Ensemble Deep Learning for Chest X-Ray Disease Detection
Ensemble Deep Learning for Multi-Class Chest X-Ray Classification: Robust Detection of Pneumonia, and Tuberculosis
Shahmeer Shahzad1,2, Sajeela1, Nouman Shah1, Shahzad Ali Khan1
- Department of Public Health, Health Services Academy, Islamabad 45500, Pakistan
- School of Computing and Mathematical Sciences, University of Leicester, LE17RH, UK
OPEN ACCESS
PUBLISHED: 30 November 2025
CITATION: Shahzad, S., Sajeela., et al., 2025. Ensemble Deep Learning for Multi-Class Chest X-Ray Classification: Robust Detection of Pneumonia, and Tuberculosis. Medical Research Archives, [online] 13(11). https://doi.org/10.18103/mra.v13i11.7065
DOI https://doi.org/10.18103/mra.v13i11.7065
ISSN 2375-1924
ABSTRACT
The most common lung diseases, such as pneumonia and tuberculosis, remain some of the biggest global health diagnosis challenges, particularly in areas with extremely poor access to expert radiologists. Chest X-rays are affordable and accessible; however, their interpretation requires a great deal of expertise, which might not be consistently available across different clinicians. Recent advances in artificial intelligence, particularly deep learning, present promising solutions by automating and improving the interpretation of radiographic images. This study presents a robust system contributing to improved diagnostic performance by processing chest radiographs using state-of-the-art deep learning techniques. Various models were trained and evaluated for the detection of tuberculosis and pneumonia. An individual best performing model could achieve an accuracy of 98.73% while the result after an ensemble of diverse deep models could achieve a test accuracy of 98.05%. That proves that diverse deep learning models can substantially improve medical image analysis, enabling the development of more reliable diagnostic tools and offering accessibility across high-resource and low-resource healthcare settings. Code and all models have been made publicly available to foster transparency and subsequent research in AI-driven medical diagnostics.
Keywords
Deep Learning, Chest X-Ray, Pneumonia, Tuberculosis, Medical Imaging
1. Introduction
Pneumonia and tuberculosis still stand among the most dangerous infections worldwide, causing millions of people either to die or be hospitalized every year. These diseases predominantly spread in low-income and middle-income countries due to underpowered health resources and a shortage of skilled radiologists. Pneumonia is still the most infectious cause of death in children five years and below, while TB ranks among the top ten causes of death globally, with over 10 million new cases reported annually as reported by WHO.
Chest X-rays, being the most available and inexpensive way of imaging, are used as the first diagnostic tool among these conditions. However, it requires extensive expertise to interpret chest radiographs correctly, and even expert radiologists disagree about the findings, which again makes diagnostic variability with potential for errors.
Recent knowledge in deep learning within artificial intelligence has given promising results for automating and improving the interpretation of medical images. CNNs have achieved state-of-the-art performance in detecting abnormalities in chest radiographs, often competitively or even surpassing human experts’ performances. They can learn complex patterns from large datasets and generalize across a diverse imaging condition.
Despite such advance, most AI systems focus on single-disease detection, such as pneumonia, and have flaws such as overfitting, class imbalance, and poor generalizability. From a clinical perspective, in real-world clinical practice, it is very important to diagnose many conditions. Addressing these challenges, combining multiple models with ensemble learning has emerged as a robust approach toward enhancing accuracy and reliability in medical image classification.
The current study proposes a deep learning ensemble system that can classify chest X-rays as normal, pneumonia, and tuberculosis. In this respect, we seek to develop a system with different architectures for CNNs, embedding strict preprocessing and rigorous methodologies during its evaluation, so that the developed tool can be deployed in environments that are both rich and poor in resources.
1.1 SUPERVISED LEARNING
Supervised learning is one of the foundational modes of machine learning where the model is trained on labelled data, consisting of pairs of inputs and corresponding correct outputs. The model learns by minimizing an error between its predictions and actual labels, usually with optimization algorithms such as stochastic gradient descent. This is quite traditionally used in classification tasks, where for example, expert-labelled radiographs guide the model to learn to recognize disease patterns.
