Cellular Stress Proteins in Mothers of Aneuploid Fetuses
Cellular stress proteins in the proteome of serum from mothers carrying aneuploid fetuses
G. Arelí Lopez Uriarte1, Laura Elia Martínez de Villarreal1, Eliel Ruiz May2, Laura Mireya Zavala Flores3, Victor Manuel Torres de la Cruz3*
- Departamento de Genética. Facultad de Medicina y Hospital Universitario. UANL. Monterrey, Nuevo León, México
- Laboratorio de Farmacología, Escuela Militar de Graduados de Sanidad, Universidad del Ejército y Fuerza Aérea, Ciudad de México CP 11200, México
- Instituto Mexicano del Seguro Social, Centro de Investigación Biomédica del Noreste. Monterrey, México.
*Corr. author: [email protected]
OPEN ACCESS
PUBLISHED: 31 January 2026
CITATION: Lpez-Uriarte GA., et al., 2026. Cellular stress proteins in the proteome of serum from mothers carrying aneuploid fetuses. Medical Research Archives, [online] 14(1).
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI: https://doi.org/10.18103/mra.v14i1.7199
ISSN 2375-1924
ABSTRACT
Background: Aneuploidy is a genetic condition characterized by an abnormal number of chromosomes in a cell. The reduced fitness of most aneuploid cells led to the hypothesis that aneuploidy induces stress due to imbalanced global gene expression. In this study, we used proteomic analysis to analyze possible cellular stress mechanisms by comparing the serum of pregnant women between those carrying aneuploid fetuses and those carrying normal fetuses. Materials and methods: Women in their second trimester of pregnancy carrying an aneuploid fetus or a healthy fetus, as determined by ultrasound, were invited to participate in the study. The women’s serum proteome was analyzed using a UPLC-QTof/MS/MS system. We identified differentially accumulated proteins between the groups and generated a network of proteins that were up accumulated in the aneuploidy group. Results: The proteome comparison between five pregnant women carrying a fetus with aneuploidy (two with T13 and three with T18) and ten mothers carrying a normal fetus (mean gestational age: 18.5 weeks) yielded 16 differentially accumulated proteins, eight from each group. The down-accumulated proteins were zinc finger protein 13 (which contains a CCCH domain), layilin, C-C chemokine receptor type 10, transcription factor TFIIB, TATA Box Binding Protein, G protein-coupled receptor, Malate dehydrogenase (MDH1), and Heavy chain mu. The up-accumulated proteins were ATP-synthase, Helicase, TAP1, Rho GTPase-activating protein 28, Microtubule-Associated Protein 4, Serotonin transporter, Plasma membrane transporter, and Interferon gamma receptor 1. Conclusion: We have demonstrated differential expressions of proteins that interfere with cell cycle, energy production, and immunity in the serum of mothers carrying aneuploid fetuses, which may represent cell stress or an aberrant cell cycle due to the disrupted stoichiometry of genes and proteins triggered by the fetal aneuploid tissues. Also, the protein networks of overexpressed proteins showed complex evolution in many biochemical pathways of the central nervous system.
Keywords: Aneuploidies. Patau syndrome, Edwards syndrome; Prenatal proteomic profile; Maternal serum; nanoUPLC-MS/MS.
Introduction
Aneuploidy is a genetic condition that arises when there’s an abnormal number of chromosomes in a cell. While we typically have two copies of each chromosome, individuals with aneuploidy may have extras or may be missing some, which can sometimes lead to challenges such as miscarriage or, in rare situations, live births accompanied by significant developmental differences(1). The complexity of aneuploidy stems from errors that can occur during the segregation of chromosomes during meiosis, resulting in gametes and embryos with aneuploid cells throughout the body a phenomenon known as constitutional aneuploidy(2). This means that aneuploidy isn’t just a single genetic condition; it encompasses a rich diversity of scenarios.
It’s noteworthy that aneuploidy is implicated in a staggering 35-70% of fetal losses during the first half of pregnancy. In fact, trisomies, which are instances where an extra chromosome is present, occur in about 0.3% of live births(3, 4). The most prevalent form among these is trisomy 21, commonly known as Down syndrome, affecting approximately 1 in every 750 live births(5). Other types of autosomal trisomies involve chromosomes 13 and 18, with frequencies of about 1 in 20,000 and 1 in 8,000 births, respectively(6, 7). The only viable human monosomy is linked to the X chromosome, known as Turner syndrome, which occurs in about 1 in 2,500 women(8).
It’s essential to understand that aneuploid fetuses may often present a range of congenital disabilities that can raise serious concerns for newborns. During pregnancy, women experience a multitude of physiological changes in their plasma proteins, driven by an intricate interplay of signals and factors, including hormones and growth factors(9). This delicate balance plays a vital role in supporting the health and development of both the mother and the fetus(2). Unfortunately, aneuploid cells tend to have lower fitness compared to normal diploid cells, leading to the interesting hypothesis that aneuploidy may create stress within the organism due to an imbalance in gene expression.
Research on aneuploid cells from diverse species and cell lines reveals intriguing patterns: there is often an up-regulation of genes associated with stress responses and a down-regulation of those linked to the cell cycle and division. A fascinating study found that various types of aneuploidies can cause similar shifts in gene expression, regardless of the specific chromosomal changes involved(10). However, it remains a mystery how stress manifests in different contexts, depending on factors like karyotype, ploidy, and the specific cell type or tissue(2, 11).
In our previous work, we took an exciting step forward by analyzing the proteomic profile of serum from pregnant women carrying a fetus with Down syndrome. We uncovered proteins connected to immune and clotting pathways these proteins didn’t directly correspond to the gene expression of the extra chromosome, but they did hint at other potential pathological processes(12). We observed intriguing links between proteotoxic, metabolic, replication, and mitotic stress states and aneuploidy, suggesting that serum from mothers with aneuploid fetuses might carry proteins indicative of cellular stress.
