Comparison of Classical vs. Modern Mycobacterial Detection
Prospective comparison of Classical and Modern Molecular mycobacterial detection techniques in cattle infected with M. avium subspecies paratuberculosis
Robert J. Greenstein1,2, Sheldon T. Brown3
- Department of Surgery James J. Peters Veterans Affairs Medical Center Bronx NY USA.
- Laboratory of Molecular Surgical Research James J. Peters Veterans Affairs Medical Center Bronx NY USA.
- ACOS Research. James J. Peters Veterans Affairs Medical Center Bronx NY USA.
OPEN ACCESS
PUBLISHED: 31 October 2025
CITATION: Greenstein, RJ., and Brown, ST., 2025. Prospective comparison of Classical and Modern Molecular mycobacterial detection techniques in cattle infected with M. avium subspecies paratuberculosis. Medical Research Archives, [online] 13(10). https://doi.org/10.18103/mra.v13i10.6979
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v13i10.6979
ISSN 2375-1924
ABSTRACT
Background: “Classical” mycobacterial identification is difficult to perform, time consuming, inefficient and often insensitive. Modern techniques, incorporating molecular methods, may be more rapid, less expensive, more sensitive and more accurate. We prospectively compared “classical” and modern mycobacterial detection techniques.
Methods: A plaque/qPCR study has been published on the blood of a herd of Johne disease (cattle infected with M. avium subspecies paratuberculosis (MAP.)) Simultaneously aliquots were submitted for “classical” mycobacterial identification. Plaque/qPCR could be completed in <3days. “Classical” cultures were followed for a minimum of 2 years.
Results: Plaque/qPCR MAP positivity were observed in 74% of Johne and 34% of Control animals. “Classical” detection on appropriate culture medium were seen in 19% in Johne disease and 18% Controls.
Conclusions: “Classical” mycobacterial detections methodology has demonstrably been supplanted by modern molecular methods.
Keywords
mycobacterial detection, M. avium, Johne disease, molecular methods, classical methods
Introduction
The identification and culture of mycobacteria has a long and storied history. In 1874, in lepromatous patients, Hansen microscopically identified bacilli. In 1882 Koch identified and cultured the tuberculosis bacillus, for which he received the 1905 Nobel prize in Medicine. Specific staining of the mycolic acid rich cell wall of mycobacteria enhanced detection of mycobacteria in 1882 by Ziehl and in 1883 by Neelsen. Over the ensuing 150 years a multitude of efforts occurred to improve mycobacterial detection and culture evolved. In the 1930’s the agar-based slopes of Lowenstein and Jensen became the norm. This led to the evolution of semi-solid agar in 1955 followed by the introduction of pure liquid culture media by Middlebrook in 1958. The introduction of radiometric 14C led to a detection system (Bactec 460®) of unsurpassed sensitivity of mycobacterial growth rates. Regrettably, because the Bactec460 system is only semi-automatic and onerous regulations about disposing of the 14C, with its half-life of 5700 years, the Bactec460 is no longer produced or supported. The present fluorometric Bactec 960® is the most prevalent extant mycobacterial growth detection system. In our experience it is less sensitive than the radiometric Bactec460. All of these detection systems were invariably time-consuming process, taking weeks, months or even years.
A potentially major improvement in the detection of viable mycobacteria uses mycobacteriophage to infect the viable mycobacteria and RT PCR to identify the specific DNA of that mycobacterium. The use of mycobacteriophage based assays has multiple advantages to the aforementioned “Classical method”, including improved detection sensitivity and speed. We previously published a phage/qPCR study on blood from a herd of cattle naturally infected with Mycobacterium avium subspecies paratuberculosis (MAP) known as Johne disease (JD) and Control animals. The phage component of the assay was completed within 36 hours of samples arriving in the laboratory. MAP was detected in 72% of the JD and, unsuspectedly, in 36% of the controls. Simultaneously to performing the phage/qPCR assay, samples were submitted for standard “Classical” mycobacterial culture (vide supra, specifically MGIT®.)
