COVID Post-Infection Therapy: Methods and Mechanisms
COVID Post-Infection Therapy: A Survey of Methods and Mechanisms
Richard R. Zito, Ph.D.1
- Research LLC; Tucson, Arizona, USA
OPEN ACCESS
PUBLISHED: 30 October 2024
CITATION: Zito, R.R., 2024. COVID Post-Infection Therapy: A Survey of Methods and Mechanisms. Medical Research Archives, [online] 12(10).
https://doi.org/10.18103/mra.v1i210.5681
COPYRIGHT: © 2024 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v1i210.5681
ISSN 2375-1924
ABSTRACT
There has been a significant reduction in the number of COVID cases world-wide since the pandemic peaked in 2022. However, new strains have developed in the two years following the peak that are significantly different genetically from their predecessors. This, combined with public apathy concerning vaccination and other preventative measures, has led to periodic outbreaks. The problem has been particularly troublesome in the U.S. So, what options are available to an individual who has developed a breakthrough infection, or has become COVID-19 positive due to negligence? This publication covers the mechanisms and side effects of COVID antiviral and anti-inflammatory drugs, interleukins, interferons, and monoclonal antibodies. The chemistry of these diverse therapeutic agents is complex, but their pharmacology, biochemistry, and even their molecular structures, are often related. These relationships will be explained, and the associated terminology demystified.
Keywords: COVID-19, antiviral drugs, anti-inflammatory drugs, interleukins, interferons, monoclonal antibodies
Background and Introduction
Globally, the number of COVID deaths and active cases has certainly declined since the “Omicron peak” in 2021 — an encouraging sign. However, here in the U.S. it is difficult to know the actual number of active COVID cases because many people are treating themselves at home when their symptoms are minor,²,³ and many COVID records are no longer being kept.⁴ Nevertheless, in spite of some eye-popping estimates,²,³ sewage samples do not seem to indicate that viral concentrations are greater than they were during the omicron peak of the pandemic.⁵,⁶,⁷ Furthermore, the number of active cases, and deaths, are bound to be a lot less due to immunity imparted to the population by vaccination and wild-type infections. However, the U.S. has seen repeated COVID outbreaks. For example, in September 2023, there has been a significant resurgence of COVID due to strains EG.5 (21.5%), FL.1.5.1 (14.5%), XBB.1.16.6 (9.2%), and others.⁸,⁹,¹⁰ These PANGO designations have been explained in the literature.¹¹ How do these outbreaks occur? In part, it is because Americans have waited two years before updating their new vaccine. It may be that COVID strains are again evolving out of the range of the protection offered by current vaccinations. Furthermore, the decay of immunity offered by wild-type infection, combined with resistance to public safety measures (e.g. unwillingness to vaccinate, wear masks, follow social distancing rules, and laxity about hand washing and sanitizing), have resulted in a more susceptible population.
So, what options are available to someone who gets an active case of COVID, either through negligence or as a breakthrough infection? Many therapies are available, none are 100% effective, and all have side effects. How do these drugs work? Is there any relationship between them? What are their limitations? How are they administered? How do they affect the human body? These are the questions that will be answered in this publication. First, however, it is necessary to discuss a few basic properties of vaccines, antibiotics and antiviral drugs.
Most Vaccines Concentrate on Developing Immunity to the Spike (S) Protein
Examination of the coronavirus model in Figure 1 shows that each virion has an outer bilayer (double layer) of lipid molecules (e.g. fatty acids or cholesterol), called an envelope, that looks like the lipid bilayer that makes up the outer membrane of any animal cell. All of a human cell’s biochemical machinery is enclosed within this membrane, and it is necessary for cellular life. Therefore, it is not possible to develop antibodies against the virion’s envelope because it looks “self”, and you can’t develop antibodies against yourself unless you have an autoimmune disease.¹² Embedded within the envelope are proteins called trans-membrane proteins that extend from the surface of the outer lipid layer to the surface of the inner lipid layer. For the coronavirus, two important trans-membrane proteins are M (for membrane) and H-E (for hemagglutinin esterase). The latter of these is a dangerous protein associated with blood clots in patients. It is the strange shape of foreign proteins such as these that trigger an immune response. However, neither of these proteins protrudes above the outer lipid layer, and they can only be seen “end-on” by attacking cells of the immune system. Only the spike, or S-protein, is accessible to immune system B-cells. Furthermore, the contents of the virion’s envelope, such as the nucleocapsid protein, are invisible to impinging immune system cells and antibodies. Therefore, trying to develop “whole virus” antibodies is futile. Only antibodies developed against S will be effective in preventing disease!
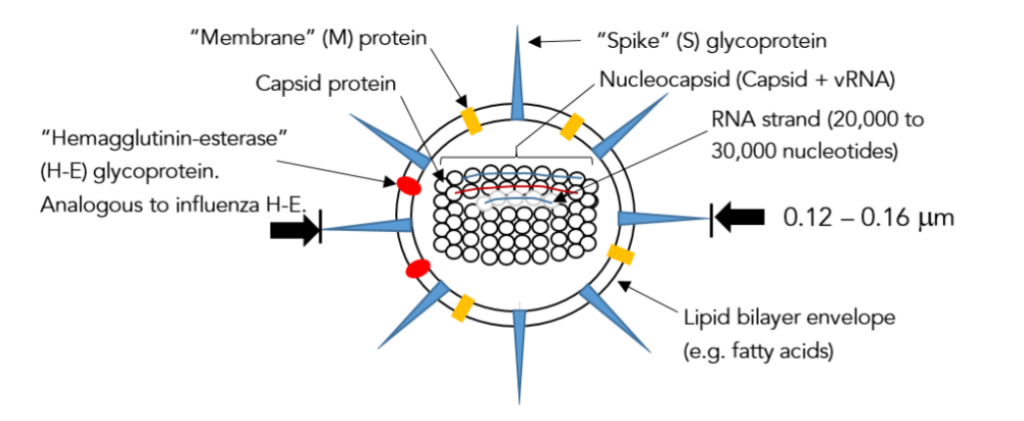
Figure 1 – Schematic of an enveloped coronavirus particle (a virion).
However, as was explained in “The Delta Variant” and “The Omicron Strain”,¹³,¹¹ evolutionary pressures have provided the coronavirus with a partial defense. Some of the amino acids that make up S are capable of binding sugar molecules, or even groups of sugar molecules. Since sugar is not normally recognized by our immune system as something dangerous, the coronavirus has disguised the most important and vulnerable part of its anatomy. However, the disguise is imperfect because not every amino acid on the surface of the S-protein binds sugar. That is why the ones that do, have a preference for groups of sugar molecules that are wide enough to cover neighboring amino acids that may not. Worse still, mutations can exchange amino acids that don’t bind sugar for ones that do. This is what happened in δ (mutations L452R and E484Q).¹³ However, the human immune system has an answer to this conundrum. If the sugar screens become too ornate, the immune system will eventually recognize them as “not self”, but by then the infection may be well underway and possibly fatal to the host.
So Many Antibiotics, So Few Antiviral Drugs
Antibiotics, as everyone knows, are effective against bacteria — technically, members of a separate kingdom of single-celled organisms called Protista that include animal-like amoeba and paramecia, plant-like bacteria, and some organisms like the euglena that have both plant and animal characteristics.¹⁴,¹⁵ It is the thick, fairly rigid, cell wall of bacteria, made of a polymer of sugars and amino acids — a very different structure from the fatty lipid outer cell membrane of an animal cell — that makes them so plant-like, and therein lies their Achilles’ heel. The antibiotic penicillin interferes with the biochemical reactions necessary to make bacterial cell walls¹⁵ — chemical reactions that have no counterpart in animal metabolism. Since bacterial cells must continue to synthesize cell wall in order to grow and divide, inhibition of this process provides a completely specific way to control the proliferation of bacteria without harming the human or animal host. Penicillin is often considered an ideal antibiotic. Unfortunately, bacteria can acquire resistance to penicillin. However, the cephalosporins, bacitracin, and vancomycin work in a similar way.¹⁵ Hence, there is a continual hunt for new antibiotics.
By comparison, antiviral drugs are harder to come by than antibiotics because enveloped viruses look so much like human cells to the immune system. The reader might think that a bacterial cell is closer to a human cell than a virus, and in many ways that is certainly true. However, on the surface, the lipid envelope of a coronavirus looks much like the lipid outer membrane of a human cell. Therefore, most things that you can think of that would destroy a virus would also damage human cells.
corona virion will also destroy its human host cell. For example, detergent will destroy the coronavirus envelope, thereby washing away the all-important spike (S) protein that is so necessary for the coronavirus to bind to a target cell. However, the same detergent will also disrupt the host cell’s outer membrane, thereby killing it! Detergent is useful outside the human body because interior cells are protected by the dead keratinocytes of the epidermis.¹⁶ Because of the surface similarities between an enveloped virus and a human cell, a more involved strategy must be employed to kill a virus than to kill a bacterium. Sooner or later during viral replication, the coronavirus must expose its chemically reactive naked RNA to the cellular environment.¹⁷ It is at this point in its life cycle that the coronavirus is most vulnerable because there are compounds that are capable of binding to viral RNA (vRNA). Such compounds are potentially antivirals because they might block the replication of vRNA by the viral protein replicase! Figure 2 schematically depicts replicase as a disk with a hole in it; a shape that is topologically equivalent to a torus, or “doughnut”. If one viral RNA strand can fit through the hole, two will emerge. However, if a molecule with the right size and shape is bound to the vRNA, passage through the hole can be effectively blocked so that transcription (copying of vRNA) cannot take place. Such a molecule is an antiviral pharmaceutical. In order to understand exactly how these therapeutics bind to vRNA it is necessary to understand nucleic acids.