Currently, we are embarking on a promising study that aims to compare serum samples from women carrying aneuploid fetuses with those carrying healthy fetuses, as assessed by normal fetal anatomy observed on ultrasound. Through proteomic analysis, we hope to explore cellular stress mechanisms further. Based on our previous findings, we are optimistic about identifying key proteomic differences that will shed light on the physiological changes associated with fetal loss. This journey promises to deepen our understanding and potentially pave the way for new insights in maternal-fetal health.
Materials and Methods
PARTICIPANTS AND SAMPLE COLLECTION
This was an observational, cross-sectional study. The population consisted of pregnant women with an aneuploid fetus and pregnant women with a healthy fetus, as determined by ultrasound, who attended the Obstetrics Clinic of the Dr. José E. González Hospital Universitario de la Universidad Autónoma de Nuevo León and the Hospital Regional Materno Infantil. The Ethics and Research Committees of the University Hospital (registration number GN14-004) approved the protocol.
Women in their second trimester of pregnancy with pregnancies in which fetal aneuploidy was suspected based on suggestive ultrasound data, such as increased nuchal translucency, fetal growth restriction, fetal heart disease, and other congenital defects, were included as cases, and pregnant women with a second-trimester fetus that was apparently healthy on the ultrasound were the controls. The exclusion criteria were multiple pregnancies, hypertensive disease, rheumatic diseases, fever, and treatment with medications.
Upon suspicion of aneuploidy based on ultrasound, patients were referred to the Genetics Department, which offers prenatal diagnosis, genetic counseling, and individualized medical follow-up. They were invited to participate in the study. Services were provided regardless of patients’ participation in the study.
PROTEOMICS STUDIES
We collected a 5 mL venous blood sample in an anticoagulant-free tube after a 4-hour fast during the second trimester of pregnancy. The blood was centrifuged at 2500 rpm and 25°C for 15 minutes to extract the serum, with the addition of 100 of protease inhibitor. It then frozen for 1 hour at -20°C and subsequently stored at -70°C. An aliquot of 25 was diluted with 75 of PBS buffer (50 mM NaH2PO4, 150 mM NaCl, pH 7.2). Protein concentration was quantified using a NanoDrop® ND-1000 spectrophotometer at 280 nm.
The depletion column was centrifuged using the Qproteome Cell Compartment Kit (Qiagen) at 500 × g, then stabilized with two aliquots of 500 PBS, and the supernatant was decanted by gravity. 100 was loaded and incubated for 10 min at room temperature, then transferred by gravity drop to a new Eppendorf tube. The total protein concentration was quantified again. A 10 kDa membrane (Millipore, Amicon Ultra-4) was conditioned with 100 of 100 mM ammonium bicarbonate (NH4HCO3) and then centrifuged at 5,000 g for 3 min. The sample was loaded onto the membrane and centrifuged at 5,000 g for 15-20 minutes at room temperature. It was then washed twice with 100 mM NH4HCO3 and centrifuged again. For trypsin digestion, 5 of 200 mM DTT (dithiothreitol) was added to 100 of the sample, and the mixture was incubated at 65°C for 45 minutes. For derivatization, 4 of 1 M iodoacetamide was added, and the mixture was incubated at room temperature for 45 min.
Sequencing-grade trypsin was prepared, then the sample was vortexed again and incubated for 24 h at 37°C in a water bath. The sample was centrifuged at 12,000 rpm for 15 minutes at room temperature. Then, 10 of MassPrep MIX 1 standard (Waters) was added, and the sample was homogenized. Subsequently, 100 was placed in a total recovery vial. All samples were subjected to chromatographic separation prior to mass spectrometry analysis using 2D-nanoUPLC (nanoAcquity Waters®). A fractionation strategy involving acid and elemental orthogonal fractionation, with six fractions and one intercalated blank injection, was performed, along with a 100% acetonitrile blank between each sample to prevent system drag or contamination. The equipment was injected with a 1.0 sample (1.0 microgram of protein) in the loading column X-Bridge C-18, 300 microns x 5 cm, 5 microns; six fractions with 11 to 65% acetonitrile was performed, applied as phase B, using 200 mM ammonium formiate as phase A. This was then passed to a trap column (a C-18, 5 microns, 150-micron x 2 cm symmetry column). The final column was a BEH analytical column (1.7 microns, 75 microns x 25 cm) at 40°C using acetonitrile gradient 95% with 0.1% formic acid as phase B.
The mass spectrometry conditions using the nano-ESI interface with a 20µ taper Tips capillary was optimized with 4000 volts for 400nL/min flow of the analytical column. The internal cone voltage was fixed in 40 Ev, and we used a small flow of nitrogen fixed at 0.9 mL/hour. The mass spectrometry acquisition method used was the protein Expression template with MSe fragmentation, running a voltage gradient of 10 to 45 eV at collision cell and V optic pathway for the TOF detector with a resolution value of 10,000 adjusted with the Glu1-fibrinopeptide m/z+2 785.8426.
DATA ANALYSIS
The raw data were acquired with MassLynx 4.1 software (Waters®). All of the fractions obtained from each sample were exported in RAW format and then processed using ProteinLynx Global Server 3.0 software (PLGS) with the MASCOT algorithm and the UniProt-SwissProt database in FASTA format for the human proteome. The reported proteins were positively identified when they met the following criteria: a 95% confidence level, more than 3 peptides per protein, and each peptide containing more than 7 precursor amino acids, which served as a filter for the previous expression calculation ratio. The sample fractions were processed to obtain a merged sample for each. The merge file per sample was pooled into two groups: a group of pregnant women with chromosomally aneuploid fetuses and a control group, and subsequent label-free expression analysis of the proteins was performed. Based on their expression levels, the proteins were categorized into three groups: up-accumulated, down-accumulated, and unique. The criterion applied to aneuploidy group: controls was an expression ratio of -1.5 to 1.0, as flags for down- and over-accumulation, respectively and a p-value of 0.6 to 1.0 like statistical significance. We identified the need to better understand the down- and up-accumulated proteins. To achieve this goal, we built networks using the PINOT virtual server v 1.1. We loaded row data set (list of Uniprot ascension ID), up-accumulated row data sets, and a mix of down- and up-accumulated row data sets. The raw data associated with the networks was exported to Cytoscape version 3.9.1 for analysis and enrichment with the String application.