The purpose of this manuscript is to report on the comparison of the “Classical method” of mycobacterial culture with our published phage/qPCR data. The “Classical method” cultures were studied for a minimum of two years following inoculation.
Methods
Except for the subculture from Pozzato medium and MGIT tubes when indicated, and cultured onto HEYM slopes, all Methods have been previously published in detail. In brief: From March 15, 2019, until January 21, 2020, we received a total of 150 blood samples from USDA herds in Ames Iowa. Of the total 49 animals, 32 had Johne and 17 were Control. Conventional blood and stool tests for MAP and/or its signal was compared with a MAP phage/qPCR assay. White blood cells were isolated from cattle whole blood using a Ficoll gradient (Becton-Dickinson: Vacutainer® CPT™ Cell Preparation Tube with Sodium Citrate (8 ml), Cat #362761), lysed and 25% of the lysate was then cultured using both Pozzato and 7H-9 media in MGIT® (Becton Dickinson) both of which contained PANTA and Mycobactin J. Occasionally so many plaques occurred, the appearance of the Petri dish appeared as a spurious “clear”. When this “clearance” occurred when plating 0.5 ml. of the lysate, we began to simultaneously plate 0.2 ml of the lysate. Some Control animals were appropriately negative in the plaques assay. Idiosyncratically occasionally Control animals were positive in the plaque assay as previously reported.
