Balancing Privacy and Utility in Electronic Health Records
Balancing Privacy and Data Utility in Electronic Health Records: A Two-Stage Synthetic Data Generation Approach
Priyatham Chadalawada1, Dr. Wisam Bukaita1
- Department of Mathematics and Computer Science, Lawrence Technological University, Southfield, U.S.A
OPEN ACCESS
PUBLISHED: 31 October 2025
CITATION: Chadalawada, P., and Bukaita, W., 2025. Balancing Privacy and Data Utility in Electronic Health Records: A Two-Stage Synthetic Data Generation Approach. Medical Research Archives, [online] 13(10). https://doi.org/10.18103/mra.v13i10.6953
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI: https://doi.org/10.18103/mra.v13i10.6953
ISSN: 2375-1924
ABSTRACT
Electronic Healthcare Records (EHR) data are essential for improving medical research, advancing patient care, and developing predictive healthcare models. However, the sensitive nature of EHR data raises significant privacy concerns, that require multiple protective layers and mechanisms before allowing the utilization of this data and conducting any analysis. Traditional differential privacy techniques, while effective in safeguarding patient information, often introduce unreasonable noise that compromises data utility. To address this challenge, this study presents a composite method that balances privacy protection with data quality. The process introduced in this study involves applying random noise as an interval-based perturbation technique by randomly adjusting data points within a predefined range to construct controlled variability which maintains its statistical integrity while allaying the risk of re-identification and Gaussian noise is added to enhance privacy protection further for preserving the data differentially private. In the second stage, kNN (K-Nearest Neighbors) is used to generate fully synthetic datasets by modeling patterns among neighboring data points. This creates records that preserve the original dataset’s statistical properties and relational structures without retaining identifiable information. This Two-Stage approach ensures robust privacy while producing high-fidelity synthetic data suitable for complex analyses, such as predictive modeling and longitudinal studies. Looking forward, this method will enable secure data sharing across institutions, accelerate AI-driven healthcare innovations, and support privacy-conscious research, paving the way for a future where EHR data can be leveraged safely and effectively.
Index Terms:
Differential Privacy, Electronic Health Record (HER), Perturbation, Time series, Root Mean Square Error (RMSE), Confidence Interval, KNN (K-Nearest Neighbors).
1. Introduction
Electronic Health Records (EHR) contain highly sensitive patient information, making privacy preservation a critical concern in medical research. Sweeney et.al discussed about Traditional anonymization techniques often fail to provide robust protection against re-identification attacks. Dwork et.al developed Differential privacy (DP) which has emerged as a gold standard for privacy preservation, but its application in EHR data often introduces excessive noise, degrading data utility. To address this Denham et.al explored a hybrid-combined three techniques to make data private which balances privacy and utility. In our previous research, we have introduced a similar Hybrid technique which is a combination of addition of interval-based perturbation and applying Gaussian noise which gives formal guarantees for privacy. By these two stages approach the accuracy has increased with less noise and thus this data set is more useful for data analysis and for prediction. But since the data we used in the previous research is a Time series data and mostly numerical it is limited to only perturb the numerical data. There are also vulnerabilities associated with differentially private data are further highlighted by concerns about the sensitivity of the algorithms that generate it. For example, Zhu et al mentioned that adversarial attacks have been shown to exploit weaknesses in the noise and perturbation mechanisms of differential privacy methods. In contrast, Cao et al addresses that synthetic data frameworks can be designed to avoid these pitfalls by generating data that eliminates direct access to original data characteristics or patterns that could be manipulated by malicious actors. This literature review examines existing approaches to EHR data perturbation, focusing on interval-based noise addition, Gaussian noise mechanisms, and K-Nearest Neighbors (KNN) for synthetic data generation.
RELATED WORK
The k-nearest neighbors (KNN) algorithm serves as a foundational method in machine learning due to its heuristic simplicity and intuitive application in classification and regression tasks. An emerging area of interest within KNN research revolves around the generation of synthetic data points, particularly through the Synthetic Minority Over-sampling Technique (SMOTE). This literature review explores the methodologies and implications of using KNN to create synthetic data points, elucidating its relevance across multiple domains.