Results
Forty-five pregnant women were referred to the Genetics department due to a suspicion of aneuploidy based on ultrasound. Of these, five pregnant women carrying a fetus with an aneuploidy (two with T13 and three with T18) agreed to participate in the study. The diagnosis was confirmed by karyotype of amniocytes. Ten mothers whose ultrasounds were consistent with a normal fetus were included as the control group. The mean gestational age was 18.5 weeks, and the median was 21.5 weeks gestational weeks.
There were 155 differentially accumulated proteins: 122 up-accumulated and 33 down-accumulated. We summarized the top-represented proteins in each group based on log E values (Table 1). We also found 30 proteins unique to the aneuploidy group. Proteins that were down-accumulated in the aneuploidy group included zinc finger protein 13 (ZC3H13), which contains the CCCH domain and is important in regulating nuclear RNA methylation; layilin (LAYN), a cell-surface receptor that is known to bind to hyaluronan and collagens; C-C chemokine receptor type 10 S(CCR10), which plays roles in the development, homeostasis, and function of the immune system; transcription factor TFIIB (GTF2B), involved in transcription initiation by RNA polymerase II; TATA Box Binding Protein (TBP)-(BRF1), with a role in transcription initiation by RNA polymerase III; G protein-coupled receptor (GLP2R), which activates adenyl-cyclase; Malate dehydrogenase (MDH1), which catalyzes the NAD/NADH-dependent reversible conversion of malate to oxaloacetate; and Heavy chain mu (IgM) (IGHM), which participates in antigen recognition molecules of B cells.
Proteins that were up-accumulated in the aneuploidy group included ATP-synthase (MT-ATP6), which is relevant for ATP synthesis; Helicase (SKIV2L), whose function is separating the two strands of DNA or RNA; (TAP1), which participates in pumping degraded cytosolic peptides across the endoplasmic reticulum; Rho GTPase-activating protein 28 (ARHGAP28), which has a role in regulating cell signaling by inactivating RhoA; a small GTPase that is involved in various cellular processes like cell movement, adhesion, and stress fiber formation; Microtubule-Associated Protein 4 (MAPK4), a component of the cytoskeleton with a role in the assembly of microtubules; Serotonin transporter (SLC6A4), which participates in removing serotonin from the synaptic cleft; Plasma membrane transporter SLC41A2, an anion transporter; and Interferon gamma receptor 1 (IFNGR), associated with susceptibility to infectious diseases.
Proteins that were unique to the aneuploidy group included Brevican, E3 ubiquitin-protein ligase Mdm2, WTIP, DGKH, kielin/chordin-like, and dystrobrevin alpha, which are mainly associated with tumor generation and cell signaling.
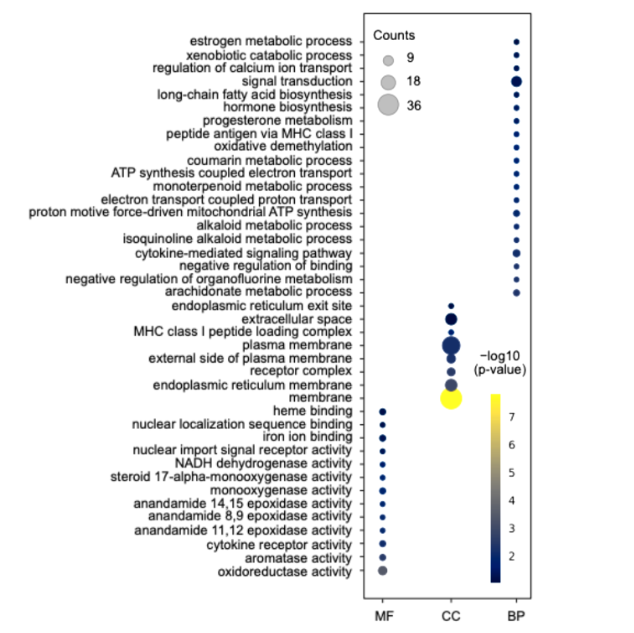
The proteins that were up accumulated in the aneuploidy group can be categorized into three groups: molecular function (MF), cellular component (CC), and biological process (BP) (Figure 1). The most abundant proteins belonged to the CC group, which is primarily associated with plasma and endoplasmic reticulum membranes. Notably, there were significant amounts of proteins related to mitochondrial function ATP synthase, NADH-ubiquinone reductase, and oxygen reductase activity; metabolism; CYP; steroid hormones; and immune pathways (IL, INF, and HLA). Fig. 1 shows the up-accumulated ontology groups associated with molecular function (MF), Cellular component (CC), and biological process (BP). The most representative for each category are Oxide reductase activity, Plasma membrane, and signal transduction, respectively.