Annotating images from large datasets can enable supervised learning to perform disease detection automatically in medical imaging. However, class imbalance, noisy labels, and changes in image acquisition conditions are but a few of the challenges that may hinder performance and generalize a model.
1.2 TRADITIONAL MACHINE LEARNING TECHNIQUES
Before deep learning took over, most medical image classification tasks were done using traditional machine learning algorithms, including Support Vector Machines, decision trees, random forests, and k-nearest neighbours. These conventional methods are too heavily dependent on handcrafted features, designed using domain-specific knowledge in texture, shape, or intensity.
Although effective in simpler tasks, many of them suffered from complex image data due to their limited capability for the extraction of hierarchical and spatial features. This limitation showed the path toward powerful models that learned features directly from raw data.
1.3 ARTIFICIAL NEURAL NETWORKS
Artificial Neural Networks consist of layers of interconnected nodes, neurons, which, through the means of weighted connections and activation functions, carry out data processing. Though effective in modeling nonlinear relationships, ANN is good to go with structured data; it lacks the abilities for image analysis because it does not preserve spatial hierarchies.
In medical imaging, ANNs laid the groundwork for more advanced architectures but were eventually surpassed by models like Convolutional Neural Networks (CNNs), which are specifically designed to handle image data.
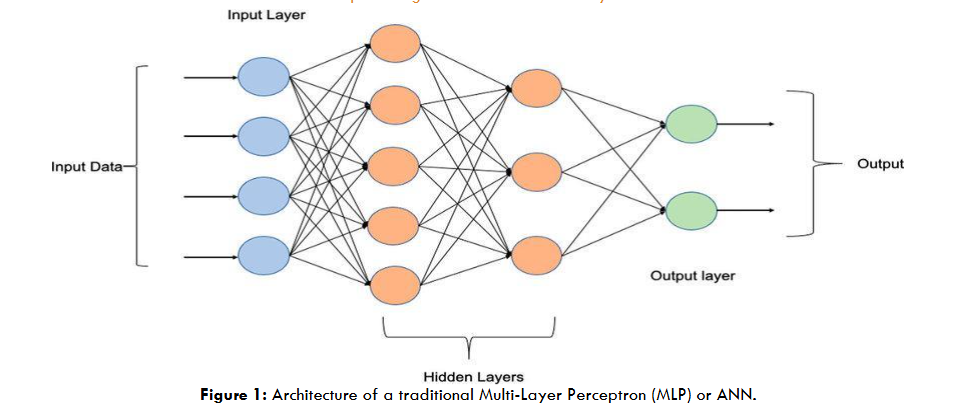
1.4 CONVOLUTIONAL NEURAL NETWORKS
The Convolutional Neural Networks are a family of deep learning models that are specifically designed to process image data. Compared to traditional neural networks, CNNs are able to learn patterns inherent in images more efficiently by incorporating convolutional layers that look at small regions of the image one at a time. Such convolutional layers help the model identify edges, textures, and other important features and then combine those features at deeper layers in order to recognize complex patterns.
The CNNs also employ pooling layers, which scale down the size of an image as it passes through a network. This helps the network to focus on the most important features of the images and enhances efficiency. This makes CNNs particularly useful for medical image analysis, such as classifying chest X-rays, where even tiny differences in texture and structure are important for an accurate diagnosis.
In chest X-ray image analysis, CNNs have achieved high performances comparable to expert radiologists in the detection of pathologies such as pneumonia and tuberculosis. Large, annotated image datasets and transfer learning, where a model is pretrained with a general dataset such as ImageNet and then fine-tuned for a medical task, allow CNNs to automatically learn the patterns of medical images without relying on human expertise explicitly.