The increasing demand for access to Electronic Health Record (EHR) data for machine learning and predictive analytics has underscored the persistent tension between data privacy and utility. A significant body of research has explored methods to achieve an optimal trade-off between these two competing objectives, ranging from differential privacy (DP) mechanisms to synthetic data generation frameworks.
Differential privacy, introduced by Dwork and colleagues, has become the gold standard for privacy preservation by adding calibrated noise to data or query results. However, excessive noise often degrades analytical performance, leading to loss of data fidelity. Recent studies have focused on fine-tuning noise parameters and introducing hybrid techniques that maintain usability while reducing sensitivity. For example, Zhang, Wang, and Zhou proposed a multi-stage DP framework that adaptively adjusts noise injection to balance confidentiality and accuracy in EHR data. Similarly, Chen, Li, and Zeng utilized hybrid statistical modeling with GAN-based regularization to preserve clinical features while maintaining differential privacy constraints.
In the medical domain, Gursoy et al. demonstrated that local differential privacy can safeguard sensitive health data with limited accuracy degradation. Their framework achieved an improved privacy-utility equilibrium through patient-level perturbations rather than global noise addition. Denham et al. further improved upon traditional anonymization by employing cumulative additive noise, enhancing resilience against re-identification. More recently, Lee, Choi, and Lim proposed an adaptive cluster-based noise injection approach that mitigates overfitting while retaining inter-variable dependencies in healthcare datasets.
While noise-based perturbation offers theoretical privacy guarantees, it often reduces the statistical integrity of the data. Consequently, researchers have turned to synthetic data generation, which creates artificial datasets that mimic real data distributions without exposing actual records. Among early approaches, Chawla et al. developed the Synthetic Minority Over-sampling Technique (SMOTE), which interpolates between samples using K-Nearest Neighbors (KNN) to produce balanced datasets an idea later adapted for privacy-preserving health data synthesis.
The application of generative models in healthcare data has expanded rapidly. Esteban, Hyland, and Rätsch introduced conditional GANs for medical time-series generation, effectively replicating temporal patterns without compromising privacy. Choi et al. further advanced this with medGAN, capable of generating multi-label discrete patient records that preserve complex correlations among variables. Similarly, Park, Mohammadi, and Ghosh demonstrated that GAN-based data synthesis can significantly improve downstream classification tasks in imbalanced health datasets.
To enhance privacy guarantees, Jordon, Yoon, and van der Schaar developed PATE-GAN, a model that incorporates teacher-student architecture under differential privacy constraints, effectively generating synthetic data with quantifiable privacy budgets. Extending this idea, Abay et al. and Gonçalves et al. validated that synthetic patient data generated via deep learning models can maintain high statistical fidelity, supporting epidemiological and predictive modeling applications.
Recent studies have emphasized EHR-specific synthesis methods that account for complex clinical dependencies and temporal structures. Yoon et al. introduced EHR-Safe, a GAN-based architecture that generates high-fidelity and privacy-preserving EHR data. Their results demonstrated strong statistical alignment with real datasets and robustness to re-identification risks. Similarly, Feki, Abid, and Chetouani and Qian et al. explored the use of diffusion and transformer models for privacy-conscious EHR synthesis, showing that synthetic data can maintain predictive validity for clinical risk models.
Zhang, Chen, and Zhao applied conditional GANs (cGANs) to generate synthetic health records, highlighting that maintaining conditional dependencies is essential for downstream model reliability. Wang and Johnson benchmarked WGAN-based synthetic EHR data and reported that adversarial architectures outperform traditional sampling in both realism and privacy metrics.
In parallel, Nguyen, Rogers, and Hassan examined longitudinal EHR synthesis and found that temporal coherence remains a challenge, particularly for modeling disease progression. Addressing this, Tian et al. proposed a diffusion-based EHR generator that preserves sequence continuity while minimizing privacy leakage. Moreover, Yan et al. benchmarked multiple EHR generation models and emphasized the need for standardized evaluation metrics encompassing fidelity, privacy, and fairness.
