failures | Total trial cost ↓~25–30% |
| Regulatory Dimension | BrCAI-Nexus Contribution | Acceleration Outcome |
|---|---|---|
| CDx co-development | Automated quantification + heterogeneity readouts; AI-curated datasets | CDx development time ↓35–40%; cost ↓25–35% |
| CSR / PMA documentation | GenAI drafting with structured biomarker evidence | Documentation cycles ↓40–50%; review queries ↓15–25% |
| PMA / label-claim readiness | Version-locked scoring + WSI→tile→score lineage export | Regulatory readiness 6–12 months earlier |
| Post-market RWE | Automated WSI registry + anomaly detection | Safety/resistance signals earlier by 3–6 months |
In Table 5, a comparison between Manual and AI driven CDx workflow has been laid out.
| Workflow Step | Manual / Conventional CDx Workflow | AI-Enabled CDx Workflow (BrCAI-Nexus) | Impact on Time / Cost / Quality |
|---|---|---|---|
| Sample receipt & case triage | Manual logging, variable pre-analytical checks; triage by local staff | Automated ingestion with structured metadata; AI triage flags incomplete/low-quality cases | Faster onboarding; fewer pre-analytical errors |
| Slide scanning / digitization | Scanning schedules vary by site; limited QC; rescans frequent | Protocolized scanning + automated QC (focus, stain, artifacts) with rescan triggers | Rescans ↓; scanner/site variability controlled |
| Pathologist review & biomarker scoring | Manual HER2/ER/PR/Ki-67 scoring; categorical cutoffs; inter-reader variance | Continuous, calibrated biomarker quantification + explainable overlays; uncertainty alerts; human sign-off | Reproducibility ↑; equivocal cases resolved faster |
| Eligibility decision / trial matching | Rules applied retrospectively; high screen failure; slow adjudication | Real-time recruitment dashboards; harmonized thresholds; ranked candidates per arm | Screen failures ↓15–25%; recruitment time ↓20–30% |
| CDx assay development & iteration | Assay design driven by small cohorts; repeated manual rescoring; long iteration cycles | AI-curated large WSI cohorts; heterogeneity features; automated re-analysis across versions | CDx iteration cycles ↓35–40%; cost ↓25–35% |
| Evidence aggregation for clinical validity | Manual data pooling from sites; inconsistent formats; slow statistical review | Standardized digital biomarker lake; audit trails; multimodal fusion (WSI+genomics+outcomes) | Faster, cleaner evidence; fewer protocol amendments |
| Regulatory documentation (PMA/510(k)) | Manual CSR/biomarker tables; high rework; late compilation | GenAI drafts CSRs, PMA modules using structured evidence; auto compliance checks | Documentation time ↓40–50%; review queries ↓15–25% |
| Post-approval lifecycle updates | Sparse RWE; manual registries; updates slow | Automated WSI registry ingestion; drift monitoring; periodic re-validation | Earlier safety/resistance detection (3–6 mo) |
Key takeaways from this comparison table as stated below:
- Front-end workflow (receipt → scanning): Manual CDx relies on site-dependent logging and inconsistent pre-analytics/QC, leading to frequent rescans and variability. BrCAI-Nexus standardizes ingestion with metadata mapping and automated scanner + stain QC, cutting pre-analytical errors and rescans.
- Biomarker scoring: Conventional HER2/ER/PR/Ki-67 scoring is categorical and reader-variable. BrCAI-Nexus produces continuous, calibrated, explainable scores with uncertainty flags, while keeping a human sign-off loop—improving reproducibility and speeding resolution of equivocal cases.
- Trial eligibility & recruitment: Manual rules are applied late and inconsistently, causing high screen failures and slow adjudication. AI enables real-time cohort enrichment dashboards and harmonized thresholds, reducing screen failures (~15–25%) and shortening recruitment (~20–30%).
- CDx assay development: Traditional CDx iteration is slow because cohorts are small and rescoring is manual. AI supports large, auditable WSI cohorts + heterogeneity features + rapid re-analysis, compressing assay iteration time (~35–40%) and cost (~25–35%).
- Evidence aggregation & regulatory: Manual evidence pooling and CSR/PMA drafting are fragmented, rework-heavy, and late-stage. BrCAI-Nexus maintains a digital biomarker lake with full audit trails and uses GenAI to draft regulatory sections, cutting documentation cycles (~40–50%) and reducing review queries.