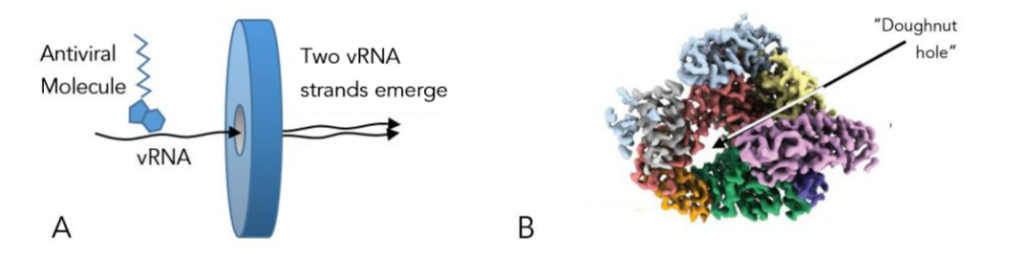
Figure 2 – A) Schematic of replicase in action. B) Constant electron density surface of the COVID replicase protein. Sub-units (sub-proteins) that make up the whole replicase protein are color coded. Modified from Yin et al.¹⁸
Nucleic Acids (RNA and DNA)
RNA, and each of the two strands of DNA, are made up of three building blocks. Phosphate (PO₄²⁻), and a five carbon sugar called ribose (“rose means sugar), make up the backbone of these information molecules of inheritance. And, bonded to the ribose of this backbone, like ribs, are molecules called bases. DNA uses four bases (Adenine [A], Guanine [G], Cytosine [C], and Thymine [T]). RNA uses A, G, and C, but not T. Instead, RNA uses Uracil (U) in place of T. These bases are depicted in Figure 3. Notice that they fall into two groups due to similarity of structure.
Replicase
Although the reader is probably beginning to get an idea of how antivirals work, the author must resist the temptation to get side-tracked or jump ahead too fast. A deeper understanding of replicase (short for RNA dependent RNA polymerase) needs to be had. Uninfected human cells do not make replicase. When a cell has to make some needed protein, it makes an RNA copy of one gene on a chromosome — a single gigantic molecule of DNA that acts as a permanent library of information from which no “books” (genes) can ever be removed. It makes this copy using an enzymatic macromolecular copying machine called polymerase (short for DNA dependent RNA polymerase). Polymerase (a cubical molecule) is very different from replicase (a torus). So, where does replicase come from? It is coded for on the vRNA of an invading coronavirus. Once a cell is infected with vRNA, that cell’s own ribosomes are “hijacked” to manufacture the protein replicase.¹³ Replicase is typically a gigantic protein molecule.
In fact, so gigantic that it is made up of 16 subunits,¹⁵ where each subunit is a complete protein in its own right. The subunits of replicase have shapes that fit together and are bound by Van der Waals forces, hydrogen bonding, and chemical bonds between amino acids of opposing subunits. One of these 16 subunits is very special indeed, it is called helicase,¹⁵ and its role in copying the vRNA genome will be discussed presently. Later, another subunit called nsp5 (non-structural protein 5) will be of interest. For now, the author wishes to direct the reader’s attention to Figure 4.
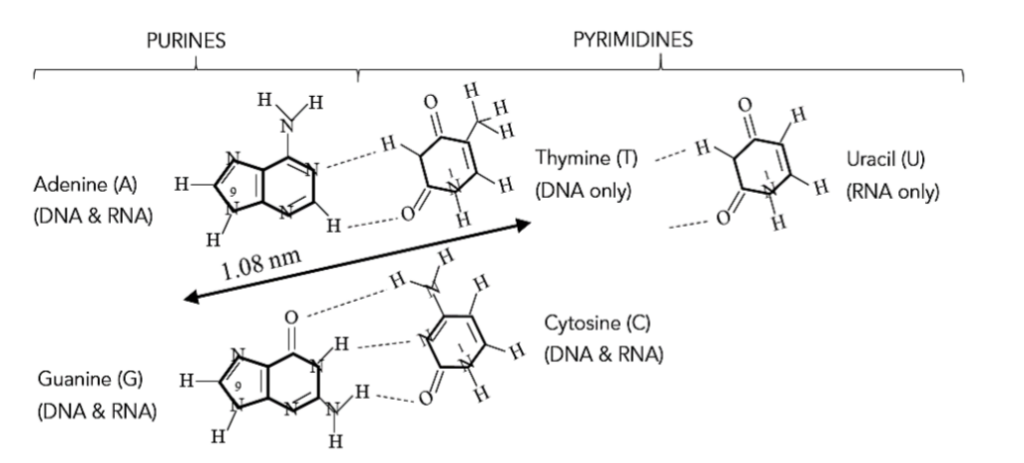
Figure 3 – Purines and pyrimidines.¹⁵ O is an oxygen atom, N is a nitrogen atom, and H is a hydrogen atom. Carbon atoms reside at unlabeled vertices. The numbers refer to the atomic position in the structures shown. All purines attach to the ribose sugar of a nucleic acid backbone at the #9 position with the release of a water molecule, while pyrimidines attach at the #1 position. The dotted lines show which atoms are involved in pyrimidine-purine bonding. These are not true chemical bonds, but weak hydrogen bonds so that the individual strands of double stranded DNA can be readily pulled apart when necessary.
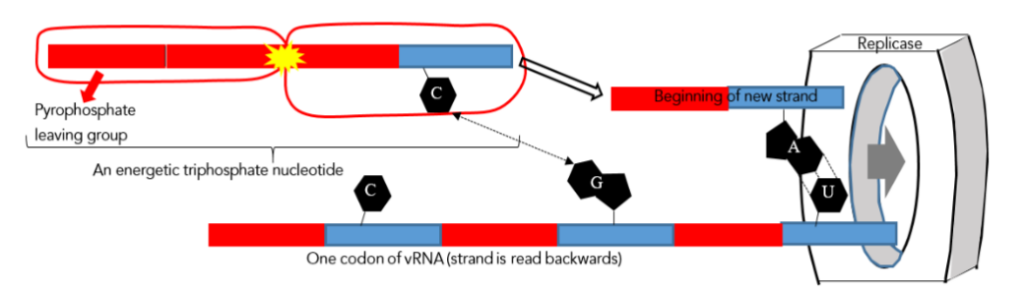
Figure 4 – A model of how a vRNA strand is copied by replicase. Each blue bar represents a molecule of ribose, and each red bar is a phosphate (PO₄²⁻) group. The ribose and phosphate are alternately bonded together chemically to form a nucleic acid “backbone”. Attached to each ribose is a base (black), five are shown. The yellow flash indicates the release of energy when a pyrophosphate group separates from a nucleotide triphosphate. All of the actions in this diagram take place within the replicase “doughnut hole”, which has been moved to the right so that the reader can clearly see each step.
A triplet of bases is called a codon, or genetic name. For example, CGU (as well as CGC, CGA, and CGG) are all genetic names for the amino acid arginine, one building block that a ribosome might use to construct a protein.¹⁵ In Figure 4, by the pyrimidine-purine bonding rule of the previous section, it is clear that adenine (A) can bond to uracil (U) in the macromolecular machine called replicase. While C bonds to G, and A to G. What happens inside replicase is that a double stranded RNA is created. Now, here is where helicase comes into play. Helicase, as its name implies, is a protein that pries apart (unwinds) this nascent double stranded helical RNA molecule. It is like a sailor’s pick (Marline spike) that is used to pry apart a jammed knot in a rope. So, what actually emerges from the replicase is the original vRNA strand plus something called a complementary strand. The complementary strand isn’t directly useful to the virus because its coding is all wrong. “A” emerges first, then C, then G, and ACG (that is how a protein synthesizing ribosome would read it, not the other way around) is one of the codons (genetic names) for the amino acid threonine¹⁵ – completely wrong. So, how do we get a duplicate vRNA like the one that was originally inserted into replicase? The answer is simplicity itself. When the complementary strand enters the cytosol (semi-fluid interior of the cell) it will eventually meet replicase again, and a second pass through this machinery will produce the desired vRNA clone (bearing mutations). Any compound that interferes with the operation of replicase, but not the distinctly different human polymerase, will be a useful antiviral agent. Two last questions must be answered before the mechanism of antiviral drugs can be completely understood.
First, where do the purine and pyrimidine bases come from? They come from food! Digestive enzymes called nucleases break food down into phosphate/ribose/base combinations called nucleotides. Several years ago an amusing survey was conducted in which people were asked if they would ever eat DNA. Most Americans said they would never eat such a thing. Fewer Europeans were fooled by this question. Presumably this says something about our respective educational systems!
Second, where does the energy come from to power chemical reactions like the extension of nucleic acid strands? If there is one thing that science teaches us, it is that there is no “free lunch” in this universe. Referring back to Figure 4, growth is not just a matter of hydrogen bonding pure bases to the vRNA strand. Instead each added base must carry with it a ribose and a phosphate to form the “backbone” of the complimentary strand.¹⁵ Where does the energy come from to form the new bonds? Each phosphate/ribose/base nucleotide that comes from food is modified to a triphosphate/ribose/base. This might seem like a trivial modification, but it is actually seminal, because the loss of two of these phosphate groups (called a pyrophosphate) releases a great deal of energy for new bond formation. Yes, but how did the monophosphate nucleotide from food get energized into a triphosphate compound in the first place? It happens when the monophosphate nucleotide interacts with an energy molecule called ATP (adenosine triphosphate) and gains a pyrophosphate group. ATP is the energy currency of the cell, and stores the energy released from the oxidation of dietary sugars. If this whole game with molecular energy seems confusing, don’t be intimidated. The bottom line is that food is the ultimate source of both the raw materials, as well as the energy, needed to build nucleic acids (RNA and DNA).