Every isolate from the 150 received samples were submitted to MGIT (7H9 based) and Pozzato culture. Both contained PANTA. Cultures were followed with serial Optical Densities (OD’s.) When turbidity was detected, cultures were subcultured onto HEYM. Slopes were placed in incubators at 37oC and regularly observed. When evidence of growth was detected (by increase in turbidity, increasing Optical Densities or Positivity on Bactec960® MGIT® tubes,) samples were sub-cultured onto Herrold’s egg yolk slopes (Herrold’s egg yolk agar with amphotericin, nalidixic acid, vancomycin, with mycobactin J. SKU 222232: GTIN 00382902222323 BD. New Jersey) Visually identified colonies, detected from 13 to 28 months following inoculation) were sub-cultured in 7H-9 to permit nucleic acid identification in the future. All cultures were supplemented with Mycobactin J. All flasks were incubated at 37oC. Identified colonies were harvested and subjected to Ziehl-Neelsen staining.
Results:
A total of 150 samples were received and assayed from 17 Control and 32 Johne animals.
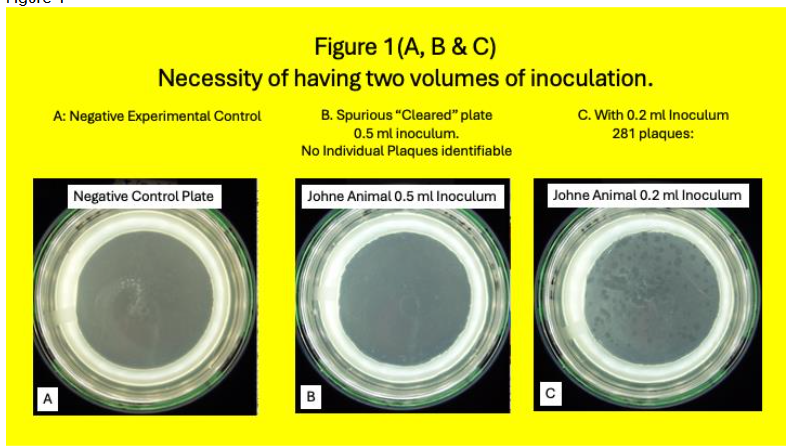
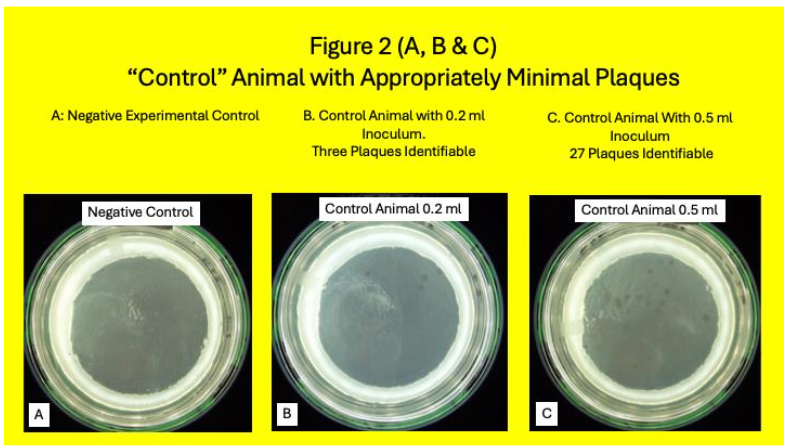
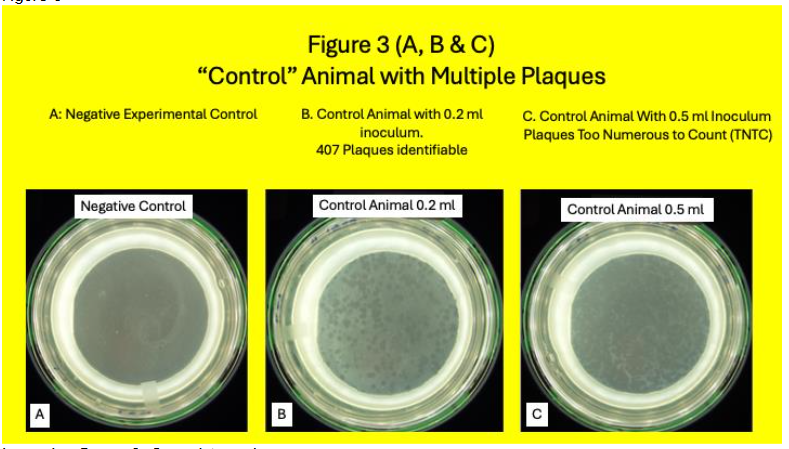
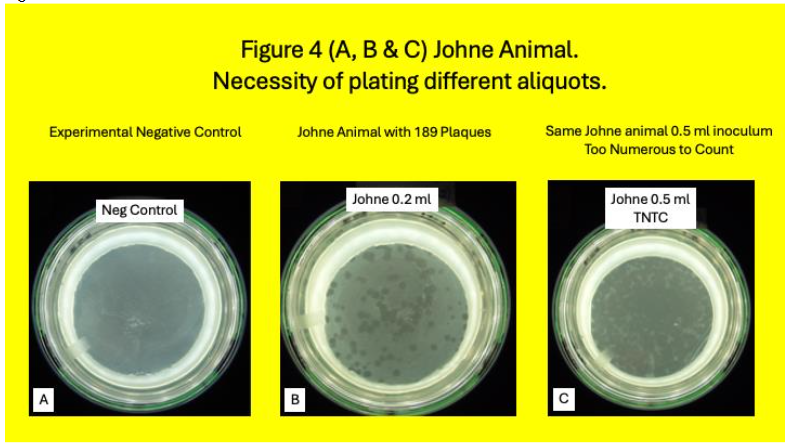
Samples were subcultured onto HEYM and followed for a minimum of 2 years. Colonies were seen HEYM slopes in 18% (3/17) Control animals (Table 1& Figure 5.) Of these Three Control animals where identifiable colonies grew, MAP plaque IS 900 qPCR was positive in one.