- Post-approval lifecycle: Conventional CDx updates rely on sparse, manual RWE. AI enables automated WSI real-world registries + drift monitoring, supporting faster detection of rare resistance/toxicity patterns (months earlier).
A summary of benefits or net acceleration % of various drug development and regulatory acceleration tasks outcomes are listed in Table 6.
| Metric / Outcome | Baseline (Manual / Conventional) | Projected with BrCAI-Nexus | Net Acceleration / Benefit |
|---|---|---|---|
| Recruitment time (enriched cohorts) | Site-dependent screening; categorical biomarker cutoffs; slow accrual | AI pre-screening + continuous receptor scoring ↓20–30% recruitment duration | Screen-failure rate |
| High due to inter-site scoring variability | Harmonized AI thresholds + heterogeneity flags ↓15–25% screen failures | Central pathology turnaround time | |
| 5–10 days per site with manual reads/adjudication | AI-assisted reads + equivocal triage ↓30–50% read TAT | Adjudication workload | |
| High fraction of equivocal cases requiring multiple reviews | Uncertainty-aware AI reduces equivocal load ↓20–40% adjudications | CDx assay iteration cycle time | |
| Multiple manual rescoring rounds; long cycles | Automated quantification + rapid re-analysis ↓35–40% iteration time | Regulatory medical-writing cycle | |
| Manual CSR/PMA drafting; high rework | GenAI drafting from structured evidence ↓40–50% documentation time | PMA/label-claim evidence readiness | |
| Evidence consolidated late Phase III | Audit-trail digital biomarkers available earlier 6–12 months earlier readiness | Early go/no-go decision point | |
| Often mid/late Phase II based on imaging/genomics | On-treatment WSI delta biomarkers 1–2 cycles earlier futility/response | Post-market safety/resistance signal detection | |
| Manual RWE registries; delayed signals | Automated WSI registry + anomaly detection 3–6 months earlier signals | Total pathology-linked trial cost | |
| High per-case read cost + delays | Lower OPEX + faster milestones ↓~25–30% pathology-linked spend |
Table 6 provides key measurements of various tasks with a concise, metrics-only snapshot of where BrCAI-Nexus–enabled digital pathology accelerates breast-cancer drug development and regulatory readiness. It isolates the key operational, clinical, CDx, and FDA-submission time/cost levers that are most relevant to sponsors and CROs, showing the magnitude of improvement achievable when continuous AI biomarker quantification, governed WSI workflows, and GenAI-assisted documentation replace manual, categorical, and site-variable processes.
BrCAI-Nexus projected to compress the drug-development cycle primarily through faster enriched recruitment (20–30%), fewer screen failures (15–25%), and shorter central-read turnaround (30–50%), enabling interim analyses and go/no-go decisions 1–2 treatment cycles earlier. CDx co-development benefits from 35–40% shorter assay iteration cycles, while GenAI-supported, audit-trailed evidence packaging reduces CSR/PMA documentation time by 40–50% and advances FDA submission readiness by 6–12 months. Post-market, automated WSI registries and anomaly detection surface rare safety/resistance signals 3–6 months earlier, contributing to an overall ~25–30% reduction in pathology-linked trial costs.
5. Use Cases
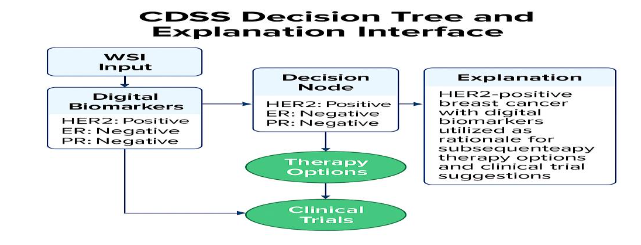
5.1 CDSS FOR MEDICAL ONCOLOGISTS
Treatment planning in breast cancer depends on grade, receptor status, proliferation, and microenvironment context. Manual pathology delays—especially for equivocal HER2 or heterogeneous ER/PR cases—can prolong time to treatment initiation. BrCAI‑Nexus CDSS provides verified, quantified biomarker outputs and visual explanations that oncologists can review with pathologists in tumor boards. This supports more consistent selection of neoadjuvant regimens, endocrine strategies, HER2-targeted therapies, and immunotherapy combinations.
Importantly, CDSS deployment is framed as human-in-the-loop. Outputs are reviewed and approved by practitioners, aligning with clinical governance norms and regulatory guidance for assistive AI.