Remdesivir and Other Antiviral Pharmaceuticals
To date, remdesivir (trade name Veklury) is the only antiviral pharmaceutical approved by the U.S. FDA (Food and Drug Administration) without qualification for the treatment of COVID infections. Therefore, this is the drug that will be discussed in most detail. The structure of remdesivir is shown in Figure 5. Notice the purine head of this complex molecule. It looks more like adenine, rather than guanine, because it lacks an oxygen atom on the hexagonal ring. In fact, this head is called pseudo-adenine, or an adenine analogue. It is thought that the triphosphate version of remdesivir, naturally enough called remdesivir triphosphate and formed by reaction with ATP (Figure 5), substitutes for true triphosphate/ribose/adenine during vRNA copying.²⁰ This results in remdesivir (naturally a monophosphate) being incorporated into the growing nascent (new) RNA strand. The nascent strand can be either complementary, or the final clone. In either case, remdesivir sometimes wins against the true adenine nucleotide in this game of genetic “musical chairs” — but not always. That is why remdesivir is not 100% effective; although its use produces earlier clinical improvement (18 vs. 23 days), and reduces the 28-day mortality (11% vs. 15%), in adult patients hospitalized with severe COVID, when those patients were treated within 10 days of symptoms as compared with a placebo.²¹
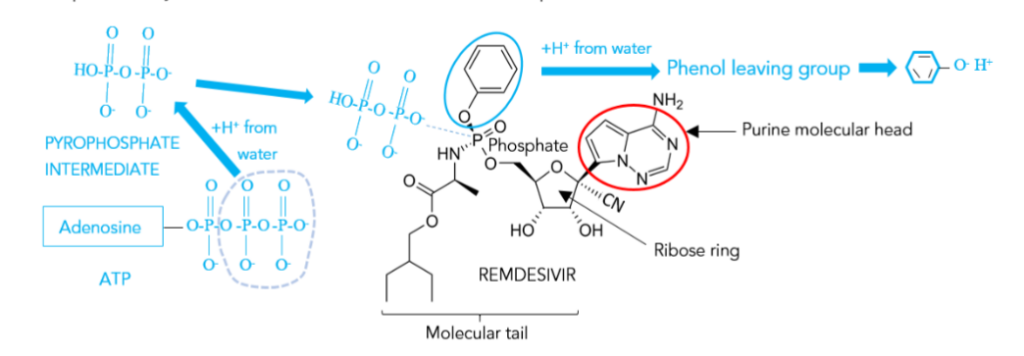
Figure 5 – The molecular structure of remdesivir (black), and a simplified model of how it reacts with cellular ATP (blue) to become a triphosphate. Ultimately, phenol must be released as a waste product (leaving group). Each unlabeled vertex is occupied by a carbon atom. Solid black wedges indicate bonds coming out of the plane of the paper towards the reader. Wedges made of dashes indicate bonds extending behind the paper plane. Common H-C bonds are not shown.
The cryo-electron microscopy of Taylor and Johnson shows what happens on the molecular level when remdesivir does win (Figure 6).²² Remdesivir seems to plug the “doughnut hole” of replicase, thereby causing the premature termination of transcription (copying of the viral genome). It is one of the astounding triumphs of modern science that instruments are now available that are capable of imaging a single protein molecule and can automatically produce a structural cartoon output (e.g. Fig. 6) from an electron density map like that in Figure 2B — work that used to take months or even years! Figure 6 begs the question “Exactly how does a molecule that looks like the one in Figure 5 plug the hole of the replicase torus?” One theory is that the cyanide group (-CN) attached to the ribose ring in Figure 5 interacts with the amino acid “S861” inside the “doughnut hole” of replicase. The “S” stands for serine, the 861st amino acid of one of the 16 subunits of replicase.²³
At this point the reader is probably wondering why remdesivir can’t be used for other viral infections since its mechanism of action is fairly general. Well, as a matter of fact, remdesivir was originally used as a therapeutic agent to treat Ebola virus infections. It was later repurposed as a COVID therapy, and is effective against all strains from α to omicron. Table 1 is a list of antiviral drugs, the diseases they are effective against, and their mechanism of action.
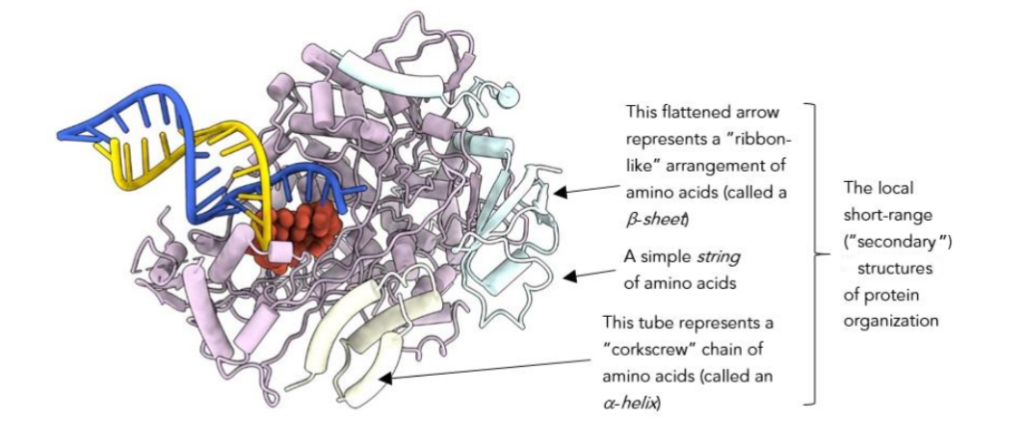
Figure 6 – Cartoon of remdesivir in action (oblique rear view). The red part of the cartoon is remdesivir plugging the hole of the replicase torus (in light blue). The emergent yellow strand is a complementary copy of whatever was fed into this macromolecular copying machine. If the input strand (dark blue) emerging from the exit end of the replicase was itself a complementary strand of the original vRNA, then the yellow output will be a copy of the original vRNA. In this cartoon transcription (copying) has been terminated prematurely by remdesivir. Figure from University of Texas modified by the author.²²
Table 1 – Antivirals and their uses.²⁰ The “Drug” column lists the generic name of antiviral drugs available in the U.S. as of 2023. These names end in “-vir”, for “virus”. The trade names of these drugs follow in parenthesis, some of which are Canadian. Some of these drugs are FDA approved. Others only have Emergency Use Authorization (EUA). The “Disease” column lists the diseases typically treated by the corresponding antiviral in the first column. The last column describes the “Mechanisms of Action” of the drugs in column 1. Note that viruses can have an RNA or DNA genome. RNA viruses (like coronavirus) are transcribed (copied) by replicase, while DNA viruses employ a polymerase.
DRUG — DISEASE — MECHANISM OF ACTION
acyclovir (Sitavig, Zovirax) — herpes (cold sore, genital, neonatal), shingles, chickenpox, herpes simplex encephalitis — A purine analogue that interferes with viral DNA synthesis.
baloxavir (Xofluza) — influenza — Endonuclease inhibitor.*
entecavir (Baraclude) — chronic hepatitis B — A guanine analogue that interferes with viral RNA replicase.
famciclovir — shingles, herpes (cold sores, genital), mucocutaneous herpes in HIV patients — Inhibits viral DNA synthesis in herpes infected cells only.
foscarnet (Foscavir) — cytomegalovirus infection (CMV), retinitis in HIV patients, treatment of acyclovir-resistant mucocutaneous herpes in immunocompromised patients — Prevents viral replication by inhibiting viral DNA polymerase and reverse transcriptase.²⁵
ganciclovir (Cytovene) — CMV infection in immunocompromised patients, HIV patients, congenital infections in neonates — Inhibits viral DNA polymerase.
lamivudine (3TC, Epivir, Epivir-HBV, Heptovir) — HIV, chronic hepatitis B — Inhibits viral DNA synthesis by inhibiting the enzyme reverse transcriptase.²⁵
molnupiravir (no trade name) — mild to moderate COVID-19 (EUA) — Nucleotide analogue.
nirmatrelvir/ritonavir (both drugs administered together) (Paxlovid) — mild to moderate COVID-19 (EUA) — Protease inhibitor.**
oseltamivir (Tamiflu) — influenza — Viral neuraminidase inhibitor.***
remdesivir (no trade name) — COVID-19 (FDA approved), ebola virus infection — Adenine analogue that interferes with RNA replicase.
valACYclovir (Valtrex) — shingles, genital herpes, chickenpox, cold sores — Interferes with viral DNA synthesis.
valGANciclovir (Valcyte) — CMV retinitis in AIDS patients, prevention of CMV in kidney/pancreas and heart transplant patients at risk — Inhibits viral DNA polymerase.
* Ribosomes, the cellular macromolecular machines that produce proteins from RNA instructions, require that the RNA be capped. That is to say, the RNA instructions must begin with a certain recognizable sequence of nucleotides. This is true regardless of whether the ribosome is translating the hosts own RNA, or foreign vRNA. Most vRNA comes with its own cap. However, some viruses, like influenza, steal a cap from a host cell mRNA. The process is called cap-snatching, and utilizes an enzyme (catalyst) called endonuclease to cleave off the necessary sequence. If endonuclease is inhibited, so is viral replication of influenza.²⁴
** A protease is a protein molecule that cuts up other proteins. Some viruses require protease activity for the assembly of new infectious daughter virions.²⁴,²⁶ It’s just one more way in which viruses hi-jack cellular machinery for their own purposes.
*** The genome of the influenza virus codes for a protein enzyme called neuraminidase that facilitates release of daughter virions from infected cells, as well as viral entry into healthy cells.²⁷ Therefore, inhibition of neuraminidase suppresses cell-to-cell release and spread of the virus. Interested readers can study a molecular cartoon of neuraminidase in Molecular Biology of the Cell.²⁸
Finally, the clinical aspects of remdesivir need to be addressed.²⁰ It’s harder than you might think to get a drug into a cell. However, remdesivir has a good disguise. The combination of purine attached to a ribose ring followed by a phosphate group (or triphosphate group) makes remdesivir look like a nucleotide (Figure 5). The immune system considers it to be “self”, and cells will let it in as a food derived nutrient. Each part of the remdesivir molecule has some specific function. It is truly a “designer drug”. Welcome to modern pharmacy!