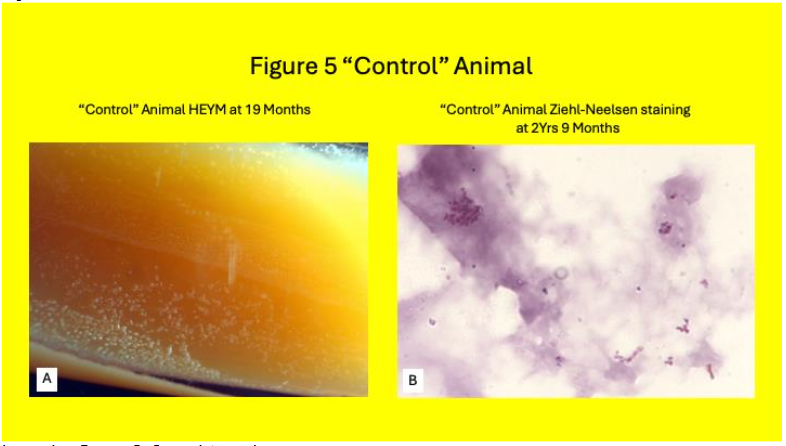
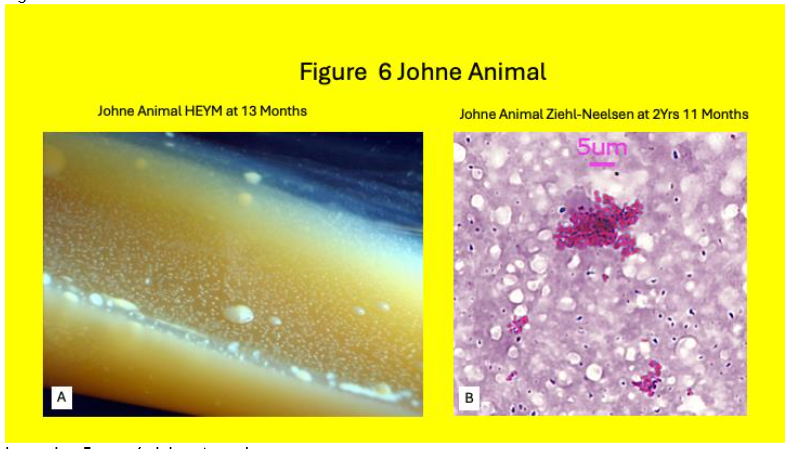
Colonies were seen HEYM slopes in 19% (6/32) Johne animals (Table 2 & Figure 6.) Of the six Johne animals with identifiable colonies on HEYM slopes, four were plaque IS900 DNA positive.
| Control Animal # | HEYM Colonies Identified | Negative Plaque # | IS 900 DNA qPCR Positive | IS 900 DNA qPCR Negative |
|---|---|---|---|---|
| 1c | – | 11 | + | |
| 2c | + | 1000 | + | |
| 3c | – | 3 | – | |
| 4c | – | 238 | + | |
| 5c | – | 688 | + | |
| 6c | – | 29 | + | |
| 7c | – | 10 | – | |
| 8c | + | 1 | – | |
| 9c | – | 151 | – | |
| 10c | + | 47 | – | |
| 11c | – | 4 | – | |
| 12c | – | 1 | – | |
| 13c | – | 8 | – | |
| 14c | – | 0 | – | |
| 15c | – | 4 | – | |
| 16c | 6 | – |
% positive (# positive/total tested) 18% (3/17)
| Johne Animal Chart # | HEYM Culture + | Culture – | Plaque # | IS 900 DNA qPCR Positive | IS 900 DNA qPCR Negative |
|---|---|---|---|---|---|
| 1j | – | 27 | + | ||
| 2j | – | 312 | + | ||
| 3j | – | 139 | – | ||
| 4j | – | 1000est | + | ||
| 5j | – | 25 | – | ||
| 6j | – | 30 | + | ||
| 7j | – | 189 | + | ||
| 8j | – | 98 | + | ||
| 9j | – | 14 | – | ||
| 10j | – | 5 | – | ||
| 11j | – | 8 | – | ||
| 12j | – | 545 | – | ||
| 13j | – | 129 | + | ||
| 14j | + | 78 | – | ||
| 15j | – | 38cal | + | ||
| 16j | – | 59 | – | ||
| 17j | – | 26 | + | ||
| 18j | + | 214 | – | ||
| 19j | – | 48 | – | ||
| 20j | + | 449 | – | ||
| 21j | + | 91 | + | ||
| 22j | – | 135 | – | ||
| 23j | + | 308 | – | ||
| 24j | + | 321 | – | ||
| 25j | – | 97 | – | ||
| 26j | – | 145 | – | ||
| 27j | – | 1000est | – | ||
| 28j | + | 359 | – | ||
| 29j | – | 229 | + | ||
| 30j | – | 17 | – | ||
| 31j | – | 64 | – | ||
| 32j | – | 5 | – |
% positive (# positive/total tested) 19% (6/32)
| Animal # | HEYM Culture Positive | ELISA +/- | IFNγ +/- | Fecal PCR +/- | Plaque IS900 qPCR +/- |
|---|---|---|---|---|---|
| 1c | – | – | – | + | |
| 2c | + | – | – | – | |
| 3c | – | – | – | – | |
| 4c | – | – | – | + | |
| 5c | – | – | – | + | |
| 6c | – | – | – | – | |
| 7c | – | – | – | – | |
| 8c | – | – | – | + | |
| 9c | + | – | – | – | |
| 10c | – | – | – | – | |
| 11c | + | – | + | – | |
| 12c | – | – | – | – | |
| 13c | – | – | – | – | |
| 14c | – | – | – | – | |
| 15c | – | – | – | – | |
| 16c | – | + | – | – | |
| 17c | – | – | – | – |