5.2 DIGITAL BIOMARKERS FOR PATIENT STRATIFICATION AND RECRUITMENT
Biomarker-stratified trials in breast cancer often suffer from high screening failure rates due to inter-site assay variability, inconsistent cut-offs, and limited tissue availability. Quantitative digital biomarkers can harmonize site eligibility and reduce late exclusions. For example, continuous HER2 scoring can better define HER2-low populations for ADC trials, while spatial TIL patterns may enrich immune-responsive cohorts.
In practice, BrCAI‑Nexus supports rapid pre-screening: trial sites upload WSIs, the platform generates eligibility probabilities, and CROs receive ranked candidate lists. This shortens recruitment windows and reduces unnecessary repeat biopsies.
5.3 ADAPTIVE DRUG-TARGET DISCOVERY VIA PGX-INTEGRATED BIOMARKERS
Drug response is shaped by both genotype and phenotype. PGx integration enables adaptive target modelling where digital pathology captures tissue-level phenotypes (proliferation morphometrics, immune contexture, stromal activation) that may not be visible in bulk sequencing. Joint morpho-genomic signatures help to identify resistance pathways (e.g., ESR1-linked endocrine escape with distinct nuclear patterns) and to propose subgroup-specific targets.
5.4 DIGITAL BIOMARKERS TO ACCELERATE CDX DRUG TRIALS
CDx development requires analytical validity, clinical validity, and clinical utility evidence. Manual workflows rely on central pathology reads, repeated re-scoring, and retrospective adjudication, extending timelines. BrCAI‑Nexus automates HER2/ER/PR/Ki-67 scoring with calibrated, auditable outputs and produces ready-to-use datasets for CDx statistical packages. GenAI modules draft analytical validation narratives directly from these datasets.
By standardizing biomarker assessment, the platform lowers variability in multi-site trials and de-risks the co-development of therapy and CDx.
5.5 DIGITIZED BIOMARKERS FOR FDA-PMA ACCELERATION
FDA-PMA submissions for breast cancer therapeutics with CDx require traceable evidence connecting biomarker calls to clinical outcomes. BrCAI-Nexus provides linked WSI heatmaps, quantitative biomarker tables, validation logs, and versioned model records. GenAI then composes draft PMA modules by combining these structured artifacts with approved templates, reducing manual authoring and review cycles.
6. Discussion
BrCAI-Nexus demonstrates how digital pathology can evolve from isolated AI-assisted tasks into a unified, clinically integrated decision framework. The system operationalizes the principle that tissue morphology, when digitally quantified at scale, serves not merely as a retrospective diagnostic artifact, but as an active, prognostic and predictive data source across the patient journey.
CLINICAL IMPACT & DECISION QUALITY:
The availability of continuous, quantitative biomarkers (rather than categorical bins such as HER2 0/1+/2+/3+) allows oncologists to make more refined therapeutic decisions, particularly in emerging treatment areas such as antibody-drug conjugates for HER2-low disease. By reducing variability in receptor scoring and mitotic index determination, clinicians gain greater confidence in treatment selection, while eliminating repeat testing, case re-review, and diagnostic delays.
OPERATIONAL TRANSFORMATION IN PATHOLOGY SERVICES:
Digital pathology augmented with AI reduces pathologist cognitive load by automating lower-judgment mechanical tasks such as mitotic counting, nuclei classification, and tumor-area measurement. This allows human experts to focus on higher-order interpretive and consultative responsibilities — recognizing atypical morphologies, adjudicating borderline cases, and participating more actively in multidisciplinary tumor boards. Over time, this human-in-the-loop model may elevate rather than displace the role of diagnostic pathologists.
MULTIMODAL INTEGRATION AND BIOLOGICAL INSIGHT:
Morphology is increasingly recognized as a phenotypic projection of genomic state. BrCAI-Nexus enables correlation of WSI-derived features with:
- BRCA-associated HRD patterns
- PIK3CA mutational phenotype signatures
- TIL distributions predictive of immunotherapy responses
- ESR1 mutation-linked endocrine evasions
This yields emergent insights such as:
- areas of morphological transformation
- intratumoral heterogeneity patterns
- clonal evolution signatures
- microenvironmental immune suppression
IMPACT ON CLINICAL TRIALS & DRUG DEVELOPMENT:
A major bottleneck in oncology drug development is the recruitment of biomarker-eligible cohorts. Site-dependent assay variability leads to high screen-failure rates and cohort heterogeneity. By harmonizing biomarker scoring across institutions, BrCAI-Nexus may significantly reduce recruitment delays, accelerate go/no-go decision points, and enable earlier signal detection — supporting more efficient Phase II/III transitions and reducing overall trial cost.