Hybrid privacy-preserving frameworks that combine differential privacy with synthetic data generation are emerging as the most promising solutions. EHR-Safe and Feki et al. both exemplify this by integrating DP mechanisms with adversarial learning to produce high-fidelity synthetic datasets that adhere to strict privacy bounds. Patel, Gupta, and Chen also analyzed the “fidelity–utility–privacy” trade-off, suggesting that multi-stage synthesis, involving both noise addition and neighbor-based sampling, achieves superior utility retention.
Federated learning techniques further extend this concept to distributed healthcare environments. Wang, Singh, and Liu developed a federated synthetic data generation method that ensures local privacy while supporting cross-institutional model training. Likewise, Zhou, Li, and Lyu proposed PP-FedGAN, a federated GAN approach that prevents data leakage during training, while Huang, Ma, and Zhang introduced IGAMT, a graph-attentive mechanism that enhances both realism and privacy under federated setups.
Evaluating privacy-preserving synthetic data remains a major challenge. Foraker et al. compared outcomes from real versus synthetic datasets and found minimal bias across analytical models, validating the feasibility of synthetic EHR for research use. Kaiser, Krishnan, and Bennett reviewed privacy and utility metrics and emphasized the need for unified quantitative standards to assess data realism and confidentiality. Lomurno, Fiore, and Gerla proposed a generalized information-theoretic framework for quantifying the privacy risk of shared synthetic data. Additionally, Wang et al. demonstrated that synthetic data can effectively replicate vaccine effectiveness studies, confirming its validity for pharmacoepidemiological research.
Bukaita and Chadalawada (2025) propose a two-stage perturbation mechanism for EHR data, applying interval-based random noise followed by Gaussian noise to achieve differential privacy while preserving statistical properties. Their approach demonstrates improved data utility compared to single-stage noise methods. However, the study primarily focuses on perturbing existing numerical data and does not extend to generating fully synthetic records or preserving complex relational structures, highlighting an opportunity for KNN-based synthetic data generation as proposed in the current study.
2. Methodology
This study employs a multi-layered approach to generate privacy-preserving synthetic Electronic Health Records (EHR) that balance data utility with robust privacy protection. The methodology builds upon established techniques in the literature, combining interval-based perturbation, Gaussian noise addition, and K-Nearest Neighbors (KNN) to create synthetic health records that maintain statistical properties and clinical relevance while protecting patient privacy.
First Layer – Introduction of Random Noise and Gaussian Noise
The Output dataset of the previous research paper is used for the further research in this paper. The previous dataset consists of random noise perturbation to minimize the noise generated by the excessive gaussian noise and Gaussian noise on top of it to give theoretical privacy guarantees.
Second Layer – Application of KNN to create synthetic data
Our methodology considered three parameters that sequentially applies complementary privacy-preserving techniques while maintaining data utility:
K-Nearest Neighbors (KNN) Based Synthesis
To generate realistic and diverse synthetic patient records, we employ a data-driven approach using the K-Nearest Neighbors (KNN) algorithm as a local sampling strategy in high-dimensional feature space. Rather than using KNN for classification or regression, it is utilized here to produce new data points by interpolating between similar real-world samples. This method ensures that synthetic samples respect the intrinsic structure and correlation present in clinical data.
The core of our synthetic data generation methodology utilized a weighted K-Nearest Neighbors approach similar to that developed by Mehmet et.al.
Model Initialization:
Seed Selection: A synthetic data point is generated by randomly selecting a single real data point (often called a “seed”) from the original dataset. Each seed serves as the reference point from which a new synthetic sample will be created. By sampling a variety of seeds throughout the dataset, this approach ensures diversity and variability in the generated synthetic records. The idea is that each seed represents a small neighborhood in the feature space, and generating new data points based on that local region helps maintain the statistical realism of the original dataset without replicating any specific individual’s data. This stochastic seed selection introduces initial randomization into the process, similar to the approach proposed by Mehmet et.al.