% positive (# positive/total tested) 18% (3/17) 0% (0/17) 12% (2/17) 0% (0/17) 35% (6/17)
In the six Johne animals with visible HEYM slope colonies, ELISA was positive in two, IFNγ was positive in five and all six of IS900 fecal PCR.
| Animal # | HEYM Culture + | ELISA +/- | IFNγ +/- | Fecal IS 900 PCR +/- | Phage / qPCR +/- |
|---|---|---|---|---|---|
| 1j | – | + | + | + | |
| 2j | – | + | – | + | |
| 3j | – | + | + | – | |
| 4j | – | – | – | + | |
| 5j | – | – | + | – | |
| 6j | – | – | – | – | |
| 7j | – | + | – | + | |
| 8j | – | + | – | – | |
| 9j | – | + | + | + | |
| 10j | + | + | + | + | |
| 11j | – | + | + | – | |
| 12j | – | + | + | – | |
| 13j | – | + | – | + | |
| 14j | + | – | + | + | |
| 15j | – | – | + | – | |
| 16j | – | + | + | – | |
| 17j | – | + | + | – | |
| 18j | + | – | + | + | |
| 19j | + | – | + | – | |
| 20j | + | – | + | + | |
| 21j | + | – | + | – | |
| 22j | + | – | + | – | |
| 23j | – | – | – | – | |
| 24j | – | – | – | – | |
| 25j | – | – | – | – | |
| 26j | – | – | – | – | |
| 27j | – | – | – | – | |
| 28j | + | – | – | – | |
| 29j | + | – | – | – | |
| 30j | + | – | – | – | |
| 31j | + | – | – | – | |
| 32j | + | – | – | – |
% positive (# positive/total tested) 19% (6/32) 31% (10/32) 69% (22/32) 75% (24/32) 72% (23/32)
Discussion:
The evolution in the detection and identification of mycobacteria has been progressive over more than 130 years. A time consuming and insensitive system has increasingly and rapidly incorporated modern, rapid and more sensitive techniques. In this study we have had an unparalleled opportunity to compare classical and modern techniques.
We clearly show that the modern, rapid techniques are more sensitive and specific than the classical methods. Additionally, the modern techniques increased sensitivity show that putatively negative Control animals may be infected. The majority of the infected control animals were without clinical symptoms. The implications of this asymptomatic colonization/cryptic infection have far reaching implications. It signifies that animals that are colonized but not clinically symptomatic can be identified early in the course of their exposure, before they are symptomatic and possibly amenable to early and possible less intrusive interventions. For example, it is well described that elevated vitamin D levels are associated with improved outcomes in many mycobacterial diseases.