Remdesivir is supplied as a powder that can be prepared for injection by dissolving 100 mg of remdesivir in 20 ml of sterile water. This procedure prevents the drug from being damaged by digestive acids and enzymes. A remdesivir solution is stable for 4 hours at room temperature, or 24 hours if refrigerated. Therefore, like COVID vaccines, there is a risk of drug deterioration due to mishandling. In the U.S., for adults and children over 12 whose weight is over 40 kg, 200 mg of remdesivir is administered on the first day of treatment. Then 100 mg (once daily) are administered starting on day 2. Treatment continues for 10 days if the patient requires invasive mechanical ventilation, but only for 5 days if no ventilation is required. If the latter patients do not improve in 5 days, the treatment may be extended for another 5 days. So, quite a lot of solution is required. Remdesivir has a half-life of 20 hours in the human body. Most of it is excreted in the urine (74%). Another 18% is found in feces. Remdesivir, like any drug, can have side effects. These include hyperglycemia, nausea, constipation, hepatic (liver) impairment, acute kidney injury, anemia, respiratory failure, hypersensitivity reactions (including anaphylaxis), fever, and infusion reactions. There is a popular rumor circulating among anti-vaxxers, they believe that therapeutics are less dangerous than vaccine. Furthermore, they believe that therapy is 100% effective and that they can afford to forgo vaccination. All of these beliefs are completely false! Therapy is slow, potentially dangerous, and the outcome is not guaranteed. Everyone should use a layered defense against COVID. Physical barriers (masks, distancing, hand-washing, etc.) are the first line of defense. Next comes vaccination. The last line of defense is therapeutics. After that, you are on your own, and you must depend on the integrity of your own immune system and your physical condition to keep you alive. There is one last point that must be mentioned. The pharmacological activity of remdesivir is down-regulated by concurrent use of chloroquine and hydroxychloroquine. Therefore, these last drugs, once thought to be useful COVID therapeutics, should be avoided when taking remdesivir.
Molnupiravir (Merck/Ridgeback COVID Chemotherapy) and Pfizer Paxlovid (Both EUA Only Drugs)
When molnupiravir was introduced it created quite a sensation in the popular press.²⁷ Technically, it is a pyrimidine analogue (instead of a purine analogue like remdesivir; see Table 1). The molecular structure of this drug is shown in Figure 7A. It has a pseudo-cytosine head (see Figure 3), and a tail containing a ribose sugar ring, but no phosphate group. The result of this deletion is that molnupiravir, if and when it is able to substitute for the natural cytosine nucleotide and bind to a guanine nucleotide via the pyrimidine-purine binding rule, induces a catastrophic number of RNA copying errors (viral error catastrophe), as might be expected. Originally intended as an influenza therapeutic, it was later repurposed for COVID-19 treatment.
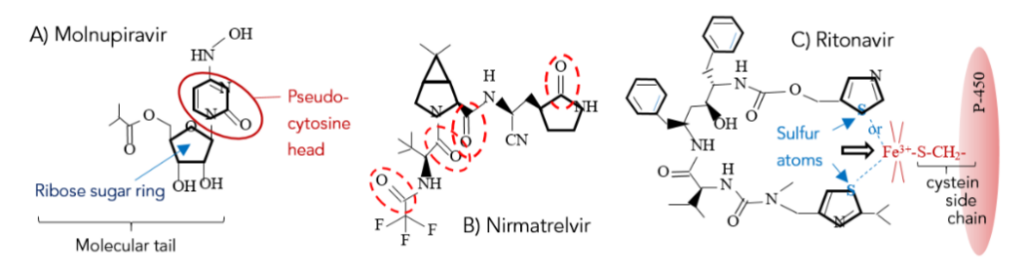
Figure 7 – A) The molecular structure of molnupiravir (C₁₃H₁₉N₃O₇; Gram Molecular Weight (GMW) = 329.309 g/mol).³¹ B) Nirmatrelvir structure (C₂₃H₃₂F₃N₅O₄; GMW = 499.535 g/mol).³² It is one of this molecule’s chain of chemically active C=O (carbonyl) bonds (encircled in red dashes) that seems to react with cysteine, an amino acid in the replicase nsp5 subunit, thereby inactivating its protease activity.³³ C) Ritonavir (C₃₇H₄₈N₆O₅S₂; GMW = 720.95 g/mol).³⁴ Note the sulfur atom (blue) at the end of each “horn” of this “horseshoe” shaped molecular structure.
conformation (one of several possible configurations), forming a kind of “pincer”. Although mechanisms are being debated,³⁵ either of ritonavir’s two sulfur atom has the potential to bond to the Fe³⁺ ion in the P-450 binding site (red).¹⁵,³⁶ P-450 is a molecule that is destructive to nirmatrelvir, and once its iron ion is bound or extracted, P-450 is inactivated.
Clinically, molnupiravir is used when remdesivir is ineffective. Currently, the drug has Emergency Use Authorization (EUA) in the U.S., but can be purchased with a prescription in the UK. It is administered orally, making it easier to use than remdesivir. Side effects include diarrhea, nausea, and dizziness. This drug should be used cautiously in men and women of reproductive potential, and used during pregnancy only if the potential maternal benefit outweighs potential fetal risk. Molnupiravir has caused fetal harm in animal studies.²⁰ Adults take 800 mg every 12 hours for 5 days, starting as soon as possible after diagnosis of COVID-19, and within 5 days of symptom onset.²⁰ U.S.)²⁰ When molnupiravir was first announced in the press, a chemist colleague of the author said, “You see, all your vaccinations were a waste, now you can just take a pill and get better!” Wrong! You don’t want to get sick in the first place, thereby avoiding chemotherapy and its side effects!
Paxlovid, a mixture of nirmatrelvir and ritonavir, was introduced without media fan-fair (thankfully) by Pfizer on Nov. 5, 2021 (see Table 1).³³ This clever drug works as follows. Recall that replicase is composed of 16 sub-proteins. When replicase is first translated from a vRNA strand into the amino acid language of proteins, 11 of these proteins are released from ribosomes (the cell’s protein manufacturing macromolecular micro-machines) as a single nascent string of amino acids that must be cut before the individual proteins can fold into the final shapes needed to form the replicase torus. This cutting is done at specific cleavage sites by nsp5. The protein nsp5 is called a protease, and nirmatrelvir (Figure 7B) is a protease inhibitor that is effective against all coronavirus strains. Therefore, in the presence of nirmatrelvir a functional replicase torus can never be formed, and without replicase the coronavirus can never multiply! What about the ritonavir? What does that do?
Ritonavir (Fig. 7C) is an HIV protease inhibitor that acts as a kind of preservative in this case. The endoplasmic reticulum¹⁷ of human cells contains a family of proteins collectively known as cytochrome P-450. They have that name because cytochromes (“cyto” from cytoplasm, “chrome” meaning color) contain a molecular structure called a heme group that, like the heme in blood, is capable of absorbing molecular oxygen (O₂) and carbon monoxide (CO) because of its iron atom. When P-450 absorbs CO, it has a peak optical absorption at a wavelength of 450 nm (blue light) — hence, the name. What is so important about P-450 is that it metabolizes xenobiotics (denoted by RH), foreign (ξένος, xenos) substances like nirmatrelvir, for eventual excretion. It is precisely the heme group’s ability to absorb, and split, O₂, that facilitates this process (heme Fe³⁺ + RH + 1e⁻ ultimately from the B vitamin riboflavin → Fe²⁺RH + O₂ from air → Fe²⁺OORH + 1e⁻ from riboflavin → Fe³⁺OORH + H₂O waste → ROH + Fe³⁺). The hydroxylation of xenobiotics (RH → ROH) renders them water soluble, the first step toward excretion. Ritonavir, beside its other activities, is also a P-450 inhibitor (Fig. 7C), thereby prolonging the half-life of nirmatrelvir in-vivo and allowing lower doses to be used!
This discussion on P-450 also raises two interesting, and frightening, questions. First, can a COVID strain evolve whose genome encodes for a protein that up-regulates (either directly or indirectly) the production of P-450; thereby neutralizing xenobiotic therapeutics before they can do their job? In that case, the dose of ritonavir would have to be increased to offset the additional P-450 production. Second, how much ritonavir can a patient tolerate?
Clinically, biochemists are always looking for drugs with less side effect. Nirmatrelvir was derived by molecular modification from drugs unglamorously called GC-376 and GC-373 that were used to treat a feline coronavirus infection that shared some similarities to human COVID-19. Neither of these last two drugs seemed to have any toxicity in human in-vitro cell cultures. However, in-vivo use of Paxlovid can have significant side effects, including hypertension (high blood pressure), diarrhea, myalgia (muscle pain), and dysgeusia (distortion of the sense of taste). The use of Paxlovid is contraindicated with the concurrent use of 26 other common drugs, and is also contraindicated for patients with severe renal or hepatic (liver) impairment. Use during pregnancy only if potential maternal benefit justifies potential fetal risk. Furthermore, safety has not been established for breast feeding or for children. There is no U.S. FDA authorization for children under 12 years old or under 40 kg. Typically, healthy adults will receive two nirmatrelvir 150 mg tablets and one ritonavir 100 mg tablet twice daily for 5 days. Treatment should be started as soon as possible after diagnosis of COVID-19 and within 5 days of symptom onset.²⁰
Dexamethasone – An Anti-Inflammatory Drug
Dexamethasone is different than the previous pharmaceuticals that targeted the viral life cycle. In fact, dexamethasone doesn’t interact with the virus at all. Instead, it targets the host’s response to the virus. Most of the damage done to the human body during COVID-19 infection is due to an overly aggressive response to the coronavirus.¹² In many severe cases, it would be helpful to “dial back” (down-regulate) the body’s inflammatory response. This can be done with steroids. Dexamethasone (“-sone”, or “-one”, refers to steroids) is a long acting corticosteroid. Steroids are a large group of substances whose most famous member is cholesterol, a fatty substance (a lipid) with an evil reputation because excess amounts of it can accumulate in arteries. However, cholesterol is actually necessary for life. It is a constituent that stiffens animal cell membranes, and more importantly for this discussion, it is the precursor for the body’s synthesis of many steroid hormones (Figure 8). Hormones are substances that allow specialized secretory cells to communicate with, and regulate the metabolism of, target cells that possess hormone specific receptors. In short, hormones are the chemical language that certain pairs of cells use to communicate, and blood vessels are the “telegraph wires” of this communication. There are five major classes of hormones: progestins (the regulators of events during pregnancy, and the precursors to all the other steroid hormones – Figure 8), glucocorticoids, mineralocorticoids, androgens (e.g. testosterone), and the estrogens. Of these, it is the glucocorticoids (Figure 8) that are of most interest, because these are the molecules that suppress inflammation.¹⁵ Glucocorticoids are produced by the adrenal cortex – hence, the suffix “corticoids” in glucocorticoids, and the prefix “cortico” in corticosteroids. Dexamethasone is a synthetic glucocorticoid that has 20 to 30 times the binding affinity for glucocorticoid receptors than its nearest natural predecessor (cortisol, Figure 8).³⁷ A general feature of natural steroid hormones is that they are not stored for release after synthesis. Therefore, the level of circulating hormone is controlled primarily by its rate of synthesis, which is often ultimately controlled by signals from the brain. As a synthetic steroid hormone, dexamethasone is administered intravenously to ease inflammation or airway edema (swelling due to collection of fluid in the intercellular spaces of tissues; a common symptom of severe COVID).²⁰
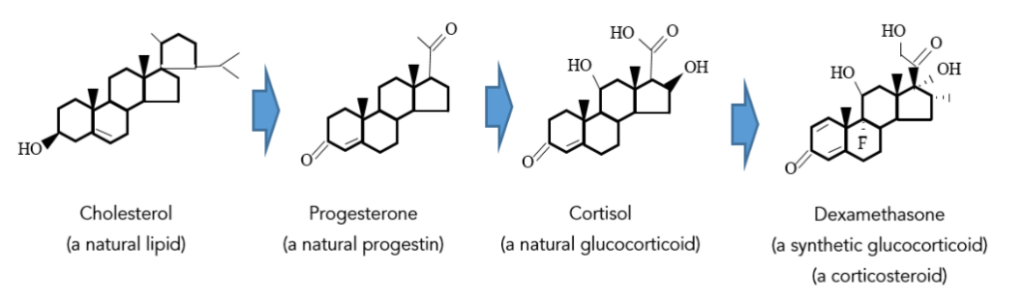
Figure 8 – The family tree of dexamethasone (C₂₂H₂₉FO₅),³⁸ starting from cholesterol.¹⁵ “Dexametha” means the “right-handed arrangement of methyl groups”; a -CH₃ group is understood to be at the end of each “dead-end” bond. “Dex” is an abbreviation of dexios (δεξιός), Greek for “right (handed)”.