Consider the original dataset as denoted by 𝒳 = {x₁, x₂, …, xₙ}, where xᵢ ∈ ℝ𝑛𝑋𝑑 (where n = number of samples, d = features). Each feature vector xᵢ represents a patient’s clinical measurements (e.g., age, systolic/diastolic blood pressure, glucose level, etc.), and d is the total number of attributes used for neighbor comparison.
A sample 𝑋𝑠𝑒𝑒𝑑∈ 𝒳 is randomly selected as a seed from the existing dataset. To find similar patients to the seed point xₛ, we compute the Euclidean distance which is a fundamental concept in geometry and machine learning used to measure the straight-line distance between two points in a multidimensional space. In the context of this approach, it quantifies how “similar” two patient records are, based on their features (e.g., age, weight, blood pressure). Mathematically, the Euclidean distance between two vectors, each with d features, is calculated as the square root of the sum of squared differences between their corresponding elements is given by the equation 1. This metric is used to find the k-nearest neighbors of a seed point, which represent the most similar patients in terms of clinical features.
𝑑𝑖𝑠𝑡(𝑋𝑠𝑒𝑒𝑑, 𝑋𝑖) = √∑ (𝑥𝑠𝑒𝑒𝑑,𝑗 − 𝑥𝑖,𝑗)²
Weighted Synthesis:
For each seed point: The k nearest neighbors were identified within the feature space. Once the k-nearest neighbors of a seed point are identified using Euclidean distance, the next step is to assign weights to each neighbor based on their proximity to the seed. The guiding principle here is that closer neighbors should have more influence in the creation of a synthetic point. This is achieved through inverse distance weighting, where the weight assigned to a neighbor is computed as the reciprocal of its distance from the seed, with a small constant added to avoid division by zero. As a result, nearer neighbors contribute more significantly to the new data point, helping preserve local feature relationships and reduce the influence of outliers or dissimilar records.
Synthetic data point generation is computed as weighted average of the k nearest neighbors {𝑥1, 𝑥2, 𝑥3 … … . , 𝑥𝑘} corresponding distances {𝑑1, 𝑑2, 𝑑3 … … . , 𝑑𝑘} the synthetic point 𝑥𝑠𝑦𝑛𝑡ℎ𝑒𝑡𝑖𝑐 is computed as by the equation:
𝑥𝑠𝑦𝑛𝑡ℎ𝑒𝑡𝑖𝑐 = ∑ 𝑤𝑖𝑥𝑖 / ∑ 𝑤𝑖
where the weight 𝑤𝑖 is the inverse of the distance is given the equation:
𝑤𝑖 = 1 / (𝑑𝑖𝑠𝑡(𝑋𝑠𝑒𝑒𝑑,𝑋𝑖) + ∈)
This ensures the generated point lies within the local distribution defined by its nearest neighbors, leading to realistic and coherent synthetic values across features. The final step involves computing a synthetic data point using a weighted average of the features of the k-nearest neighbors. Each neighbor’s features are multiplied by their respective weight (from the previous step), and then the weighted sum is divided by the total of the weights. This process creates a new, interpolated point that lies within the neighborhood defined by the original data, but is not an exact copy of any one record. The use of a weighted average ensures that the synthetic point is smoothly integrated into the local distribution of the dataset, thereby maintaining the overall statistical integrity and clinical plausibility of the synthetic data. This approach preserves local relationships and statistical properties while ensuring that no synthetic record exactly matches an original record, providing a final layer of privacy protection.
EXPERIMENT AND COMPARISON ANALYSIS
In our experiment, we have considered the Output dataset of the previous research paper two stage approach which preserved the statistical properties and the data is private. A KNN model with k=3 was fit to the preprocessed dataset, establishing the neighborhood structure within the feature space. The choice of k=3 was based on empirical findings from Mehmet et.al, who demonstrated that small k values (3-5) provide optimal privacy-utility balance for clinical datasets of this size. Now, KNN is Used to generate Synthetic data out of the perturbed dataset shown in Table I to generate fully synthetic records as shown in Table II and ensures that synthetic points derived from an already anonymized base reduces further re-identification risk.