In the Johne animals, all that were positive on HEYM had also been positive on fecal IS900 PCR. However, 17 animals that were fecal IS900 positive were not positive on HEYM: again, attesting to the lack of diagnostic sensitivity of Classical mycobacterial detection methods. In contrast, none of the Control animals were fecal PCR positive, whereas 5/17 Control animals were positive, attesting the sensitivity of the plaque/qPCR assay over Classical mycobacterial detection methods.
One of the most powerful aspects of the plaque/qPCR methodology is that it can only detect living mycobacteria. A viable mycobacterium is necessary for the mycobacteriophage to infect, replicate and burst releasing the DNA of the infected mycobacterium to be identified during the subsequent qPCR process. Thus, the concern that PCR is only detecting the DNA of dead mycobacteria DNA in milk will no longer be acceptable.
This technology has the potential to address a century long mycobacteriologist dilemma. Although Leprosy has been considered to be a mycobacterial infection since 1882, it has never fulfilled Kochs postulates. As an obligate intracellular parasite, because of massive genomic deletion, it cannot be grown in pure culture and therefore the second and fourth postulates cannot be met. The diagnosis of Leprosy is clinical. It is relatively simple with lepromatous leprosy but is clinical for tuberculoid leprosy, where the organism is not detectable. Multiple serum diagnostic tests to confirm an M. Leprae infection are proposed. However, the phage/qPCR technology discussed in this manuscript has the potential to identify living M. leprae and render null and void Koch’s unmet postulates.
Conclusions:
Clinically useful detection of viable MAP in blood is feasible within 48 hours using a rapid phage/qPCR assay. In contrast, although of intellectual and academic interest, attempting to culture mycobacteria from blood has no role in clinical practice. In a miniscule number of cases, it may be of use in mycobacterial nucleic acid analyses.
Funding: This study was performed at the James J. Peters Veterans Affairs Medical Center Bronx NY 10468. Funding was by the Bronx Veterans Affairs Research Foundation Inc (BVRF Inc.) Extramural funds were provided by C. Heffner.
Conflicts of Interests: Neither of the authors have conflicting interests. Neither the BVRF Inc. nor C. Heffner had any role in the design or performance of the studies, wrote the manuscript or made any decision to publish.
Acknowledgements We thank L. Su for laboratory assistance and all the authors who contributed to the published manuscript.
References
1. Hansen GHA. Indberetning til det norske medicinske Selskap i Christiania om en med Understottelse av Selkabet foretage Reise for at anstille Undersogelser angaaende Spedalskhedens Aarsager. Norsk Magazin for Laegevidenskaben 1874;4:1–88. (In Norwegian).
2. Hansen GHA. Bacillus leprae. Quarterly Journal of the Microscopical Science 1880;20:92–102.
3. Koch R. Die Aetiologie der Tuberculose. Berliner Klinische Wochenschrift 1882;19:221–230. (In German).
4. Koch R. Die Aetiologie der Tuberkulose. Mittbeilungen aus dem Kaiserlichen Gesundbeitsamte 1884;2:1–88. (In German).
5. Dulberger CL, Rubin EJ, Boutte CC. The mycobacterial cell envelope — a moving target. Nature Reviews Microbiology 2020;18(1):47–59. Doi: 10.1038/s41579-019-0273-7.
6. Marrakchi H, Lanéelle M-A, Daffé M. Mycolic Acids: Structures, Biosynthesis, and Beyond. Chemistry & Biology 2014;21(1):67–85. Doi: https://doi.org/10.1016/j.chembiol.2013.11.011.
7. Ziehl F. Zur Farbung des Tuberkelbacillus. Dtsch, med Wschr, 1882;8:451.
8. Neelsen FCA. Ein casuistischer Bietrag zur Lehre von der Tuberkulose. Zbl med Wiss 1883;21(497-501).
9. Lowenstein E. Die Zachtung der Tubekelba zillen aus dem stramenden Blute. Zentralbl Bakteriol Parasitenkd Infektionskr Hyg Abt I Orig 1931;1:120–127.