| Protein | Function | Accumulate Type |
|---|---|---|
| Zinc finger protein 13 (ZC3H13) | Regulates nuclear RNA methylation | Down-accumulated |
| Layilin (LAYN) | Cell-surface receptor binding to hyaluronan | Down-accumulated |
| C-C chemokine receptor type 10 (CCR10) | Immune system function | Down-accumulated |
| Transcription factor TFIIB (GTF2B) | Transcription initiation by RNA polymerase II | Down-accumulated |
| TATA Box Binding Protein (TBP)-(BRF1) | Transcription initiation by RNA polymerase III | Down-accumulated |
| G protein-coupled receptor (GLP2R) | Activates adenyl-cyclase | Down-accumulated |
| Malate dehydrogenase (MDH1) | Catalyzes conversion of malate to oxaloacetate | Down-accumulated |
| Heavy chain mu (IgM) (IGHM) | Antigen recognition molecules of B cells | Down-accumulated |
| ATP-synthase (MT-ATP6) | ATP synthesis | Up-accumulated |
| Helicase (SKIV2L) | Separates DNA or RNA strands | Up-accumulated |
| TAP1 | Pumps degraded cytosolic peptides | Up-accumulated |
| Rho GTPase-activating protein 28 (ARHGAP28) | Regulates cell signaling | Up-accumulated |
| Microtubule-Associated Protein 4 (MAPK4) | Assembly of microtubules | Up-accumulated |
| Serotonin transporter (SLC6A4) | Removes serotonin from synaptic cleft | Up-accumulated |
| Plasma membrane transporter (SLC41A2) | Anion transporter | Up-accumulated |
| Interferon gamma receptor 1 (IFNGR) | Susceptibility to infectious diseases | Up-accumulated |
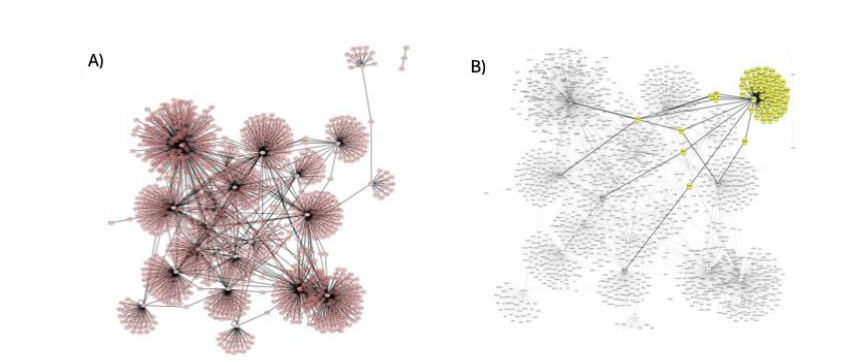
Our analysis of protein networks examined relationships within the up-accumulated proteins and between the up- and down-accumulated proteins. We confirmed the connection to the up-accumulated proteins, building the network with 1,547 nodes and 28 central nodes. String enrichment showed relevant connections with top score values for the group, which validated the biochemical relationship (Figure 2).
We also built a network of differentially accumulated proteins, with 1,656 nodes and 26 central nodes or subnetworks, as shown in Figure 3. String enrichment showed top accumulated scores for some edge connections with values of 21 for CFH (Uniprot ID P08603) and (Uniprot ID P02741), suggesting strong evidence of connection. We also observed mutual regulation of expression, as shown in the OIP5 subnetwork in Figure 3B. This explains how some down-accumulated proteins can lead to up-accumulated proteins and vice versa, while other connected proteins show significant biochemical changes.

Discussion
In fetuses with gene alterations, the equilibrium between the mother, fetus, and the placenta is abnormal, and relevant markers can be identified to determine a specific pathogenesis or its severity (9). Aneuploidy disrupts the stoichiometry of genes and proteins within a cell, altering gene dosage and the resulting protein products, leading to various cellular consequences. Overexpression of genes associated with aneuploid chromosomes disrupts proteostasis(13,14).
In the present study, a comparison of serum from mothers carrying aneuploid fetuses revealed differential expression of several proteins compared with serum from mothers with normal fetuses. There were up- and down-accumulated proteins associated with transcription, energy, immunity, cell cycle, and other essential cellular functions that may reflect cellular stress due to gene expression in the aneuploid fetus.
Enzymes involved in transcription, such as TFIIB (GTF2B) and TATA Box-Binding Protein (BRF1)-associated factors, were downregulated. It has been reported that either down- or up-accumulated transcription factors in aneuploidies can disrupt gene expression patterns and contribute to cellular stress, impacting cell growth and function(15); thus, these proteins may be considered markers of cellular stress.
Both TFIIB (GTF2B) and TATA Box-Binding Protein (BRF1)-associated factors have been linked to various developmental and cellular issues, including maintaining standard cellular structure and function, cell growth, and altered immune responses. Reduced levels of TFIIB (GTF2B) could decrease overall transcription rates across many genes by affecting RNA polymerase II recruitment, thereby impacting various cellular processes. This protein is vital for maintaining the standard cellular structure and function, and it also serves as the basis for developmental changes, such as differentiation and morphogenesis. The GTF2B gene regulates cell apoptosis and invasion in trophoblast tissues and has been reported to be involved in the pathological process of preeclampsia(16).
Under expression of TATA Box Binding Protein (BRF1), a subunit of the RNA polymerase III transcription initiation factor, is associated with defects in cell growth and proliferation, and may even lead to embryonic lethality, suggesting that it plays a crucial role in cellular processes essential for proper development and homeostasis. Mutations in BRF1 reduce protein activity and cause neurodevelopmental anomalies; therefore, BRF1-mediated Pol III transcription is required for normal cerebellar and cognitive development(15).
Helicase (SKIV2L), a cell-cycle protein, was over accumulated in the aneuploidy group. Helicases play a crucial role in various cellular processes by unwinding DNA or RNA and breaking the hydrogen bonds that hold the double helix together. This unwinding is essential for DNA replication, where each strand serves as a template for the synthesis of a new DNA molecule. Helicases are also involved in DNA repair, RNA transcription, and other cellular processes that require access to the genetic material(17). Given that helicases play a crucial role in regulating and maintaining chromosomal DNA, it is logical to assume that their dysfunction leads to unfavorable states(18). When a helicase protein is over-represented, it suggests that the cell’s demand for its unwinding activity might be increased, or that its regulation has been disrupted. The over-representation of specific DNA repair helicases may indicate an increased need to repair DNA damage, potentially resulting from exposure to toxins or other stressors(18).
Studies of trisomy 21 have indicated a potential link between increased variability in gene expression and the presence of extra or missing chromosomes. While this is not a direct increase in helicase expression, some research suggests that aneuploidy can lead to a global, and sometimes stochastic, increase in gene expression variability, which can affect cell-type-specific genes and potentially impact phenotypic outcomes. This variability can be observed in both single-cell and single-nucleus RNA sequencing datasets(18). In our study, the overrepresentation of helicase could indicate disrupted regulation and/or cellular stress.
Another up-accumulated protein in the aneuploidy group is the Rho GTPase-activating protein (ARHGAP28). Rho GTPase-activating proteins (RhoGAPs) are negative regulators (i.e., inactivators) of Rho GTPase enzymes, which are key regulators of the cell cytoskeleton and cell division. The cytoskeleton is essential for cell adhesion, migration, and contraction, and the microtubule-associated protein; it has been suggested that this may lead to the cellular stress reported in aneuploid tissues(2, 20).