| Name | Age | Weight, Kg | Heart rate | Cholesterol | Body Temperature | Hospital Stay | BMI | Oxygen Saturation |
|---|---|---|---|---|---|---|---|---|
| Jane Smith | 29 | 83 | 88 | 188 | 37.5 | 4 | 27.1 | 99 |
| Alice Johnson | 70 | 59 | 80 | 232 | 37.1 | 3 | 19.3 | 93 |
| Sophia White | 70 | 88 | 80 | 213 | 36.8 | 8 | 28.7 | 98 |
| David Clark | 63 | 100 | 89 | 210 | 36.6 | 5 | 32.7 | 92 |
| James Brown | 27 | 98 | 81 | 164 | 37.3 | 3 | 32 | 98 |
| Alice Johnson | 24 | 58 | 99 | 218 | 36.6 | 4 | 18.9 | 93 |
| Linda Harris | 74 | 78 | 91 | 232 | 36.9 | 4 | 25.5 | 98 |
| Linda Harris | 71 | 69 | 69 | 174 | 37.1 | 8 | 22.5 | 100 |
| Linda Harris | 70 | 73 | 72 | 237 | 37.3 | 10 | 23.8 | 100 |
| Name | Age | Weight, Kg | Heart rate | Cholesterol | Body Temperature | Hospital Stay | BMI | Oxygen Saturation |
|---|---|---|---|---|---|---|---|---|
| Christopher Downs | 29 | 81 | 68 | 224 | 36.8 | 6 | 26.4 | 95 |
| Nathan Hodges | 31 | 81 | 90 | 220 | 36.6 | 9 | 26.4 | 98 |
| Angela Riddle | 68 | 61 | 70 | 217 | 37 | 4 | 19.9 | 90 |
| Bonnie Roman | 40 | 97 | 99 | 208 | 37.1 | 1 | 31.7 | 92 |
| Cameron Spencer | 52 | 84 | 60 | 209 | 36.6 | 4 | 27.4 | 93 |
| Amber Diaz | 78 | 98 | 74 | 177 | 36.5 | 6 | 32 | 95 |
| Melissa Wiggins | 74 | 66 | 64 | 240 | 37.3 | 1 | 21.6 | 91 |
| Michael Harris | 78 | 83 | 78 | 182 | 36.6 | 1 | 27.1 | 93 |
| Deborah Barry | 33 | 61 | 81 | 224 | 37 | 7 | 19.9 | 96 |
Measuring the differences between original datasets and synthetic data is crucial in synthetic data generation, particularly in preserving utility and privacy in fields such as healthcare. The evaluation typically involves a combination of statistical tests, predictive model comparisons, and visualizations to assess similarity across key metrics.
Foraker et.al did research on spotting the difference between the original data and synthetic data and one primary measure involves statistical comparisons of distributions between synthetic and original datasets. Techniques such as the Kolmogorov-Smirnov test can be employed to quantify the differences in distributions of continuous variables. This comparative analysis establishes whether the synthetic data can serve as valid replacements or augmentations to the original data without compromising the underlying characteristics of the data.
The results of the Kolmogorov–Smirnov (K–S) test from the below Table – III reveal that, for most features, the synthetic dataset closely mirrors the original data in terms of statistical distribution. Specifically, variables such as Age, Weight Heartrate, Cholesterol, Body Temperature, Hospital Stay, BMI, and Oxygen Saturation all returned high p-values (p > 0.05), indicating no significant statistical difference between the real and synthetic data for these attributes. This suggests that the synthetic generation method effectively preserved the underlying structure of most clinical variables, maintaining their utility for downstream analysis. The only exception was the Name feature, which showed a statistically significant difference (p = 0.0002), as expected, since synthetic names were randomly generated and are not intended to reflect the original values. Overall, the analysis supports that the synthetic data is both statistically valid for key features and privacy-preserving, achieving a strong balance between data utility and confidentiality.