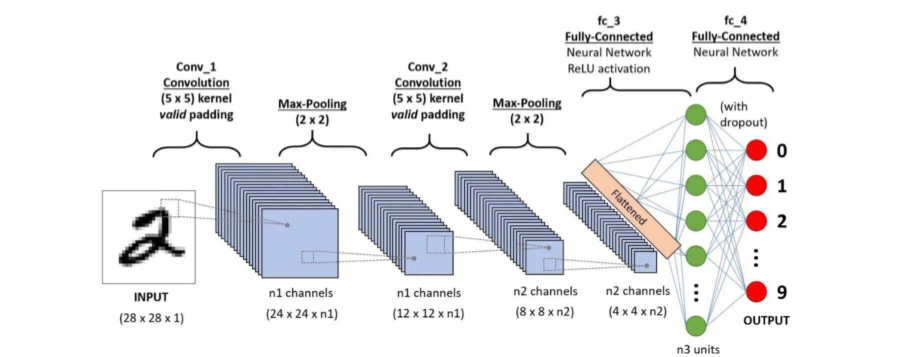
1.5 ENSEMBLE LEARNING IN DEEP LEARNING
Ensemble learning in machine learning refers to the process of combining multiple models for improving the overall performance of a model. Rather than relying on a single model, ensemble methods take advantage of several diverse models, reducing the risk of errors and making better predictions. Ensemble techniques generally come in a few different types:
- Bagging – Bootstrap Aggregating: It involves training multiple models on different random subsets of the data. Their predictions are then combined by averaging. A well-known example is Random Forests.
- Boosting: The models are trained sequentially, each model correcting the errors of the previously trained model. Two very common algorithms of boosting include AdaBoost and Gradient Boosting.
- Stacking: The predictions of several models are combined using another model, called a meta-model, which learns the best way of combining them.
- Voting: Several models make their prediction, and the final result is determined by majority vote (hard voting) or the average of their probabilities (soft voting).
Ensemble methods bear particular usefulness in medical image analysis, considering the problems of data variability and class imbalance. For instance, several different neural networks combined can improve predictive accuracy and provide more reliable results, which is crucial in disease diagnosis from chest X-rays.
1.6 RELATED WORK
Deep learning has been making significant strides in the analysis of chest X-rays, with several studies showing the power of CNNs in detecting such conditions as pneumonia and tuberculosis. A particular example is CheXNet, a deep learning model from Rajpurkar et al. (2017), which attained radiologist-level performance for detecting pneumonia. This inspired further research, including expert-labeled datasets like CheXpert, containing chest X-ray images that enable the evaluation of more advanced models.
Many studies have highlighted the effectiveness of deep learning models for automated medical image classification. Lakhani & Sundaram (2017) demonstrated that deep CNNs were effective in the detection of TB, surpassing traditional methods. Their research on tuberculosis detection illustrated that CNNs could learn complex patterns in radiographs and identify features linked to various diseases, enabling more accurate diagnoses in resource-limited settings.
Pasa et al. (2019) developed efficient deep network architectures for fast chest X-ray tuberculosis screening, demonstrating that deep models could be applied to tuberculosis detection with high accuracy while maintaining computational efficiency. Their model was optimized to run on resource-constrained devices, which is crucial for deployment in low-resource settings. Similarly, Litjens et al. (2021) conducted a comprehensive review of deep learning applications in chest X-ray analysis, concluding that CNNs offer an automated and reliable alternative to traditional image analysis methods, especially for detecting pneumonia and tuberculosis.
Other notable works, such as those by Irvin et al. (2019), further strengthened the case for deep learning in medical image analysis by presenting CheXpert, a large chest radiograph dataset with uncertainty labels and expert comparisons. The dataset enabled researchers to develop more reliable and scalable models for detecting pneumonia, which has since been widely adopted by researchers for evaluating deep learning models.
Furthermore, studies like that of Ahsan et al. (2022) highlighted the importance of using explainable AI (XAI) in medical image analysis to improve interpretability, allowing clinicians to understand the rationale behind model predictions. This is especially important for critical conditions such as tuberculosis, where transparent decision-making is vital for patient safety. Their work underlined the value of XAI in providing explanations for model outputs and fostering trust among medical professionals.
Murphy et al. (2023) showed that deep learning-based detection of active pulmonary tuberculosis at chest radiography can match the clinical performance of radiologists, reinforcing the potential of AI systems in complementing human expertise in diagnosing tuberculosis. Their study also emphasized the potential of AI to identify subtle features that are difficult for even experienced radiologists to discern, which is crucial in detecting early-stage tuberculosis.
The impact of ensemble models in improving medical image classification has also been explored. For example, Baltruschat et al. (2019) demonstrated that ensemble methods, which combine multiple CNN models, could outperform individual models in terms of accuracy and reliability. Their work suggested that ensemble techniques could help mitigate the risk of overfitting and increase model generalization, which is crucial in clinical applications where patient data can vary widely.