The increased sensitivity of aneuploid cells to energy stress and disruption of protein homeostasis can lead to impaired cell proliferation and growth. The overexpression of this protein in the aneuploidy group may reflect impaired cell division.
Regarding proteins related to immunity, there were up- and down-accumulated proteins in each group. TAP1 (Transporter associated with Antigen Processing 1) (TAP1) has a vital role in the TAP complex (transporter associated with antigen processing), where it transports peptides from the cytoplasm to the endoplasmic reticulum, where they bind to MHC class I molecules. During pregnancy, the TAP1 protein exhibits unique expression patterns in placental tissues, particularly in trophoblasts, where it plays a crucial role in immune system interactions. While TAP1 is vital for antigen presentation by MHC class I molecules and for immune responses, its overaccumulation in certain placental cell types, such as extravillous cytotrophoblasts, is essential for successful placentation and fetal-maternal immune tolerance(21). The over-accumulation of this protein has been analyzed in cancer, showing a dual effect. At the same time, it can enhance the immune response and potentially improve prognosis; it can also contribute to tumor progression and metastasis, especially in the context of aneuploidy and immune evasion strategies employed by cancer cells(22). The overaccumulation of TAP1 observed in the present study may indicate cellular stress.
Another up-accumulated immune-related protein was Interferon gamma receptor 1 (IFNGR1), which functions as a receptor for interferon gamma (IFN-γ), a cytokine crucial for immune responses. It plays a key role in activating immune cells and regulating immune responses against infections. Overexpression of IFNGR1 can significantly alter cellular responses to IFN-γ signaling. While IFNGR1 expression is typically sufficient and can be upregulated by various factors, its overexpression can lead to diverse outcomes depending on the cellular context, including altered immune responses, increased sensitivity to IFN-γ, and potential implications in cancer progression. While IFN-γ is generally beneficial, excessive stimulation of IFN-γ can also be pathogenic. Overexpression of IFNGR1, leading to heightened IFN-γ signaling, could contribute to inflammatory conditions or exacerbate existing inflammatory responses(23).
In a previous study, it was demonstrated that individuals with trisomy 21 can exhibit overexpression of IFN receptors, increased IFN hypersensitivity, altered expression of pro- and anti-inflammatory cytokines, and impaired chemotaxis of mononuclear cells, all of which can impair innate immune responses. Other studies have identified that adaptive immunity in individuals with trisomy 21 is also impacted by altered AIRE expression, thymic hypoplasia, and cortical atrophy, which can collectively contribute to higher rates of apoptosis in B and T cells, T helper cells that are more resistant to T regulatory (Treg) cell suppression, and an altered immunoglobulin (Ig) pattern of increased IgG but decreased IgM and IgE levels(24). In the present study, the IFN gamma receptor was also overexpressed in the serum of mothers carrying fetuses with T13 and T18 aneuploidies, suggesting an impaired immune response due to cellular stress.
There were two down-accumulated proteins related to immunity: the heavy chain of the IgM (IgHM) and chemokine receptor type 10 (CCR10). The IgM gene encodes the mu (μ) heavy chain of the immunoglobulin M (IgM) antibody, which is a crucial component of the body’s adaptive immune response. Down-accumulation of the IgM protein can occur due to a variety of factors, including chromosomal instability, altered gene expression patterns, and disruptions in the unfolded protein response (UPR). This can lead to immune dysregulation within the tumor microenvironment, potentially affecting cancer progression(25). The underexpression of this protein may be a marker of cellular stress.
The C-C chemokine receptor type 10 (CCR10), along with CCR3, is the receptor for chemokine CCL28, which plays a critical role in the establishment and maintenance of pregnancy by influencing the migration and proliferation of cells at the maternal-conceptus interface. These three proteins are expressed in the uterus during pregnancy. They are involved in the recruitment of immune cells and the migration of trophectoderm cells, which are crucial for successful implantation and placental development(26). Their under expression, in addition to cell stress, may reflect the process involved in the high frequency of abortion in these analyses.
Proteins affecting cell cycle and other functions were also down- and up-accumulated. Layilin, a protein that acts as a surface receptor for hyaluronan (HA), plays a vital role in cell adhesion, motility, regulation of cell spreading, and migration. Its expression levels can vary across cell types and under different conditions, although downregulation, as observed in our study, is uncommon. Aneuploidy can affect gene expression. It has been reported that under expression of layilin can impact cell behavior and potentially contribute to specific disease states(27).
Its essential role in diverse cell functions may reflect the altered development of the aneuploid fetuses.
MAP4 is a major microtubule-associated protein that stabilizes microtubules and regulates their dynamics. In normal cells, this is crucial for proper chromosome segregation during mitosis. It is involved in various cellular processes relevant to pregnancy, including cell migration and the immune response(28). In aneuploid cells, however, aberrant microtubule dynamics can contribute to chromosome missegregation and the development of aneuploidy. Overaccumulation of the MAP4 protein, which stabilizes microtubules, can affect cellular processes related to cell growth, migration, and, potentially, cancer development(29). The over-accumulation of MAP4 identified in our study may indicate the perpetuation of chromosome missegregation and reflect altered cellular processes.
Two channel transporters were found to be up accumulated. One is the SLC6A4 protein, also known as the serotonin transporter, which plays a crucial role in pregnancy by regulating serotonin levels and influencing both maternal and fetal health. The second is SLC41A2; this magnesium transporter helps regulate the movement of magnesium ions in and out of cells.
SLC6A4 is involved in placental development, fetal neurodevelopment, and the mother’s mental health during and after pregnancy. In aneuploid tissues, the function of SLC6A4 can be impacted, potentially leading to altered serotonin signaling and contributing to various neurological and neuropsychiatric conditions. In patients with Down syndrome, increased levels of serotonin transporter have been reported, suggesting a potential link between the genetic changes in Down syndrome and disruptions in serotonin pathways, which could contribute to the various developmental and cognitive challenges associated with the condition(30,31).