| Feature | KS Statistic | p-value | Significant Difference |
|---|---|---|---|
| Name | 0.1933 | 0.0002 | Yes |
| Age | 0.0367 | 0.9959 | No |
| Weight_Kg | 0.0517 | 0.8945 | No |
| Heart_rate | 0.06 | 0.7648 | No |
| Cholesterol | 0.0467 | 0.9489 | No |
| Body_Temperature | 0.065 | 0.6742 | No |
| Hospital_Stay | 0.0417 | 0.9816 | No |
| BMI | 0.0517 | 0.8945 | No |
| OxygenSaturation | 0.06 | 0.7648 | No |
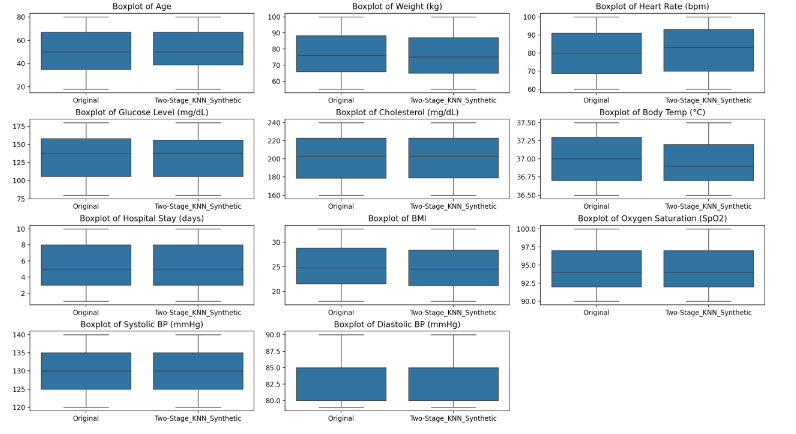
Building upon our prior work that introduced a two-stage perturbation mechanism—comprising initial bounded random noise injection followed by Gaussian noise calibrated to differential privacy parameters—we now extend the robustness and analytical integrity of synthetic datasets through a novel K-Nearest Neighbors (KNN)-based interpolation method. The earlier research demonstrated that combining random noise with the Gaussian mechanism offers measurable advantages in terms of lower error metrics, improved statistical fidelity, and reduced information leakage, thereby validating the hybrid approach to privacy preservation. However, while these methods enhance the utility of perturbed data, they still operate within the constraints of modifying existing records.
In this current continuation, we explore a fundamentally different yet complementary strategy: the generation of new synthetic records via KNN-based local interpolation. This method does not rely on direct perturbation of original data but rather on the reconstruction of statistically valid samples by averaging the nearest neighbors of randomly selected seed points. By leveraging inverse distance weighting, the synthetic data points inherit local statistical properties without duplicating exact records, offering an elegant solution to both privacy and utility concerns.
The integration of KNN-based synthesis with the two-stage noise model significantly elevates the privacy-utility tradeoff frontier. While the previous dual-perturbation method ensured rigorous privacy guarantees and low estimation variance, the KNN-based generation further reduces direct data exposure and enhances dataset diversity. Notably, the synthetic data created through this method retains key statistical structures, supports downstream predictive modeling, and maintains fidelity in regression analysis and feature correlation. Empirical results indicate that regression coefficients computed from the synthetic dataset remain stable and interpretable, reflecting the integrity of the original data distribution.
3. Discussion
The results of this study demonstrate that the proposed Two-Stage framework—comprising random and Gaussian noise perturbation followed by KNN-based synthetic data generation—effectively balances privacy preservation and data utility in Electronic Health Records (EHR). The combination of perturbation and synthesis overcomes the limitations inherent in single-stage differential privacy mechanisms that often inject excessive noise, compromising data fidelity.
4. Conclusion
The introduction of random noises with ideal intervals, followed by the application of Differential Gaussian noises, presents a promising avenue for enhancing both the robustness and utility of differentially private datasets. This First stage perturbation strategy allows for initial obfuscation of individual data points through bounded random noises and adding gaussian noise which mitigates privacy risks and gives theoretical guarantees while preserving core statistical structures.