10. Lowenstein E. Der Kulturelle Nachweis von Tuberkelbakterien in Milch auf Malachitgrun Einarboden. Ann Inst Pasteur 1933;50:161.
11. Jensen KA. Rinzuchtung und Typenbestim mung von Tuberkelbazillentammen. Zentralbl Bakteriol Parasitenkd Infektionskr Hyg Abt I Orig 1932;125.
12. Jensen KA. Reinzüchtung und Typenbestimmung von Tuberkelbazillenstämmen: Ein
Vereinfachung der Methoden für die Praxis. Zentralbl Bakt 1932;125:222–239.
13. Knox R. SEMI-SOLID AGAR MEDIA FOR RAPID CULTURE OF TUBERCLE BACILLI. The Lancet 1955;266(6881):110–112. Doi: 10.1016/S0140-6736(55)92114-0.
14. Middlebrook G, Cohn ML. Bacteriology of tuberculosis: laboratory methods. Am J Public Health Nations Health 1958;48(7):844–53. Doi: 10.2105/ajph.48.7.844.
15. Middlebrook G, Reggiardo Z, Tigertt WD. Automatable radiometric detection of growth of Mycobacterium tuberculosis in selective media. The American review of respiratory disease 1977;115(6):1066–9. (http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=122228).
16. Greenstein RJ, Su L, Haroutunian V, Shahidi A, Brown ST. On the Action of Methotrexate and 6-Mercaptopurine on M. avium subspecies paratuberculosis. PLoS ONE 2007;2(1):e161. (In eng). Doi: 10.1371/journal.pone.0000161.
17. Greenstein RJ, Su L, Shahidi A, Brown WD, Clifford A, Brown ST. Unanticipated Mycobacterium tuberculosis complex culture inhibition by immune modulators, immune suppressants, a growth enhancer, and vitamins A and D: clinical implications. Int J Infect Dis 2014;26C:37–43. Doi: 10.1016/j.ijid.2014.01.026.
18. Shin SJ, Collins MT. Thiopurine drugs azathioprine and 6-mercaptopurine inhibit Mycobacterium paratuberculosis growth in vitro. Antimicrob Agents Chemother 2008;52(2):418–26. (Research Support, Non-U.S. Gov’t). Doi: 10.1128/AAC.00678-07.
19. Krishnan MY, Manning EJ, Collins MT. Effects of interactions of antibacterial drugs with each other and with 6-mercaptopurine on in vitro growth of Mycobacterium avium subspecies paratuberculosis. J Antimicrob Chemother 2009;64(5):1018–23. (http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=19759042).
20. Chiodini RJ, Van Kruiningen HJ, Merkal RS, Thayer Jr. WR, Coutu JA. Characteristics of an unclassified Mycobacterium species isolated from patients with Crohn’s disease. J Clin Microbiol 1984;20(5):966–971.
21. Wilson SM, al-Suwaidi Z, McNerney R, Porter J, Drobniewski F. Evaluation of a new rapid bacteriophage-based method for the drug susceptibility testing of Mycobacterium tuberculosis. Nat Med 1997;3(4):465–8. Doi: 10.1038/nm0497-465.
22. Jacobs WR, Jr., Barletta RG, Udani R, et al. Rapid assessment of drug susceptibilities of Mycobacterium tuberculosis by means of luciferase reporter phages. Science 1993;260(5109):819–22. Doi: 10.1126/science.8484123.
23. Carrière C, Riska PF, Zimhony O, et al. Conditionally replicating luciferase reporter phages: improved sensitivity for rapid detection and assessment of drug susceptibility of Mycobacterium tuberculosis. Journal of Clinical Microbiology 1997;35(12):3232–3239. Doi: doi:10.1128/jcm.35.12.3232-3239.1997.
24. Swift BM, Rees CE. Detecting mycobacteria in cattle blood. Vet Rec 2013;173(21):522–3. Doi: 10.1136/vr.f7067.
25. Foddai A, Elliott CT, Grant IR. Optimization of a phage amplification assay to permit accurate enumeration of viable Mycobacterium avium subsp. paratuberculosis cells. Appl Environ Microbiol 2009;75(12):3896–902. (http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=19395561).