Now that the nature of steroids (natural and synthetic) has been explained, the next step is to discuss what happens after they bind to cellular receptors. When you get a common cold, and you blow your nose, what comes out? Well, some water comes out, but it’s certainly not pure water. It is full of dead neutrophils (white blood cells of the adaptive immune system that stain a neutral pink). These phagocytes (phago-, φάγω, eater-, cells) are the most common immune cells in blood. There is a popular rumor that the color of the nasal discharge tells you if you have a bacterial or viral infection. Actually, the color only tells you the magnitude of the infection. The darker the discharge, the more neutrophils it contains. These neutrophils have gobbled up virions and then committed suicide so that other immune cells can scavenge their bodies – virus and all. This programmed cell death is called apoptosis (apo-, ἀπό-λυτος, dead). Glucocorticoids, like dexamethasone, down-regulate the genes responsible for apoptosis after binding to neutrophil receptors.³⁹
Other effects of the corticosteroid-receptor combination include reduction of neutrophil migratory behavior, and their binding to the epithelial cells (the lining) of the lungs.⁴⁰ Also, anti-inflammatory genes, like those that ultimately produce interleukin-10, are up-regulated.³⁹ The interleukins will be discussed in more detail in the next section. Finally, an infection triggers the formation of a whole family of inflammatory agents called eicosanoids.¹⁵ Like hormones, the eicosanoids exert specific physiologic effects on target cells. However, they only act locally near their site of synthesis (for example, around a bee sting). By contrast, hormones can affect cells throughout the body. Some experts consider eicosanoids to be local hormones. However, unlike hormones that are derived from cholesterol, eicosanoids are derived from arachidonic acid (Figure 9), a 20-carbon polyunsaturated fatty acid; eicos (είκοσι) means 20, and the suffix -oid is short for the Greek oidos (είδος) meaning form, hence an “eicos-an-oid” is a molecule with a “20 carbon form”. Another of dexamethasone’s jobs is to down-regulate the production of arachidonic acid and, therefore, all its eicosanoid derivatives.³⁹
At this point the reader may wonder why the human body has a two-tier hormone system (a global endocrine hormone system and a local eicosanoid system). Why not just let the endocrine system take care of everything? The answer is simply to prevent a trivial injury, like a minor cut or an insect bite (note: spiders, arachnids, do not contain arachidonic acid in their venom, it’s made by your own body if you are bitten), from sequestering too many resources from the whole body – something which adrenaline could easily do! It’s a forced austerity, a kind of biological Balanced Budget Amendment!
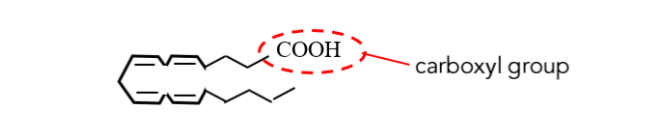
Figure 9 – Arachidonic acid, the precursor to a series of inflammatory eicosanoids, in its “hairpin” molecular configuration. Reduce the production of arachidonic acid and you reduce inflammation. Arachidonic acid can also act as a molecular messenger in its own right. Note the pincer on the open end of this molecule, a device exhibited in Figure 7C, but this time one of the pincers ends in an acidic carboxylic group. Arachidonic acid can undergo an acid/base reaction with any basic side chain of an amino acid in an appropriate receptor to signal inflammation. It is curious how hormones and eicosanoids (local hormones) are both derived from lipids – but different lipids!
Finally, the clinical aspects of dexamethasone and other corticosteroids need to be discussed. Initially, when the use of dexamethasone was piloted in the UK as a COVID therapy, some physicians thought that perhaps it only prolongs the inevitable death, thereby increasing the patients suffering.⁴¹ However, as of 2023 this has not proven to be the case. Instead,
“multiple randomized trials indicate that systemic corticosteroid therapy improves clinical outcomes and reduces mortality in hospitalized patients with COVID-19 who require supplemental oxygen presumably by mitigating the COVID-19-induced systemic inflammatory response that can lead to lung injury and multi-system organ dysfunction.”⁴²
This positive assessment comes as a bright light in a story that has so often been dark and disappointing. Currently, the U.S. NIH (National Institute of Health) is recommending 6 mg of dexamethasone intravenously once daily for up to 10 days or until hospital discharge (whichever comes first) for all COVID-19 patients who require mechanical ventilation.⁴³ However, in patients who do not require supplemental oxygen, the use of corticosteroids may actually increase mortality!⁴² This is why large clinical trials, with control groups and placebos, are necessary.
Interleukins
Interleukins, as well as all the therapeutics in succeeding sections, are qualitatively different from those discussed in preceding sections in the sense that they are not simple molecules whose detailed structure can be easily diagrammed and comprehended. Instead, we are now in the realm of giant protein molecules, and will have to content ourselves with more general structural cartoons; although detailed structural maps do exist for use by specialists. The interleukins are the smallest of these natural proteins (Figure 10), and they can also be used as therapeutic agents.
The interleukins are the signaling molecules for the human immune system. The name comes from inter, meaning between, and leukocyte, the technical name for a white blood cell – leuko is Greek for white (λευκός); the English cognate is light. So, interleukins are the chemical signals that move between white blood cells. Although free-floating mature white blood cells may appear to be independent amoeba-like entities because they are not physically anchored to other tissues, they are connected by the interleukin signaling system! The reader might wonder why the human body has so many different types of signaling molecules — there are endocrine hormones, eicosanoids, and now interleukins. Everything in nature has a purpose, and here it is to prevent “cross-talk” between the various systems within the body. It would be very bad news, for example, if a signal to produce more B-cells also initiated the global release of arachidonic acid! There are no telephone “party-lines” allowed in the human body! With these preliminaries in mind, consider two relevant examples of interleukin signaling.
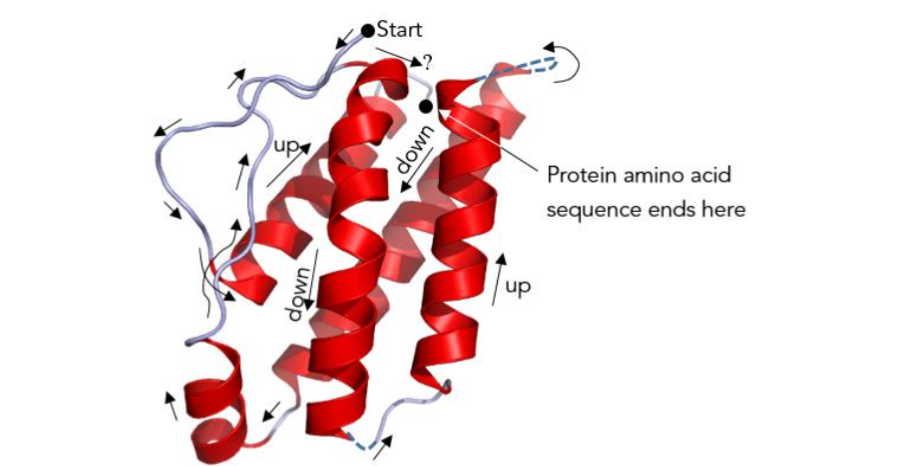
Figure 10 – A cartoon of interleukin-2 (denoted by IL-2 in the literature) with a molecular weight of about 16,000 Daltons. Some amino acid sequences of a nascent protein released from a ribosome will naturally fold into the “corkscrew” shaped motif shown. It is called an α-helix, and was represented in Figure 6 as a sausage shaped tube. IL-2 has a bundle of four such secondary structures (secondary because they form after the primary sequence of amino acids forms) — a very biochemically recognizable up-down-up-down anti-parallel stacking that is characteristic of an even larger family of signaling molecules called cytokines (cyto from cytoplasm, and kine from the Greek κινέω meaning to move or kinetic in English). Figure modified by the author.⁴⁴
Vaccination, or wild infection, causes B-cells (both infection memory cells, and effector types that produce antibodies) to proliferate. How does that happen? When immune system helper T-cells (produced in the Thymus) recognize an antigen bound to a B-cell, they release the signal protein interleukin-2 (IL-2) that stimulates B-cell reproduction.¹⁵ IL-2 has been tested as a post-infection therapeutic agent to boost the immune response to COVID-19. Early experiments in China had shown recombinant IL-2 (i.e. IL-2 artificially produced by an unlikely host such as a yeast¹⁷) to be a useful adjunct when used with other therapies.⁴⁵ More traditionally, interleukin-2 has been used to treat cancer, both systemically and locally, by injection; because IL-2 also increases the cell killing activity of the immune system’s NK (Natural Killer) cells that are so effective against tumors. Systemic anticancer use of IL-2 is attended by a variety of side-effects ranging from minor to severe, including: flu-like symptoms (fever, headache, muscle and joint pain, fatigue), nausea/vomiting, dry itchy skin or rash, weakness or shortness of breath, diarrhea, low blood pressure, drowsiness or confusion, loss of appetite, breathing problems, serious infections, seizures, allergic reactions, heart problems, kidney failure, or a variety of other complications.⁴⁵
In the previous section on dexamethasone, it was mentioned that interleukin-10 (IL-10) was an anti-inflammatory agent. Why? One reason is that IL-10 inhibits pro-inflammatory cytokine production (see Figure 10 caption), thereby preventing, or at least reducing, something called the cytokine storm; the uncontrolled and excessive release of signaling cytokines (hypercytokinemia). It is the subsequent over-reaction of the immune system that kills so many COVID-19 patients. Twenty to thirty percent of those with hypercytokinemia may die.⁴⁶ Almost all have to be treated in an intensive care unit. Low oxygen saturation levels (in tissues) and shortness of breath are co-morbidity factors. Another reason is that IL-10 prevents maturation of two types of white blood cells (macrophages and dendritic cells), thereby limiting immune response. What about the other interleukins?