Magnesium is essential for many cellular functions, including DNA replication, RNA synthesis, and protein folding. In the context of aneuploidy, the role of SLC41A2 in magnesium transport suggests a potential connection. Overexpression of SLC41A2 could increase magnesium uptake and potentially influence cell proliferation. Studies show that higher levels of SLC41A2 expression correlate with faster cell growth rates, particularly in magnesium-deficient conditions(32).
Zinc Finger CCCH-Type Containing 13 (ZC3H13) protein plays an essential role in the regulation of RNA methylation in the nucleus, specifically in the modification of N6-methyladenosine (m6A). Numerous studies have shown that m6A modifications play essential roles in multiple biological and pathological processes: hematopoietic development, central nervous and system development, the adaptive and innate immune system, carcinogenesis and the tumor microenvironment, as well as gametogenesis and early embryo development. Dysfunction generally results in various diseases due to abnormal m6A modifications of the target gene(33). Methylation of m6A plays a critical role in both normal embryonic development and the establishment and maintenance of the maternal-fetal interface during pregnancy. Dysregulation of m6A can contribute to gestational diseases and adverse birth outcomes(34). Disrupting ZC3H13 function can impair the self-renewal and differentiation of mesenchymal stem cells, potentially leading to developmental defects. Specifically, ZC3H13 depletion has been shown to trigger mESC differentiation, which is not ideal during early embryonic development(35).
The Glucagon-Like Peptide-2 Receptor (GLP2R) is a G protein-coupled receptor that mediates the effects of glucagon-like peptide-2 (GLP-2), a hormone produced in the gut that plays a crucial role in maintaining intestinal health and function. It promotes intestinal growth, enhances nutrient absorption, and improves the gut barrier function(36). In a previous study, the GLP2R gene was found to be methylated in SGA patients(37). The down-accumulation of this protein in our study may be related to the low weight and low size frequently observed in children with T13 and T18.
The over-expressed PPI network shows important connections. We found 22 central nodes of the clusters, 20 of which have biochemical functions in the nervous system. Examples include Q72417, P51116, P51114, and Q13521, which are involved in intellectual disability syndrome(38). Some of them had total scores over 5 in the String enrichment report.
Furthermore, the PPI composed of overexpressed and underexpressed proteins had 26 clusters, of which 4 are of significant biochemical relevance, such as DGAT1 (075907), which is associated with gliomas and cerebral tumors(39), and the other central node, MDH1 is related to the synapsis process and intellectual disability(40), SCL38A7(Q9NVC3) is an amino acids transporter involved in synapsis and intellectual disability(41) and TMEM234 is related to Meckel syndrome subtype(42).
We analyzed the potential mutual regulation of expression between the two groups. Here we show an example as evidence of this possibility: the OIP5(043482) protein appears to be an intermediate connection between three other central node proteins: MDC1(Q5JP55), NCKAP5L(Q9HCH0) and IL16(Q9UP18). The DYRK1A protein may be under-expressed consequently of OIP5 overexpression, and it has been reported in gliomas. This protein is also involved in the autosomal dominant 7C(MRD7) seek and the gene is in 21q22.13.
Conclusion
In the present study, we have demonstrated the differential accumulation in the serum of mothers carrying aneuploid fetuses of proteins that could interfere with the cell cycle, energy production, and immunity. This may represent cell stress or an aberrant cell cycle due to the disrupted stoichiometry of genes and proteins triggered by the fetal aneuploid tissues.
Conflicts of Interest: The authors declare no conflicts of interest.
References
- Terry Hassold, Heather Hall, Patricia Hunt, The origin of human aneuploidy: where we have been, where we are going, Human Molecular Genetics, Volume 16, Issue R2, 15 October 2007, Pages R203-R208, https://doi.org/10.1093/hmg/ddm243 S.
- Zhu J, Tsai HJ, Gordon MR, Li R. Cellular Stress Associated with Aneuploidy. Dev Cell. 2018 Feb 26; 44(4):420-431. doi: 10.1016/j.devcel.2018.02.002. PMID: 29486194; PMCID: PMC6529225.
- Hassold, T., Hunt, P. To err (meiotically) is human: the genesis of human aneuploidy. Nat Rev Genet 2, 280-291 (2001). https://doi.org/10.1038/35066065
- Laura Kasak, Kristiina Rull, Maris Laan, Chapter 21 – Genetics and Genomics of Recurrent Pregnancy Loss, Editor(s): Peter C.K. Leung, Jie Qiao, Human Reproductive and Prenatal Genetics, Academic Press, 2019, Pages 463-494, ISBN 9780128135709 https://doi.org/10.1016/B978-0-12-813570-9.00021-8.
- Creasy MR, Crolla JA. Prenatal mortality of trisomy 21 (Down syndrome). Lancet 1974. i:473.
- Presson AP, Partyka G, Jensen KM, Devine OJ, Rasmussen SA, McCabe LL, McCabe ER. Current estimate of Down Syndrome population prevalence in the United States. J Pediatr. 2013 Oct;163(4):1163-8. doi: 10.1016/j.jpeds.2013.06.013. Epub 2013 Jul 23. PMID: 23885965; PMCID: PMC4445685.
- Satgé D, Nishi M, Sirvent N, Vekemans M, Chenard MP, Barnes A. A tumor profile in Patau syndrome (trisomy 13). Am J Med Genet A. 2017 Aug;173(8):2088-2096. doi: 10.1002/ajmg.a.38294. Epub 2017 May 25. PMID: 28544599.
- Cereda A, Carey JC. The trisomy 18 syndrome. Orphanet J Rare Dis. 2012 Oct 23;7:81. doi: 10.1186/1750-1172-7-81. PMID: 23088440; PMCID: PMC3520824.
- Cho WK. Lifelong medical challenges and immunogenetics of Turner syndrome. Clin Exp Pediatr. 2024 Nov;67(11):560-568. doi: 10.3345/cep.2024.00430. Epub 2024 Jul 31. PMID: 39091155; PMCID: PMC11551602.