Hooda et al (2022) presented a review of recent advances in applying deep learning to tuberculosis screening from chest X-rays over a recent five-year period (2016–2021). They identified and compared methodical contributions, datasets, model architectures (mainly CNN-based), and highlighted both promising results and ongoing challenges such as data heterogeneity, localization beyond simple binary classification, and generalization of models across different settings.
Khan et al. (2023) examined the spatial distribution of tuberculosis cases and abnormal X-rays detected through active case-finding in Karachi, Pakistan, using deep learning models. Their study showed that deep learning models could enhance the efficiency of case-finding strategies, especially in underdeveloped regions where access to skilled radiologists is limited. This research emphasized the need for integrating AI tools into healthcare systems to improve diagnostic accuracy and early detection.
Khan et al. evaluated the diagnostic accuracy of the CAD4TB system for tuberculosis screening using chest radiographs in Karachi’s private healthcare sector. Their study, involving 6,845 individuals with presumptive TB, demonstrated that CAD4TB achieved high sensitivity (65.8–97.3%) and negative predictive value (93.1–98.4%).
Qin et al. (2021) explored the use of deep learning algorithms as a triage test for pulmonary tuberculosis, demonstrating that AI could effectively prioritize patients for further evaluation based on the analysis of chest X-rays. Their study highlighted the potential for AI-based tools to optimize the triage process, especially in high-volume healthcare settings. Similarly, Qin et al. (2023) conducted a systematic literature review on deep learning for tuberculosis detection, summarizing the strengths and limitations of various models used in this field.
Ensemble methods have gained popularity for improving model performance in challenging medical image tasks. For instance, Qin et al. (2023) emphasized the advantages of combining multiple deep learning models, which can address the inherent variability in medical images and lead to more accurate diagnoses. Their review advocated for the widespread use of ensemble models to improve diagnostic outcomes in clinical practice.
Hanson et al (2025). developed a comprehensive framework for integrating artificial intelligence (AI) into radiology practice, aiming to enhance diagnostic accuracy, reduce radiologist burnout, and improve patient outcomes. Their approach emphasized the critical role of human intervention, ethical oversight, and structured workflows to ensure the safe and effective use of AI technologies in clinical settings. The framework was implemented through pilot projects at the University of Miami Miller School of Medicine, evaluating infrastructure, staffing, and financial feasibility. The study demonstrated that the proposed framework facilitated the integration of AI into radiology practice, leading to improved diagnostic precision and workflow efficiency.
This work by testing the leading CNN architectures ResNet50/101, DenseNet121, and EfficientNet-B0/B4 on a dataset containing pneumonia and tuberculosis cases, by applying ensemble techniques supported by rigorous evaluation, this study aims to advance AI applications in chest X-ray analysis and develop systems that can be deployed across diverse healthcare environments.
2. Methods
2.1 DATASET
A curated dataset of chest X-ray images was used, with each image labeled as ‘Normal’, ‘Pneumonia’, or ‘Tuberculosis’. The data was sourced from the publicly available Kaggle dataset “Pneumonia-TB Dataset”, which aggregates images from multiple open-access sources. The dataset is distributed under a permissive license for research use. The images were split into training (90%), validation (10%), and a held-out test set, maintaining class balance through stratified sampling. The final dataset sizes were 10,401 images for training, 1,156 for validation, and 1,284 for testing. Data loaders were configured with a batch size of 32 and shuffling enabled for training.
| Pathology Label | Train | Validation | Test | Total | Percentage |
|---|---|---|---|---|---|
| Normal | 4101 | 469 | 508 | 5078 | 37.3% |
| Pneumonia | 3469 | 372 | 427 | 4268 | 31.0% |
| Tuberculosis | 2831 | 315 | 349 | 3495 | 25.7% |
| Total | 10401 | 1156 | 1284 | 12841 | 100% |
Dataset sizes: Training set = 10,401; Validation set = 1,156; Test set = 1,284.
This table summarizes the dataset composition, highlighting the balance across normal, pneumonia, and tuberculosis cases. Maintaining class balance was essential to ensure fair model training and evaluation.