26. Stewart LD, Foddai A, Elliott CT, Grant IR. Development of a novel phage-mediated immunoassay for the rapid detection of viable Mycobacterium avium subsp. paratuberculosis. J Appl Microbiol 2013;115(3):808–17. (In eng). Doi: 10.1111/jam.12275.
27. Greenstein RJ, Su L, Grant IR, et al. Comparison of a mycobacterial phage assay to detect viable Mycobacterium avium subspecies paratuberculosis with standard diagnostic modalities in cattle with naturally infected Johne disease. Gut pathogens 2021;13(1):30. Doi: 10.1186/s13099-021-00425-5.
28. Pozzato N, Gwozdz J, Gastaldelli M, Capello K, Dal Ben C, Stefani E. Evaluation of a rapid and inexpensive liquid culture system for the detection of Mycobacterium avium subsp. paratuberculosis in bovine faeces. Journal of microbiological methods 2011;84(3):413–7. Doi: 10.1016/j.mimet.2011.01.019.
29. Twort FW, Ingram GLY. Zbl Bakt 1914;73(277).
30. Francis J, Macturk HM, Madinaveitia J, Snow GA. Mycobactin, a growth factor for Mycobacterium johnei. I. Isolation from Mycobacterium phlei. Biochem J 1953;55(4):596–607. (http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=13115341).
31. Saleeby CW. The Action of Sunlight. Nature 1922;109(2723):11–12. Doi: 10.1038/109011b0.
32. Bruce D, Ooi JH, Yu S, Cantorna MT. Vitamin D and host resistance to infection? Putting the cart in front of the horse. Exp Biol Med (Maywood) 2010;235(8): 921–7. (http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=20660091).
33. Salahuddin N, Ali F, Hasan Z, Rao N, Aqeel M, Mahmood F. Vitamin D accelerates clinical recovery from tuberculosis: results of the SUCCINCT Study [Supplementary Cholecalciferol in recovery from tuberculosis]. A randomized, placebo-controlled, clinical trial of vitamin D supplementation in patients with pulmonary tuberculosis’. BMC Infectious Diseases 2013;13(1):22. Doi: 10.1186/1471-2334-13-22.
34. Stewart-Tull DES. Mycobacterium leprae – The bacteriologist’s enigma. In: Ratledge C, Stanford J, eds. The Biology of the Mycobacteria, Volume 1: Physiology, Identification, and Classification. 1 ed. New York: Academic Press; 1982:273–307.
35. Cole ST, Eiglmeier K, Parkhill J, et al. Massive gene decay in the leprosy bacillus. Nature 2001;409(6823):1007–11. (Research Support, Non-U.S. Gov’t) (In eng). Doi: 10.1038/35059006.
36. Ridley DS, Jopling WH. A classification of leprosy for research purposes. Lepr Rev 1962;33:119–28. (http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=14492126).
37. Ridley DS, Jopling WH. Classification of leprosy according to immunity – a five-group system. Int J Lepr Other Mycobact Dis 1966;34:255–273.
38. Lima MIS, Correa MBC, Moraes E, et al. HSP60 mimetic peptides from Mycobacterium leprae as new antigens for immunodiagnosis of Leprosy. AMB Express 2023;13(1):120. Doi: 10.1186/s13568-023-01625-9.
39. Alban SM, de Moura JF, Thomaz-Soccol V, et al. Phage display and synthetic peptides as promising biotechnological tools for the serological diagnosis of leprosy. PLoS One 2014;9(8):e106222. (In eng). Doi: 10.1371/journal.pone.0106222.
40. Sengupta U. Recent Laboratory Advances in Diagnostics and Monitoring Response to Treatment in Leprosy. Indian Dermatol Online J 2019;10(2):106–114. (In eng). Doi: 10.4103/idoj.IDOJ_260_18