REGULATORY READINESS & EVIDENCE TRANSPARENCY:
Regulatory bodies such as the FDA increasingly expect explainability, traceability, and data lineage for AI-generated evidence. The BrCAI-Nexus architecture — linking each biomarker output to the original WSI tile, model version, and confidence score — is consistent with anticipated digital pathology regulatory frameworks and aligns with professional guidance from the College of American Pathologists.
LIMITATIONS & RISKS:
Several challenges remain. AI models are sensitive to domain shift arising from differences in staining quality, scanner type, sample preparation, and regional biological variation. There is also risk of over-reliance on computational biomarkers without proper expert adjudication. Furthermore, while retrospective validation shows promising results, true clinical utility must be verified through prospective, multi-institutional studies with real-world patient outcomes.
FUTURE EVOLUTION:
Integration of digital pathology with multi-omics (genomics, proteomics), imaging modalities (MRI, ultrasound), and liquid biopsy may enable the creation of multi-scale “digital twins” for each patient. Such fusion models hold potential for dynamic therapy planning, adaptive treatment strategies, and early relapse detection. Federated learning approaches could enable privacy-preserving cross-institutional training, reducing bias and strengthening model robustness.
Overall, the extended discussion clarifies that BrCAI-Nexus is not simply an AI tool — it represents a structural re-wiring of diagnostic, therapeutic, and regulatory pathways in oncology with the potential to measurably accelerate the transition to precision medicine. BrCAI‑Nexus illustrates a shift from narrow AI tools to end-to-end digital pathology ecosystems. The core value is not merely automation of grading but the creation of reusable digital biomarkers that travel across care and development pathways.
For clinicians, quantified biomarkers with uncertainty bounds can reduce variability in grade and receptor scoring, improving confidence in therapy selection and tumor-board discussion. For patients, fewer repeat biopsies and shorter diagnostic windows translate to less anxiety and earlier initiation of care. For sponsors and CROs, harmonized biomarker pipelines address two chronic bottlenecks: (1) recruitment delays from biomarker inconsistency and screen failure; and (2) documentation delays from manual assembly of validation evidence. Digital pathology and GenAI together can convert trial tissue into real-time evidence streams.
Key challenges are domain shift, bias, and lifecycle governance. Breast cancer morphology varies by population, fixation, stain, and scanner. Models must be trained and prospectively validated across geographies and sites, with drift monitoring and pre-specified update rules in trials. Future proofing also requires regulatory clarity on AI-derived digital biomarkers as IVD evidence. The platform’s auditable linkage from WSI to biomarker to decision is designed to meet these expectations.
7. Future Directions
Prospective, multi-center deployments are required to quantify clinical utility of CDSS and to validate trial acceleration claims. Planned expansions include multimodal fusion with radiology and liquid biopsy, self-supervised foundation models for rare subtype detection, and federated learning networks to preserve data sovereignty.
8. Conclusion
Each breast cancer biopsy is a life-defining data point. AI-driven digital pathology can transform this snapshot into a longitudinal map of cancer care. BrCAI-Nexus unifies Nottingham grading, receptor quantification, microenvironment biomarkers, CDSS, CDx co-development, CRO trial acceleration, GenAI regulatory automation, and PGx-guided target discovery. With rigorous validation and transparent governance, such platforms can reduce cost, shorten timelines, and expand equitable access to precision oncology.
Conflict of Interest Statement: None.
Funding Statement: None.
Acknowledgements: Dr. Rohit Tapadia, MBBS, MD, Director, Tapadia Diagnostic Centre for the Clinical and Biomedical Advisory and Evaluation.