2.2 PREPROCESSING
To ensure consistency and enhance model generalization, a comprehensive preprocessing pipeline was applied to the chest X-ray images. The following steps were performed:
- Color Conversion: All images were converted to RGB format using a custom ConvertToRGB class to ensure compatibility with models pretrained on ImageNet, which expect three-channel input.
- Resizing: Images were resized to 224×224 pixels to match the input size required by the chosen CNN architectures.
- Data Augmentation (Training Only): To improve generalization and reduce overfitting, several augmentation techniques were applied to the training images:
- Random horizontal flipping (p=0.5)
- Random rotation (up to ±10 degrees)
- Random affine transformations (translation up to 5% in both x and y directions)
- Gaussian blur (kernel size 3, sigma between 0.1 and 2.0)
- Normalization: Pixel values were normalized using the standard ImageNet mean ([0.485, 0.456, 0.406]) and standard deviation ([0.229, 0.224, 0.225]) to align with the pretrained model expectations.
The preprocessing pipelines were as follows:
Training Transformations:
train_transforms = transforms.Compose([
ConvertToRGB(),
transforms.Resize((224, 224)),
transforms.RandomHorizontalFlip(p=0.5),
transforms.RandomRotation(degrees=10)
transforms.RandomAffine(degrees=0,
translate=(0.05, 0.05)),
transforms.GaussianBlur(kernel_size=3, sigma=(0.1,
2.0)),
transforms.ToTensor(),
transforms.Normalize(mean=imagenet_mean,
std=imagenet_std)])Validation/Test Transformations:
test_transforms = transforms.Compose([
ConvertToRGB(),
transforms.Resize((224, 224)),
transforms.ToTensor(),
transforms.Normalize(mean=imagenet_mean,
std=imagenet_std)])These transformations helped standardize the input data and augment the training set, resulting in improved model robustness and performance.
2.3 MODEL ARCHITECTURES
The ensemble comprised five state-of-the-art convolutional neural network architectures: ResNet50, DenseNet121, EfficientNet-B0, EfficientNet-B4, and ResNet101. All models were initialized with ImageNet-pretrained weights. The classifier head of each model was replaced with a custom multi-layer perceptron (MLP) consisting of several fully connected layers with dropout and ReLU activations for regularization. For ResNet and DenseNet, only the final layers were fine-tuned, while for EfficientNet models, the entire classifier block was retrained. Each model was trained independently, and their predictions were later combined in an ensemble.
2.2.1 DenseNet121
DenseNet121 (Densely Connected Convolutional Networks) introduces direct connections from any layer to all subsequent layers, ensuring maximum information flow between layers. Each layer receives inputs from all preceding layers, which encourages feature reuse and mitigates the vanishing gradient problem. This architecture is particularly parameter-efficient and enables deeper networks to be trained effectively. In medical imaging, DenseNet121 excels at capturing fine-grained features, making it suitable for detecting subtle anomalies in chest X-rays.
2.2.2 ResNet50
ResNet50 is a 50-layer deep residual network that utilizes residual connections (or skip connections) to allow gradients to flow directly through the network, addressing the degradation problem in deep networks. These connections enable the model to learn identity mappings, making it easier to train very deep architectures. ResNet50 is known for its robustness and ability to extract hierarchical features, making it effective for medical image classification tasks.
2.2.3 ResNet101
ResNet101 extends the ResNet architecture to 101 layers, further enhancing its capacity to learn complex representations. Like ResNet50, it leverages residual connections to facilitate the training of very deep networks. The increased depth allows ResNet101 to model intricate patterns in medical images, making it highly effective for challenging classification problems such as distinguishing between normal, pneumonia, and tuberculosis cases in chest X-rays.
2.2.4 EfficientNet-B0
EfficientNet-B0 is the baseline model in the EfficientNet family, which scales depth, width, and resolution using a compound coefficient. It employs mobile inverted bottleneck convolution (MBConv) blocks and squeeze-and-excitation optimization, achieving high accuracy with fewer parameters and lower computational cost. EfficientNet-B0 is particularly useful in resource-constrained environments while maintaining strong performance on image classification tasks.