When a helper T-cell recognizes an antigen on the surface of a B-cell, it doesn’t just stimulate the B-cell to reproduce. It also does the same to itself! Two types of daughters are possible; Tₕ1 and Tₕ2. It is the Tₕ2 cells that are of interest to the reader in this section because these are the ones that secrete interleukins. IL-4, IL-5, and IL-13 defend against extracellular pathogens; those that are outside the Tₕ2 cell membrane. They do this by stimulating B-cells to make antibodies as usual, but the main antibody they produce is called IgE. Exactly what IgE is will be explained more thoroughly in a subsequent section on antibodies. For now, the reader only needs to know that IgE can bind to mast cells, causing release of histamine, an irritating substance that can cause sneezing, coughing, and diarrhea.²⁸ This is nature’s way of expelling extracellular microbes from the body’s epithelial surfaces (lining of the lungs, etc.). Although IL-4, IL-5, and IL-13 are not used as drugs, they do play a role in the course of COVID and patient discomfort.
Still other interleukins are involved in the destruction of bacteria, or the expulsion of worms – none of which are of concern here. What about Tₕ1, what do those daughters do? These cells are concerned with activating macrophages to kill captured microbes (like viruses) within their phagosomes (digestive organelles). Tₕ1 cells also activate cytotoxic T-cells to kill infected cells. So, Tₕ1 cells defend against intracellular pathogens, and in that case the most important activating chemical signal they secrete is interferon-γ. Therefore, it is the interferons that will be discussed next.
Interferons
Interferons, like interleukins, are cytokine signaling molecules. Unlike interleukins, however, they can also affect the behavior of cells that are not part of the immune system. There are three main classes of interferons: α, β, and γ. All are of interest for the treatment of COVID, especially γ. The molecular weight of interferon-γ is between 40,000 and 70,000 Daltons as determined by molecular sieve chromatography (Figure 11). However, γ exists as a dimer, so the molecular weight of each monomer may be in the 20,000 to 35,000 Dalton range (Figure 11).⁴⁷ In either case, interferons are definitely heavier than interleukins. Naturally, interferons get their name from the fact that they interfere with viral replication in several ways. The ability of γ to activate macrophages and cytotoxic T-cells of the immune system by binding to receptors on their surface has already been mentioned. However, some interferons can affect the behavior of epithelial cells on the lining of the lungs. When epithelial cells detect the right kind of interferon (there are 21), they slow down their protein production. However, viruses need proteins to multiply – they need to make more S-proteins, nucleocapsid proteins, etc. Therefore, viral replication slows. Many cells may also die from protein starvation. However, that doesn’t matter, so long as ribosomal protein output isn’t sequestered by enemy viruses. It’s biochemical scorched earth warfare, and you begin to feel less hungry when you are infected. You may even lose your sense of taste or smell (a common COVID symptom) – food just doesn’t smell as tempting, or taste as good, anymore. The body’s overall strategy here is to slow down the infection while the adaptive immune system (white blood cell count, antibody levels, etc.) comes up to speed. However, this strategy is somewhat counter-productive since the body needs adequate nourishment to build a new army of white blood cells with antibody chemical weapons specially adapted to a patient’s particular virus. In any case, these are just a few of the many biochemical pathways that can be controlled by interferons in the body’s fight against COVID. Interferons can be powerful therapeutics, but they can also be dangerous toxins, as the reader will soon realize.
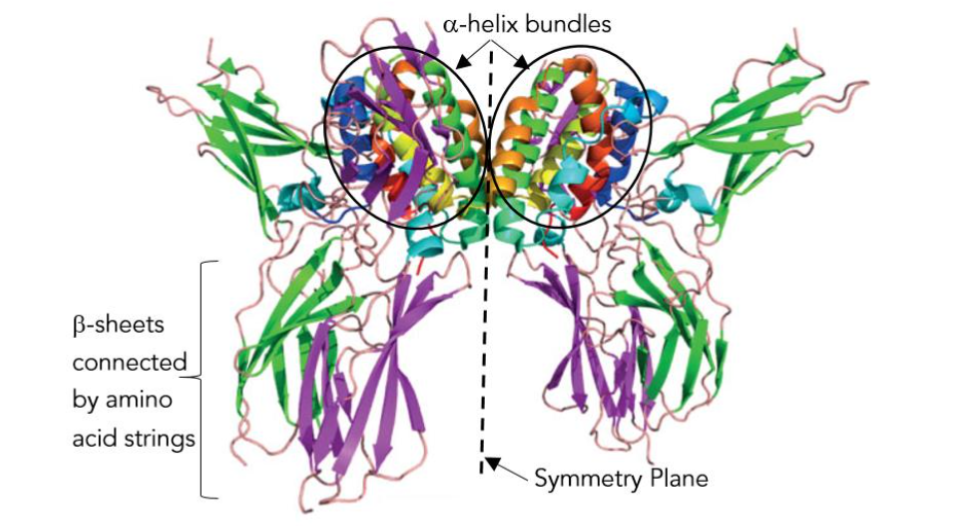
Figure 11 – The “butterfly” structure of interferon-γ. The right and left halves are the two monomers of this dimer (like the S1 and S2 monomers of the S-protein dimer). Each monomer is made up of amino acid strings, β-sheets (flat ribbons of amino acids), and an α-helix (amino acid corkscrew) bundle (cf. Figure 10). Figure modified by the author.⁴⁸ With two receptor binding sites, even minute quantities of γ produce major physiological effects!
The competition between a virus and the human immune system can be brought into sharper focus by realizing that the reproductive cycle of a virus ranges from 8 hours for the picornavirus (small RNA virus – polio) to more than 72 hours for some herpesviruses (cytomegalovirus), with the virus yield per cell ranging from several thousand for poxvirus to more than 100,000 for poliol.⁴⁹ By contrast, the adaptive immune system can take several days to reach full operational capacity, but interferons can be detected in the blood stream just a few hours after an initial wild inoculation – long before any symptoms. So, interferon levels spike in a time interval commensurate with the viral life cycle.¹²
Clinically, several interferons, including α, β, and γ, have been evaluated for treatment of COVID-19. Although α-2a and α-2b have been approved by the FDA for treatment of hepatitis B and C, and β-1a for the treatment of relapsing forms of multiple sclerosis (MS), there are currently (Dec. 2023) no such approvals for COVID-19. Interferon λ is not currently approved or authorized by the FDA for any use. Interferon use to treat COVID-19 is limited to clinical trials only.⁵⁰ Interferon β-1a has a half-life in the body of 69 hours when administered subcutaneously, but its mechanism of metabolism and excretion is unknown.⁵⁰ This is the realm of experimental medicine. Interferons have many, many, serious side effects, including chest pain, edema, hypertension, a variety of dermatological conditions, sinusitis, vision abnormalities, constipation, nausea, vomiting, abdominal pain, autoimmune hepatitis, up-regulation of liver enzymes, cystitis (inflammation of the urinary bladder), erectile dysfunction, polyuria, urinary incontinence, a variety of serious blood disorders, injection-site reactions and necrosis, myalgia (muscle pain), arthralgia (joint pain), drug-induced lupus erythematosus, muscle spasms, up-regulation of seizures with β-1a, depression, dizziness, fatigue, headache, incoordination, rigors (rigidity, stiffness), suicidal ideation (thoughts), dyspnea (labored breathing), upper respiratory tract infection, hypersensitivity reactions including anaphylaxis and angioedema (giant hives), and a few relatively minor side effects like drowsiness, chills, fever, flu-like symptoms, pain, increased sweating, and dry mouth. One might wonder what concentrations of β-1a could produce so many draconian side effects? Prefilled syringes of Avonex (trade name) contain only 30 μg / 0.5 ml of solution!²⁰ Even in these multi-microgram doses, interferon signaling molecules produce consequences throughout the body, and may affect a patient’s mind as well. There is no free lunch!