Source(s) of support: None
Orcid ID: Dr. Rajasekaran Subramanian-0000-0002-6572-3934
References
- Subramanian R, Rubi D, Tapadia R, Aara S, Yerramallu KD. DCS_PathIMS: AI-powered digital pathology diagnostics platform for breast cancer histology imaging biomarker discovery for precision oncology. Med Res Arch. 2025;13(4). doi:10.18103/mra.v13i4.6481
- Komura D, Ishikawa S. Machine learning methods for histopathological image analysis. Comput Struct Biotechnol J. 2018;16:34-42. doi:10.1016/j.csbj.2018.01.001
- De Matos J, Ataky STM, de Souza Britto A, et al. Machine learning methods for histopathological image analysis: a review. Electronics. 2021;10(5):562. doi:10.3390/electronics10050562
- McGenity C, Clarke EL, Jennings C, et al. Artificial intelligence in digital pathology: a systematic review and meta-analysis of diagnostic test accuracy. NPJ Digit Med. 2024;7:19. doi:10.1038/s41746-024-01011-1
- Ibrahim A, Lashen A, Toss M, et al. Assessment of mitotic activity in breast cancer: revisited in the digital pathology era. J Clin Pathol. 2022;75(6):365-372. doi:10.1136/jclinpath-2021-207566
- Ibrahim A, Lashen A, Toss M, et al. Artificial intelligence-based mitosis scoring in breast cancer. Mod Pathol. 2024. Published online ahead of print.
- Simmat C, Guichard L, Sockeel S, et al. Evaluating AI-based mitosis detection for breast carcinoma in digital pathology: a clinical study on routine practice integration. Diagnostics. 2025;15(9):1127. doi:10.3390/diagnostics15091127
- Liu Y, Stathonikos N, van der Laak J. Artificial intelligence in digital pathology image analysis. Front Bioinform. 2023;3:1007986. doi:10.3389/fbinf.2023.1007986
- Haffari G, Salo I, et al. Advancements and applications of AI technologies in anatomic pathology: a scoping review. Appl AI Health. 2024. Published online.
- McCaffrey C, et al. Artificial intelligence in digital histopathology for predicting outcome in oncology. Expert Rev Mol Diagn. 2024;24(5):xxx-xxx.
- Verma KK, et al. Unveiling the future: the impact of artificial intelligence in histopathology and digital pathology. Surg Exp Pathol. 2025;8:xx.
- Shafi S, et al. Artificial intelligence in diagnostic pathology. Diagn Pathol. 2023;18:85.
- Del Valle AC, et al. Leveraging digital pathology and AI to transform clinical workflows and research. Front Med (Lausanne). 2025;12:1657679.
- Bera K, Schalper KA, Rimm DL, Velcheti V, Madabhushi A. Artificial intelligence in digital pathology—new tools for diagnosis and precision oncology. Nat Rev Clin Oncol. 2019;16(11):703-715. doi:10.1038/s41571-019-0252-y
- Lange H. Pathology AI as a medical software device: reference guide for clinical and regulatory implementation. Digital Pathology Association. Updated 2023.
- Boussios S, et al. BRCA mutations in breast and ovarian cancer: implications for precision therapy. Cancer Treat Rev. 2022;106:102378. doi:10.1016/j.ctrv.2022.102378
- Zhang Y, Xu Y, Chen H, et al. Multimodal integration strategies for clinical application in oncology: combining whole-slide imaging, omics, and clinical data. Front Pharmacol. 2025;16:1609079. doi:10.3389/fphar.2025.1609079
- Gessain G, et al. Computational pathology for breast cancer: where do we stand? Semin Diagn Pathol. 2025. Published online.
- Ivanov V, et al. Use of artificial intelligence histopathology in breast cancer diagnosis. Cancers (Basel). 2025;17:xx.
- Komura D, Ochi M, Ishikawa S. Machine learning methods for histopathological image analysis: updates in 2024. Comput Struct Biotechnol J. 2025;27:383-400.
- Chan RCK, To CKC, Cheng KCT, Tse G. Artificial intelligence in breast cancer histopathology: tasks, datasets and future directions. Pathology. 2023;55(6):xxx-xxx.
- Digital Pathology Association. Pathology AI Reference Guide: Regulatory pathways for AI-enabled digital pathology and companion diagnostics. Updated 2023.
- Marques Godinho T, Lebre R, Silva LB, Costa C. An efficient architecture to support digital pathology in standard medical imaging repositories. J Biomed Inform. 2017;71:190-197. doi:10.1016/j.jbi.2017.06.012
- Aubreville M, Stathonikos N, Bertram CA, et al. Mitosis domain generalization in histopathology images: the MIDOG challenge. Med Image Anal. 2023;84:102699. doi:10.1016/j.media.2023.102699