2.2.5 EfficientNet-B4
EfficientNet-B4 is a larger and more powerful variant in the EfficientNet family, offering increased depth, width, and input resolution compared to B0. This allows it to capture more complex patterns and achieve higher accuracy, albeit with greater computational requirements. EfficientNet-B4 is well-suited for tasks where high accuracy is critical and computational resources are available.
2.2.6 Classifier Head Modification Across All Models
To adapt each pretrained backbone for our 3-class chest X-ray classification task, we replaced the original classifier (final fully connected layer) in each model with a custom multi-layer perceptron (MLP) head. The illustration below summarizes the original and custom classifier heads:
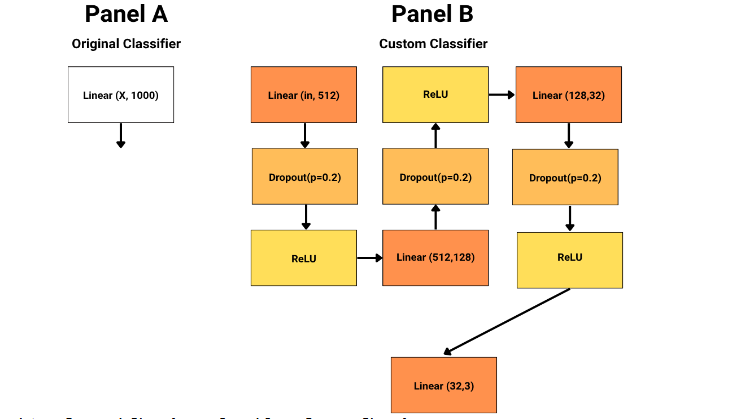
This modification enabled more expressive feature transformation and regularization before the final 3-class output, improving the models’ ability to capture complex patterns relevant to chest X-ray pathology classification.
2.3 TRAINING PROCEDURE
Training was performed using the AdamW optimizer with a learning rate of 1e-4 and weight decay of 1e-4. Cosine annealing learning rate scheduling was used, and early stopping was implemented with a patience of 3 epochs to prevent overfitting. At each epoch, batch-wise accuracy and loss were tracked for both training and validation sets. The best model weights were saved based on validation accuracy. Training was conducted on GPU hardware when available, and all experiments were run with a fixed random seed for reproducibility.
2.4 EVALUATION METRICS AND VISUALIZATION
Model performance was assessed using accuracy, precision, recall, and F1-score for each class, as well as confusion matrices, receiver operating characteristic (ROC) curves, and precision-recall curves. Results were visualized with plots showing training and validation loss and accuracy over epochs, confusion matrices, and ROC and precision-recall curves for each class. A summary comparison of all models’ metrics was also generated to facilitate direct comparison of their performance.
2.5 ENSEMBLE TECHNIQUE
To further enhance classification performance and robustness, we employed an ensemble technique that combines the predictions of all five deep learning models (DenseNet121, ResNet50, ResNet101, EfficientNet-B0, and EfficientNet-B4). Specifically, we used the soft voting mechanism, a probabilistic approach that leverages the confidence scores (softmax probabilities) produced by each model for every class.
Soft Voting Mechanism:
- In soft voting, each model outputs a probability distribution over the possible classes for a given input image. Instead of simply taking a majority vote (hard voting) on the predicted class labels, soft voting aggregates the predicted probabilities from all models and averages them for each class. The final ensemble prediction is the class with the highest average probability.
- Soft voting incorporates the confidence of each model, making the ensemble more sensitive to subtle distinctions between classes.
- It can correct for overconfident or underconfident predictions by any single model, leading to more calibrated and reliable outputs.
- In medical image classification, where misclassification can have serious consequences, soft voting helps reduce the risk of systematic errors and leverages the complementary strengths of diverse architectures.
Soft voting was chosen over hard voting (majority voting) because it better utilizes the full predictive information from each model, rather than just their top choices. This approach has been shown to improve generalization and robustness, especially in challenging, imbalanced, or noisy datasets such as chest X-ray images.