Monoclonal Antibodies
Antibodies are the protein secretions of the immune system’s effector B-cells. The smallest of these proteins has an enormous molecular mass of 152,000 Daltons (atomic mass units).¹³ This section must necessarily begin with a more in-depth discussion of the exact nature of antibodies. The simplest antibodies (IgG or γ-globulin, IgD, and IgE) are “Y”-shaped molecules called monomers (Figure 12). The more complex antibodies (IgA and IgM) are made up of several monomers and are called dimers and pentamers, respectively (Figure 13). What is so amazing about antibodies is that the amino acid composition of the antigen binding site is highly variable, and antibodies exist that are capable of binding to virtually any antigen!¹⁵ A simple calculation explains why this is so. The variable regions of a monomer may have a total mass of about 46,000 Daltons (Figure 12), which is equivalent to about 418 amino acids, since the average molecular weight of the 20 human amino acids is 110 Daltons. Each of these 418 locations can be occupied by any one of 20 amino acids. Therefore, the total number of possible antibody monomers is 20⁴¹⁸, or 10⁵⁴⁴ (i.e. a 1 followed by 544 zeros)! When the human body is invaded by something foreign (an antigen), like a virus, a strange protein, or a toxic chemical, it only takes one antibody to fit (bind) closely to the irritant – like a glove on a hand. After that, the body makes more copies of that particular antibody, until the circulatory system is flooded with an army of antibody clones capable of inactivating a particular antigenic protein, or marking a virus for digestion by a phagocyte (one type of white blood cell). Many years ago, the author saw a science fiction movie called “The Andromeda Strain”. In that story a foreign germ is brought to Earth from space. The earthlings have no defense against it, and everyone who is infected dies. Not necessarily! Even if a germ from space reached Earth, there would eventually be one antibody that would fit it, and once found, it could be multiplied by the methods to be explained next.
Each antibody-producing B-cell (B lymphocyte produced in the Bone marrow and the lymphatic system’s spleen) is specialized for the synthesis and secretion of one, and only one, antibody – say, for example, an antibody directed against the coronavirus S-protein binding site. If one such B-cell could be cultured outside the body, its daughters would all be clones (identical replicas) of their one parent (hence the term monoclonal), and all would be capable of secreting that one desirable antibody for harvesting.¹⁵ The problem is that B-cells cannot be grown in a petri dish. Here is where coronavirus story intersects with some exciting cancer research, because cancerous cells can be grown in a petri dish or test tube, and will proliferate indefinitely. They are not subject to the apoptosis (programmed cell death) discussed above in connection with dexamethasone. Cancer cells can continue to divide forever beyond the Hayflick limit; a preset number of cellular divisions which a particular cell line cannot exceed (usually about 55, except for stem cells). In 1975, Kohler and Milstein discovered that it was possible to fuse cancerous B-cells from a mouse with myeloma (a cancerous proliferation of white blood cells), with normal B-cells capable of expressing a specific antibody. The resulting line of cells, called a hybridoma, is both immortal and capable of expressing a specific antibody. With this technology, mass production of monoclonal antibodies proceeds as follows. A healthy laboratory mouse is injected with an antigen – like the S-protein, for example. After a while, the mouse begins to produce antibodies. Its spleen is then removed, and a suspension is made of its cells – some of which are B-cells that secrete S-protein antibodies. This suspension is then mixed with a suspension of myeloma B-cells in a test tube. After a while some B-cell fusions will take place forming hybridomas. All the cells are then transferred to a medium in petri dishes that only support the growth of the hybridoma. The hybridomas then multiply indefinitely and secrete antibodies that can be harvested and purified. In reality, the mouse’ immune system will respond to several parts of the S-protein (not just the binding site), and each is called an antigenic determinant. So, there will actually be several types of B-cells in the spleen suspension, each producing one specific monoclonal antibody.
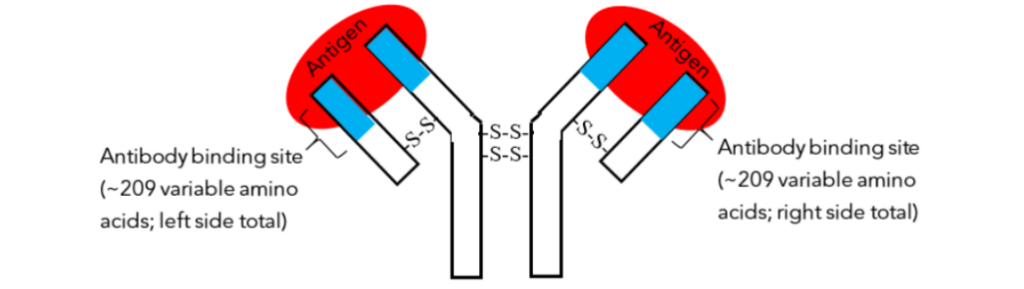
Figure 12 – The structure of a monomer. Only general regions can be depicted due to molecular size and complexity. The pieces of the “Y” shaped molecule are held together by disulfide bonds (-S-S- bonds). Two antigens (red) are bound to the antibody binding sites (blue). The binding sites have a variable amino acid composition that can match almost any antigen.
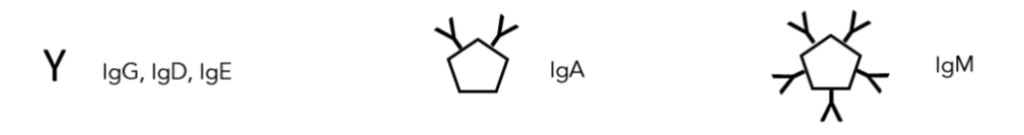
Figure 13 – The five classes of antibodies. IgG (for Immunoglobulin G), IgD, and IgE are all monomers, but IgA and IgM are a dimer and a pentamer, respectively. The globins are simply molecules that bind to other things. Hence, hemoglobin binds oxygen, and immunoglobulins bind antigens; while the word globin itself comes from the Latin globulus, meaning little ball, a reference to hemoglobin bearing red blood cells. IgM is the heaviest protective protein that will be discussed in this publication, with a molecular weight approaching one million Daltons. Antibodies have multiple binding sites (from 2 to 10) so they can clump antigens together for precipitation, or digestion by still larger white blood cells.
As of March 6, 2023, there are four monoclonal antibody products on the market that have received Emergency Use Authorization (EAU) by the U.S. FDA for treatment of COVID-19. These are: 1) bamlanivimab plus etesevimab, 2) casirivimab plus imdevimab, 3) sotrovimab, and 4) bebtelovimab. The suffix “mab” stands for Monoclonal AntiBody, while “vi” denotes “virus”. All of these products target the all-important spike (S) protein, as might be expected. Although MABs have been shown to have clinical benefits in treating COVID-19, their application is limited. The problem with all these drugs is that the coronavirus mutates so rapidly,¹¹ that the MABs are soon outdated. In fact, laboratory studies have shown that activity even against specific variants and sub-variants can vary dramatically from locality to locality due to minor local S-protein variations, thereby rendering some areas resistant to MAB therapy.⁵¹ In short, MABs are too specific. EUA authorization is only for treatment of outpatients with mild to moderate COVID-19.¹⁵ The author wonders why MABs would be used for mild COVID since this is not an emergency use?
Conclusion
Fighting the pandemic has been greatly complicated in the U.S. by political rivalries. The political left has been skeptical of therapeutics, and not without some justifiable cause when one considers both side effects and the number of rumored cures. Worse still, the scientific literature on therapeutics is steeped in a mystifying jargon (which the author has attempted to elucidate) that comes from a language and script that few Americans understand. While the political right has been skeptical about vaccination, in an atmosphere where understandable scientific explanations have been lacking (even from the U.S. CDC). In reality, both vaccines and therapeutics are needed. The confrontation regarding the use of masks has been particularly contentious. Few people wear them. Not for any scientific reason, but because those who do are intimidated by the ridicule of those who don’t. Statements like, “Halloween is over!”, or “I don’t dance with men who wear masks!” are powerful psychological deterrents. Dr. Frances Collins, former director of the National Institute of Health (NIH), head of the Human Genome Project, and recipient of the Presidential Medal of Freedom (2007) by former President George W. Bush, summed up the current situation in the U.S. this way in a Dec. 3, 2021 interview for Nature, just 16 days before his retirement from public service:⁵²
“We’re in a really bad place. If science happens to produce a result that a political perspective doesn’t like, then science has to be attacked. …This has turned out to be a more severe situation than I would have imagined a year ago. I wish we had more insights from behavioral social science research into how this has come to pass, and why it could have gotten so completely widespread. I want to call this out as one of my most major concerns as I stepped down from the NIH, of looking at the situation in our nation. Somewhere along the way, our political hyperpolarization began having a lot of really dangerous consequences, where in many instances we seem to have lost our sense of how to tell the difference between a fact and an opinion — or some Facebook post that’s, frankly, a lie. That’s truly dangerous. That’s another epidemic that is not going to go away even if we triumph over COVID-19. We need to figure out what happened here, and how to bring ourselves back to a place where our nation has a more stable future.”
– Stay Well!
Conflict of interest statement:
The author declares that he had no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
1. Worldometer. Daily Deaths. Nov. 16, 2023. Accessed Nov. 16 www.worldometers.info
2. Qasmieh SA et al. The prevalence of SARS-CoV-2 infection and long COVID in U.S, adults during the BA.4, BA.5 surge, June-July 2022. Preventative Medicine, Apr. 2023. Accessed Mar. 20, 2023 https://pubmed.ncbi.nlm.nih.gov
3. Sidharthan C. Study shows COVID-19 rates were likely forty-times higher than CDC estimates during BA.4/BA.5 dominated period in the U.S. News: Medical & Life Sciences, Feb. 27, 2023. Accessed Mar. 20, 2023 https://www.news-medical.net
4. Innes S. Arizona reduces COVID to ‘common’. Arizona Republic, Oct. 9, 2023; p. 1A, 10A.
5. Prater E. The CDC has guided the U.S. COVID epidemic to a soft landing – a manufactured conclusion that flies in the face of science some experts say. Fortune, Health·Coronavirus, Aug. 21, 2022. Accessed Dec 21, 2022 https://fortune.com
6. Prater E. COVID is at or near record levels around the country. Scientists say one (pretty gross) indicator doesn’t lie. Fortune Well, July 26, 2022. Accessed Mar. 20, 2023 https://fortune.com
7. Shah S et al. Wastewater surveillance to infer COVID-19 transmission. NIH (National Institute of Health), 2023. Accessed Mar 20, 2023 www.ncbi.nih.gov
8. Nebraska Medicine. What COVID variants are going around in September 2023. Nebraska Medicine, Sept 7, 2023. Accessed Sept. 26, 2023 www.nebraskamed.com
9. Maxmen A. COVID booster may not reach most vulnerable. Arizona Daily Star, Sept 22, 2023, p. A1, A2.
10. Mandavilli A. In Hospitals, Viruses Are Everywhere. Mask Mandates Are Not., New York Times. Sept 25, 2023, No. 59,922, p. A1, A23.