- Romero R, Erez O, Maymon E, Chaemsaithong P, Xu Z, Pacora P, Chaiworapongsa T, Done B, Hassan SS, Tarca AL. The maternal plasma proteome changes as a function of gestational age in normal pregnancy: a longitudinal study. Am J Obstet Gynecol. 2017 Jul;217(1):67.e1-67.e21. doi: 10.1016/j.ajog.2017.02.037. Epub 2017 Mar 3. PMID: 28263753; PMCID: PMC5813489.
- Eduardo M. Torres, Maitreya J. Dunham, and Angelika Amon Transcriptional consequences of aneuploidy, July 16, 2012 109 (31) 12644-12649 https://doi.org/10.1073/pnas.1209227109
- Dürrbaum M, Storchová Z. Effects of aneuploidy on gene expression: implications for cancer. FEBS J. 2016 Mar;283(5):791-802. doi: 10.1111/febs.13591. Epub 2015 Dec 8. PMID: 26555863.
- López Uriarte GA, Burciaga Flores CH, Torres de la Cruz VM, Medina Aguado MM, Gómez Puente VM, Romero Gutiérrez LN, Martínez de Villarreal LE. Proteomic profile of serum of pregnant women carrying a fetus with Down syndrome using nano uplc Q-tof ms/ms technology. J Matern Fetal Neonatal Med. 2018 Jun;31(11):1483-1489. doi: 10.1080/14767058.2017.1319923. Epub 2017 May 3. PMID: 28412848.
- Zhang S, Wang R, Huang C, Zhang L, Sun L. Modulation of Global Gene Expression by Aneuploidy and CNV of Dosage Sensitive Regulatory Genes. Genes (Basel). 2021 Oct 12;12(10):1606. doi: 10.3390/genes12101606. PMID: 34681000; PMCID: PMC8535535.
- Kojima S, Cimini D. Aneuploidy and gene expression: is there dosage compensation? Epigenomics. 2019 Dec;11(16):1827-1837. doi: 10.2217/epi-2019-0135. Epub 2019 Nov 22. PMID: 31755744; PMCID: PMC7132608.
- Borck G, Hög F, Dentici ML, Tan PL, Sowada N, Medeira A, Gueneau L, Thiele H, Kousi M, Lepri F, Wenzeck L, Blumenthal I, Radicioni A, Schwarzenberg TL, Mandriani B, Fischetto R, Morris-Rosendahl DJ, Altmüller J, Reymond A, Nürnberg P, Merla G, Dallapiccola B, Katsanis N, Cramer P, Kubisch C. BRF1 mutations alter RNA polymerase III-dependent transcription and cause neurodevelopmental anomalies. Genome Res. 2015 Feb;25(2):155-66.
- Zhou W, Wang H, Yang Y, Guo F, Yu B, Su Z. Trophoblast Cell Subtypes and Dysfunction in the Placenta of Individuals with Preeclampsia Revealed by Single-Cell RNA Sequencing. Mol Cells. 2022 May 31;45(5):317-328. doi: 10.14348/molcells.2021.0211. PMID: 35289305; PMCID: PMC9095508.
- Xin Qin, Jing Wang, Xing Wang, Tao Huang, Zhiqing Fang, Lei Yan, Yidong Fan, Dawei Xu, Widespread genomic/molecular alterations of DNA helicases and their clinical/therapeutic implications across human cancer, Biomedicine & Pharmacotherapy, Volume 158, 2023,114193, ISSN 0753-3322, https://doi.org/10.1016/j.biopha.2022.114193
- Uchiumi F, Seki M, Furuichi Y. Helicases and human diseases. Front Genet. 2015 Feb 12;6:39. doi: 10.3389/fgene.2015.00039. PMID: 25729389; PMCID: PMC4325929.
- Upadhya S, Klein JA, Nathanson A, Holton KM, Barrett LE. Single-cell analyses reveal increased gene expression variability in human neurodevelopmental conditions. Am J Hum Genet. 2025 Apr 3;112(4):876-891. doi: 10.1016/j.ajhg.2025.02.011. Epub 2025 Mar 7. PMID: 40056913; PMCID: PMC12081229.
- Sun Young Moon, Yi Zheng, Rho GTPase-activating proteins in cell regulation, Trends in Cell Biology, Volume 13, Issue 1, 2003, Pages 13-22, ISSN 0962-8924, https://doi.org/10.1016/S0962-8924(02)00004-1.
- Clover, L.M., Sargent, I.L., Townsend, A., Tampé, R. and Redman, C.W.G. (1995), Expression of TAP1 by human trophoblast. Eur. J. Immunol., 25: 543-548. https://doi.org/10.1002/eji.1830250236.
- Wang Z-D, Tian X, Wang Y, Wang J-J, Ye S-Q, Huang Y-Q, Qu Y-Y, Chang K, Shi G-H, Ye D-W and Gu C-Y (2022) The expression and prognostic value of transporter 1, ATP binding cassette subfamily B member in clear cell renal cell cancer with experimental validation. Front. Oncol. 12:1013790. doi: 10.3389/fonc.2022.1013790.
- Crisler WJ, Eshleman EM, Lenz LL. Ligand-induced IFNGR1 down-regulation calibrates myeloid cell IFNγ responsiveness. Life Sci Alliance. 2019 Oct 4;2(5):e201900447. doi: 10.26508/lsa.201900447. PMID: 31585982 PMCID: PMC6778285.
- Hom, B., Boyd, N.K., Vogel, B.N. et al. Down Syndrome and Autoimmune Disease. Clinic Rev Allerg Immunol 66, 261-273 (2024). https://doi.org/10.1007/s12016-024-08996-2.
- Xian S, Dosset M, Almanza G, Searles S, Sahani P, Waller TC, Jepsen K, Carter H, Zanetti M. The unfolded protein response links tumor aneuploidy to local immune dysregulation. EMBO Rep. 2021 Dec 6;22(12):e52509. doi: 10.15252/embr.202152509. Epub 2021 Oct 26. PMID: 34698427; PMCID: PMC8647024.