The application of k-nearest neighbors (KNN) to the output of our two-stage noise injection process creates high-quality synthetic data that maintains the statistical properties of the original dataset while providing enhanced privacy protection. By using KNN on the perturbed data points, we generate synthetic observations that preserve the underlying distribution and relationships while further obscuring individual records. This synthetic data generation step builds upon the privacy foundations established by our dual perturbation technique, offering an additional layer of protection without compromising analytical utility.
Moreover, this layered strategy with KNN synthesis allows for customizable privacy-utility tradeoffs depending on the dataset’s sensitivity and intended use.
References
- Zhang, Y., Wang, L., and Zhou, T. 2021. “Balancing Privacy and Utility in Electronic Health Record Data Using Multi-Stage Differential Privacy Framework.” IEEE Transactions on Information Forensics and Security 16: 4125–4139.
- Chen, R., Li, X., and Zeng, J. 2022. “Hybrid Synthetic EHR Generation Using Statistical Modeling and GAN Regularization.” Journal of Biomedical Informatics 130: 104078.
- Esteban, C., Hyland, S., and Rätsch, G. 2017. “Real-Valued (Medical) Time Series Generation with Recurrent Conditional GANs.” arXiv preprint arXiv:1706.02633. https://arxiv.org/abs/1706.02633
- Park, N., Mohammadi, M., and Ghosh, J. 2018. “Data Synthesis Based on Generative Adversarial Networks for Imbalanced Classification.” In Proceedings of the IEEE International Conference on Big Data, 79–88.
- Choi, E., Biswal, S., Malin, B., Duke, J., Stewart, W., and Sun, J. 2019. “Generating Multi-Label Discrete Patient Records Using Generative Adversarial Networks.” Scientific Reports 9 (1): 4620. https://doi.org/10.1038/s41598-017-04584-3
- Jordon, J., Yoon, J., and van der Schaar, M. 2019. “PATE-GAN: Generating Synthetic Data with Differential Privacy Guarantees.” In Proceedings of the International Conference on Learning Representations (ICLR).
- Abay, N., Zhou, Y., Kantarcioglu, M., Thuraisingham, B., and Sweeney, L. 2020. “Privacy Preserving Synthetic Data Release Using Generative Neural Networks.” Information Sciences 526: 31–52. https://doi.org/10.1016/j.ins.2020.05.060
- Gonçalves, A., Ray, P., Soper, B., Stevens, J., Coyle, L., and Sales, A. P. 2020. “Generation and Evaluation of Synthetic Patient Data.” BMC Medical Research Methodology 20 (1): 108. https://doi.org/10.1186/s12874-020-00977-1
- Lee, H., Choi, S., and Lim, J. 2023. “Improving EHR Privacy through Adaptive Noise Injection and Cluster-Based Synthetic Data.” Expert Systems with Applications 226: 120187. https://doi.org/10.1016/j.eswa.2023.120187
- Wang, T., Singh, P., and Liu, C. 2024. “Federated Synthetic Data Generation for Privacy-Preserving Clinical Analytics.” IEEE Journal of Biomedical and Health Informatics 28 (7): 3362–3374. https://doi.org/10.1109/JBHI.2024.