11. Zito RR. The Omicron Strain. Presented at the Annual International System Safety Conference, Aug. 28 – Sep. 1, 2023, Portland, Oregon, USA.
12. Dettmer P. Immune. Random House, New York, 2021: 162-67, 86-203, 219-221, 251-257.
13. Zito RR. The Delta Variant. J. System Safety, Fall 2022; 57(3): 7-34.
14. Wessells NK, Hopson JL. Biology. Random House, New York, N.Y., 1988: 8.
15. Mathews CK, van Holde KE. Biochemistry, 2ng ed. The Benjamin/Cummings Pub. Co. Inc., Menlo Park, CA, 1996: 3,15, 88, 91-92, 132, 146-147, 246-247, 248-251, 256-257, 326, 549-550, 678-688, 582-583, 683-688, 692-696, 885-7.
16. Tortora GJ, Derrickson B. Principles of Anatomy and Physiology, 11th ed. John Wiley, New York, NY, 2006: 149.
17. Zito RR. COVID-19, J. System Safety, Fall/ Winter 2020; 54(2): 10-25.
18. Yin W, et al. Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science, May 1, 2020; 368 (6498): 1499-1504. doi: 10.1126/Science.abc1560.
19. Sola I, Almazán F, Zúñiga S, Enjuanes L. Continuous and Discontinuous RNA Synthesis in Coronaviruses. Annu. Rev. Virol., Nov 2015; 2(1): 265-288. doi: 10.1146/annurev-virology-199114-055218, PMCID: PMC6025776 | NIHMSID, NIHMS975955 │ PMID: 26958916.
20. Vallerand AH, Sanoski CA. Davis’s Drug Guide for Nurses, 18th Ed., F.A. Davis, Philadelphia, PA, 2023: 102-105, 210-211, 373-380, 518-320, 563-564, 613-614, 627-630, 682-684, 724-725, 763-765, 973-975, 1106-1108, 1269-1273, 1401-1402.
21. Mehta RM, Bansal S, Bysani S, and Kalpakam H. A shorter symptom onset to remdesivir treatment (SORT) interval is associated with a lower mortality in moderate-to-severe COVID-19: A real world analysis. Int. J. Infect. Dis., May 2021; 106: 71-77. Published online Feb 26, 2021. doi: 10.1016/j.ijid.2021.02.092, PMCID: PMC7908839 | PMID: 33647517. Nat. Lib. of Med. Accessed Sept 25, 2023 https://www.ncbi.nlm.nih.gov
22. UTNEWS. Scientists Discover How Remdesivir Works to Inhibit Coronavirus. UTNEWS: Science & Technology, U. Texas, Austin, TX, Jan 28, 2021. Accessed Sept 25, 2023 https://news.utexas.edu
23. Eastman RT, et al. Remdesivir: A Review of Its Discovery and Development Leading to Emergency Use Authorization for Treatment of COVID-19. ACS Cent. Sci., May 27, 2020; 6(5): 672-683. Published online 2020 May 4, www.ncbi.nlm.nih.gov. doi: 10.1021/acscentsci.0c00489, PMCID: PMC7202249 | PMID: 32483554.
24. Strauss JA, Strauss EG. Viruses and Human Disease, Academic Press, San Diego, 2002: 149, 201.
25. Zito RR. Vaccine Safety. J. System Safety, Fall 2021; 57(1): 10-25.
26. Longhitano L et al. Proteasome inhibitors as a possible therapy for SARS. NIH, May 2020. Accessed Oct 13, 2023 https://ncbi.nlm.nih.gov
27. Matrosovich MN, Matrosovich TY, Gray T, Roberts NA, Klenk HD. Neuraminidase Is Important for the Initiation of Influenza Virus Infection in Human Airway Epithelium. J. Virol., Nov 2004; 78(22): 12665-12667, doi:10.1128/JVI.78.22.12665-12667.2004, PMCID: PMC525087 | PMID:15507653. Accessed Nov 20, 2023 www.ncbi.nlm.nih.gov
28. Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. The Cell, 4th Ed., Garland Science, New York, NY, 2002: 148, 1412-1413.
29. Hopkins JS, McKay AB. Merck Covid-19 Pill Cuts Risks of Hospitalizations and Death. The Wall Street Journal, Oct 2-3, 2021; Vol. CCLXXVIII, No. 79: A1, A6.
30. PubChem. Molnupiravir. 2023. Accessed Nov 27, 2023 https://pubchem.ncbi.nlm.gov
31. Wikipedia. Molnupiravir. 2023. Accessed Nov. 27, 2023 https://en.m.wikipedia.org
32. Wikipedia. Nirmatrelvir. 2023. Accessed Nov 27, 2023 https://en.m.wikipedia.org
33. Carr F. The Biochemistry and Broad Utility of Pfizer’s New COVID-19 Drug; Paxlovid. Biological Sciences, (Fall) Dec 8, 2021. Accessed Nov 27, 2023 https://sites.dartmouth.edu
34. Wikipedia. Ritonavir. 2023. Accessed Nov 29, 2023 https://en.m.wikipedia.org
35. Loos NHC, Beijnen JH, Schinkel AH. The Mechanism-Based Inactivation of CYP3A4 by Ritonavir: What Mechanism? Int. J. Mol. Sci., Sept 2022; 23(17): 9866. Published online Aug 30 2022, doi: 10.3390/ijms23179866, PMCID: PMC9456214 │ PMID: 36077262.
36. Davydov R et al. The role of the proximal cysteine hydrogen bonding interaction in cytochrome P450 2B4 studied by cryoreduction/EPR/ENDOR spectroscopy. Biochemistry, Feb 16, 2016; 55(6): 869-883. Published online Feb. 3, 2016, Pubmed Central, https://www.ncbi.nlm.nih.gov. doi: 10.1021 /acs. biochem. 5b00744, PMCID: PMC4834902 | NIHMSID: NIHMS775304 | PMID: 26750753.
37. Zabirowicz ES, Tong TJ. “Pharmacology of Postoperative Nausea and Vomiting”, in Pharmacology and Physiology for Anesthesia, 3nd Ed., 2019. Available online from ScienceDirect. Accessed Dec 3, 2023 www.sciencedirect.com
38. Wikipedia. Dexamethasone. Nov 13, 2023. Accessed Dec 3, 2023 www.en.m.wikipedia.org
39. DrugBank. Dexamethasone. 2023. Accessed Dec 13, 2023 https://go.drugbank.com
40. Stroever SJ, et al. Medication Use Among Patients With COVID-19 in a Large National Dataset: Cerner Real-World Data. Clin. Ther. June 2021; 43(6): 173-196. Published online Apr. 15, 2021. Accessed Dec 13, 2023. doi: 10.1016/j.clinthera.2 021.03.024, PMCID: PMC8049452 | PMID 33958234
41. Mueller B, Rabin RC (2020, June 17). Hope Meets Caution as Steroid Is Said to Reduce Virus Deaths. The New York Times, June 17, 2020; No. 58727: A1, A6.
42. NIH. Systemic Corticosteroids, in National Institute of Health COVID-19 Treatment Guidelines. NIH, July 21, 2023. Accessed Dec 2, 2023 www.covid19treatmentguidelines.nih.gov
43. NIH (2023b, Oct. 10). Therapeutic Management of Hospitalized Adults With COVID-19. NIH, Oct 10, 2023. Accessed Dec 2, 2023 www.https://covid19treatmentguidelines.nih.gov
44. Wikipedia. Interleukin-2. Aug 12, 2023. Accessed Dec 19, 2023 https://en.m.wikipedia.org
45. Zhu ME, Wang Q, Zhou S, Wang B, Ke L, He P. Recombinant interleukin-2 stimulates lymphocyte recovery in patients with severe COVID-19. Published online in Experimental and Therapeutic Medicine, Jan. 18, 2021; article number 227. Accessed Sept 30, 2021 https://doi.org/10.3892/etm.2021.9658
46. Ramatillah DL, et al. Impact of cytokine storms on severity of COVID-19 disease in a private hospital in West Jakarta prior to vaccination. PLOS One, 17(1), e0262438, published online Jan. 25, 2022. Accessed 2023. doi: 10.1371/journal.pone.0 262438, PMCID: PMC8789122 | PMID: 35077495.
47. Yip YK, Barrowclough BS. Molecular Weight of Human Gamma Interferon Is Similar to that of Other Human Interferons. Science, Jan 22, 1982; 215(4531): 411-413. Accessed 2023 https://www.science.org.
48. Walter MR. The Role of Structure in the Biology of Interferon Signaling. Front. Immunol., Nov 12, 2020. Accessed 2023 https://www.frontiersin.org
49. Roizman B (1996). Multiplication. In Medical Microbiology, 4th Ed., Chapter 42, Baron S (editor), NIH, National Library of Medicine. Accessed Dec 22, 2023 https://www.ncbi.nlm.nih.gov
50. NIH. COVID-19 Treatment Guidelines – Interferons. Accessed Dec 22, 2023 www.https://covid19treatmentguide lines.nih.gov
51. NIH. COVID-19 Treatment Guidelines: Anti-SARS-CoV-2 Monoclonal Antibodies. NIH, March 6, 2023. Accessed Dec 17, 2023 treatmentguidelines.nih.gov
52. Subbaraman N. Science misinformation alarms Francis Collins as he leaves top NIH job, Nature, Dec 3, 2021; 600: 372-373. Accessed Dec 19, 2022 www.nature.com. doi: https://doi.org/10.1038/ d41586-021-03611-2.