- Choi Y, Seo H, Han J, Yoo I, Kim J, Ka H. Chemokine (C-C motif) Ligand 28 and Its Receptor CCR10: Expression and Function at the Maternal-Conceptus Interface in Pigs. Biol Reprod. 2016 Oct 1;95(4):84. doi: 10.1095/biolreprod.116.141903. Epub 2016 Aug 31. PMID: 27580990.
- Pan JH, Zhou H, Cooper L, Huang JL, Zhu SB, Zhao XX, Ding H, Pan YL, Rong L. LAYN Is a Prognostic Biomarker and Correlated With Immune Infiltrates in Gastric and Colon Cancers. Front Immunol. 2019 Jan 29;10:6. doi: 10.3389/fimmu.2019.00006. PMID: 30761122; PMCID: PMC6362421.
- Sánchez C, Díaz-Nido J, Avila J. Phosphorylation of microtubule-associated protein 2 (MAP2) and its relevance for the regulation of the neuronal cytoskeleton function. Prog Neurobiol. 2000 Jun;61(2):133-68. doi: 10.1016/s0301-0082(99)00046-5. PMID: 10704996.
- Bulinski JC, McGraw TE, Gruber D, Nguyen HL, Sheetz MP. Overexpression of MAP4 inhibits organelle motility and trafficking in vivo. J Cell Sci. 1997 Dec;110 (Pt 24):3055-64. doi: 10.1242/jcs.110.24.3055. PMID: 9365275.
- Alasmari F, Ahmad A, Alsanea S, Hammad AM, Al-Qerem W. Current insights and prospects for the pathogenesis and treatment of clinical manifestations associated with Down syndrome through neurotransmitter, inflammatory, and oxidative stress pathways. Front Pharmacol. 2025 Apr 28;16:1592277. doi: 10.3389/fphar.2025.1592277. PMID: 40356974; PMCID: PMC12066560.
- Gulesserian, T., Engidawork, E., Cairns, N., & Lubec, G. (2000). Increased protein levels of serotonin transporter in frontal cortex of patients with Down syndrome. Neuroscience letters, 296(1), 53-57.
- Sahni J, Nelson B, Scharenberg AM. SLC41A2 encodes a plasma-membrane Mg2+ transporter. Biochem J. 2007 Jan 15;401(2):505-13. doi: 10.1042/BJ20060673. PMID: 16984228; PMCID: PMC1820800.
- Liu H, Zheng J, Liao A. The regulation and potential roles of m6A modifications in early embryonic development and immune tolerance at the maternal-fetal interface. Front Immunol. 2022 Sep 26;13:988130. doi: 10.3389/fimmu.2022.988130. PMID: 36225914; PMCID: PMC9549360.
- Wu S, Liu K, Zhou B, Wu S. N6-methyladenosine modifications in maternal-fetal crosstalk and gestational diseases. Front Cell Dev Biol. 2023 Mar 16;11:1164706. doi: 10.3389/fcell.2023.1164706. PMID: 37009476; PMCID: PMC10060529.
- Wen J, Lv R, Ma H, Shen H, He C, Wang J, Jiao F, Liu H, Yang P, Tan L, Lan F, Shi YG, He C, Shi Y, Diao J. Zc3h13 Regulates Nuclear RNA m6A Methylation and Mouse Embryonic Stem Cell Self-Renewal. Mol Cell. 2018 Mar 15;69(6):1028-1038.e6. doi: 10.1016/j.molcel.2018.02.015. PMID: 29547716; PMCID: PMC5858226.
- Eric D. Shina, Daniel J. Druckerb, C and Patricia L. Brubakera, B. Glucagon-like peptide 2: an update. Current Opinion in Endocrinology & Diabetes 2005, 12:63-71.
- Angulo M, Ramirez-Montaño D, Torres-Canchala L, García X, Lemus R, Aristizabal AM, Floyd-Aristizábal D, Dávalos DM, Diaz-Ordoñez L, Pachajoa H. Methylation Status of GLP2R, LEP and IRS2 in Small for Gestational Age Children with and without Catch-up Growth. J Clin Res Pediatr Endocrinol. 2021 Jun 2;13(2):136-145. doi: 10.4274/jcrpe.galenos.2020.0070. Epub 2020 Sep 17. PMID: 32936762; PMCID: PMC8186343.
- Fatemi SH., Folsom TD. The Role of Fragile X Mental Retardation Protein in Major Mental Disorders. Neuropharmacology. 2011 June; 60(7-8): 1221-1226. doi:10.1016/j.neuropharm.2010.11.011.
- Xian Ch., Feng G., Meixia P., Xiaoning Wu., Yaogang Z., Chunyan W., Zhihua T., Chunming Ch., Riu Z., Vinay P., Craig H., Xiaokui M., Xianlin H., Arnab Ch., Deliang G., Targeting DGAT1 Ameliorates Glioblastoma by Increasing Fat Catabolism and Oxidative Stress. Cell Metabolism 32, 229-242 August 4, 2020, https://doi.org/10.1016/j.cmet.2020.06.002.
- Broeks M., Shamseldin HE., Alhashem A., Hashem M., Abdulwahab F., Alshedi T., Alobaid I., Zuartkruis F., Westland D., Fuchs S., Verhoeven-Duif NM., Jans JJM., Alkuraya FS. MDH1 deficiency is a metabolic disorder of the malate aspartate shuttle associated with early onset severe encephalopathy. Human Genetics (2019) 138:1247-1257 https://doi.org/10.1007/s00439-019-02063-z.
- R Bachmann-Gagescu, J C Dempsey, I G Phelps, BJO Roak, D M Knutzen, T C Rue, G E Ishak, C R Isabella, N Gorden, J Adkins, E A Boyle, NdeLacy, DO Day, A Alswaid, Radha Ramadevi A, L Lingappa, C Lourenço, L Martorell, À Garcia-Cazorla, H Ozyürek, G Haliloğlu, B Tuysuz, M Topçu, U P Chance, M A Parisi, I A Glass, J Shendure, D Doherty. Joubert syndrome: a model for untangling recessive disorders with extreme genetic heterogeneity. J Med Genet 2015;52:514-522. doi:10.1136/jmedgenet-2015-103087.