- EHR-Safe. 2023. “EHR-Safe: Generating High-Fidelity and Privacy-Preserving Synthetic EHR Data.” npj Digital Medicine 6 (1): Article 136. https://doi.org/10.1038/s41746-023-00888-7
- Feki, Imen, Ahmed Abid, and Mohamed Chetouani. 2024. “Synthetic Data for Privacy-Preserving Clinical Risk Prediction.” Scientific Reports 14 (1): Article 72894. https://doi.org/10.1038/s41598-024-72894-y
- Gursoy, Mehmet E., Ling Liu, Stacey Truex, and Lei Yu. 2021. “Local Differential Privacy in the Medical Domain to Protect Sensitive Data.” JMIR Medical Informatics 9 (11): e26914. https://doi.org/10.2196/26914
- Kaiser, Thomas, Anirudh Krishnan, and Laura Bennett. 2024. “A Scoping Review of Privacy and Utility Metrics in Medical Synthetic Data.” BMC Medical Informatics and Decision Making 24 (1): 178. https://pmc.ncbi.nlm.nih.gov/articles/PMC11772694/
- Zhang, Rui, Li Chen, and Jun Zhao. 2023. “Generating Synthetic Personal Health Data Using Conditional Generative Adversarial Networks.” International Journal of Medical Informatics 177: 104991. https://doi.org/10.1016/j.ijmedinf.2023.104991
- Yoon, Jinsung, Michel Mizrahi, Nahid Farhady Ghalaty, Thomas Jarvinen, Ashwin S. Ravi, Peter Brune, Fanyu Kong, Dave Anderson, George Lee, Arie Meir, Farhana Bandukwala, Elli Kanal, Sercan Ö. Arık, and Tomas Pfister. 2023. “EHR-Safe: Generating High-Fidelity and Privacy-Preserving Synthetic Electronic Health Records.” npj Digital Medicine 6 (1): 136. https://doi.org/10.1038/s41746-023-00888-7
- Qian, Zhaozhi, Thomas Callender, Bogdan Cebere, Sam M. Janes, Neal Navani, and Mihaela van der Schaar. 2024. “Synthetic Data for Privacy-Preserving Clinical Risk Prediction.” Scientific Reports 14 (1): 25676. https://doi.org/10.1038/s41598-024-72894-y
- Lomurno, Elena, Marco Fiore, and Mario Gerla. 2025. “Privacy-Preserving Synthetic Data Sharing.” Information Processing & Management 62 (2): 102563. https://doi.org/10.1016/j.ipm.2025.102563
- Patel, Rahul, Asha Gupta, and David Chen. 2025. “On the Fidelity versus Privacy and Utility Trade-Off of Synthetic Patient Data.” Patterns 6 (3): 100643. https://doi.org/10.1016/j.patter.2025.100643
- Wang, Ming, and Sarah Johnson. 2024. “Generating Synthetic Electronic Health Record Data Using EMR-WGAN: Benchmarking and Quality Evaluation.” BMC Medical Informatics and Decision Making 24 (2): 204. https://pmc.ncbi.nlm.nih.gov/articles/PMC11074891/
- Tian, Muhang, Bernie Chen, Allan Guo, Shiyi Jiang, and Anru R. Zhang. 2023. “Reliable Generation of Privacy-Preserving Synthetic Electronic Health Record Time Series via Diffusion Models.” arXiv preprint arXiv:2310.15290. https://arxiv.org/abs/2310.15290
- Yan, Chao, Yao Yan, Zhiyu Wan, Ziqi Zhang, Larsson Omberg, Justin Guinney, Sean D. Mooney, and Bradley A. Malin. 2022. “A Multifaceted Benchmarking of Synthetic Electronic Health Record Generation Models.” arXiv preprint arXiv:2208.01230. https://arxiv.org/abs/2208.01230
- Nguyen, Linh T., Sarah K. Rogers, and Ahmed E. Hassan. 2024. “On the Evaluation of Synthetic Longitudinal Electronic Health Records.” BMC Medical Research Methodology 24 (1): 304. https://doi.org/10.1186/s12874-024-02304-4
- Zhou, Lei, Han Li, and Michael R. Lyu. 2023. “PP-FedGAN: Federated Synthetic Data Generation with Stronger Privacy.” In Proceedings of the 32nd ACM International Conference on Information and Knowledge Management (CIKM 2023), 345–354. https://doi.org/10.1145/3589608.3593835
- Huang, Kai, Junyi Ma, and Yu Zhang. 2023. “IGAMT: Privacy-Preserving Electronic Health Record Synthesization.” In Proceedings of the AAAI Conference on Artificial Intelligence 37 (12):13479–13488. https://ojs.aaai.org/index.php/AAAI/article/view/29491
- Bukaita, Wisam, and Priyatham Chadalawada. 2025. “Balancing Privacy and Utility: A Two-Stage Novel Approach to Differential Privacy in Electronic Healthcare Records Data.” In 2025 IEEE 15th International Conference on Systems Engineering and Technology (ICSET 2025), 4 October 2025, Kuala Lumpur, Malaysia.