3. Results
3.1 MODEL PERFORMANCE SUMMARY
| Model | Best Validation Accuracy (%) | Test Accuracy (%) | Minimum Validation Loss |
|---|---|---|---|
| EfficientNet-B0 | 98.73 | 97.66 | 0.044 |
| ResNet50 | 98.56 | 97.82 | 0.056 |
| DenseNet121 | 98.56 | 97.74 | 0.049 |
| ResNet101 | 98.48 | 96.88 | 0.043 |
| EfficientNet-B4 | 97.80 | 96.96 | 0.064 |
| Ensemble | – | 98.05 | – |
The ensemble outperforms all individual models on the test set, achieving the highest test accuracy (98.05%).
3.2 PER-CLASS METRICS
| Model | Class | Precision | Recall | F1-score | Support |
|---|---|---|---|---|---|
| DenseNet121 | Normal | 0.98 | 0.96 | 0.97 | 508 |
| DenseNet121 | Pneumonia | 0.96 | 0.98 | 0.97 | 427 |
| DenseNet121 | TB | 1.00 | 0.99 | 0.99 | 349 |
| ResNet50 | Normal | 0.99 | 0.96 | 0.97 | 508 |
| ResNet50 | Pneumonia | 0.95 | 0.99 | 0.97 | 427 |
| ResNet50 | TB | 0.99 | 1.00 | 1.00 | 349 |
| EfficientNet-B0 | Normal | 0.98 | 0.96 | 0.97 | 508 |
| EfficientNet-B0 | Pneumonia | 0.97 | 0.98 | 0.98 | 427 |
| EfficientNet-B0 | TB | 0.98 | 0.99 | 0.99 | 349 |
| ResNet101 | Normal | 0.99 | 0.93 | 0.96 | 508 |
| ResNet101 | Pneumonia | 0.93 | 0.99 | 0.96 | 427 |
| ResNet101 | TB | 0.99 | 1.00 | 0.99 | 349 |
| EfficientNet-B4 | Normal | 0.97 | 0.95 | 0.96 | 508 |
| EfficientNet-B4 | Pneumonia | 0.96 | 0.98 | 0.97 | 427 |
| EfficientNet-B4 | TB | 0.97 | 0.98 | 0.98 | 349 |
| Ensemble | Normal | 0.99 | 0.96 | 0.98 | 508 |
| Ensemble | Pneumonia | 0.97 | 0.99 | 0.98 | 427 |
| Ensemble | TB | 0.99 | 1.00 | 0.99 | 349 |
The ensemble achieves the highest or equal F1-score for all classes, especially for TB (F1 = 0.99), and improves the overall accuracy compared to individual models.
3.3 ENSEMBLE MODEL PLOTS
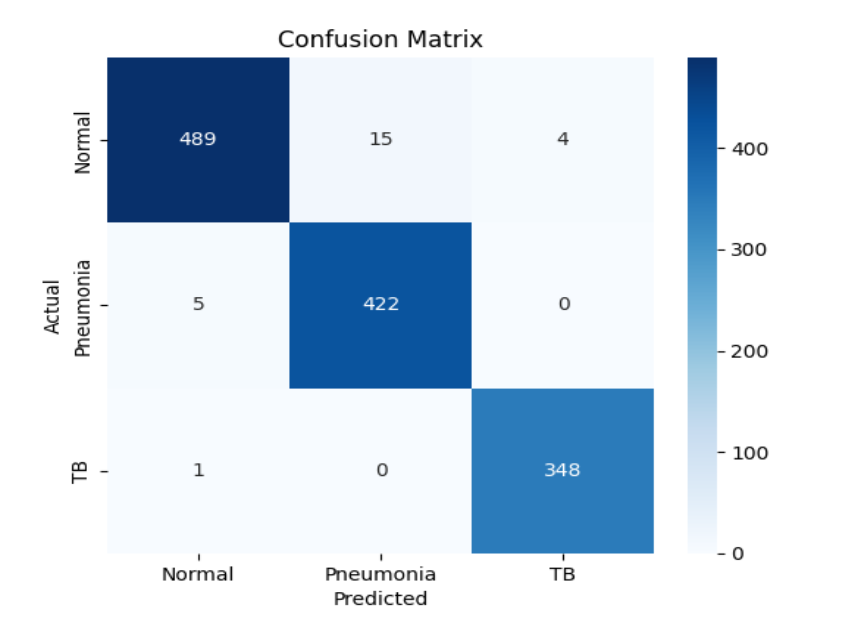
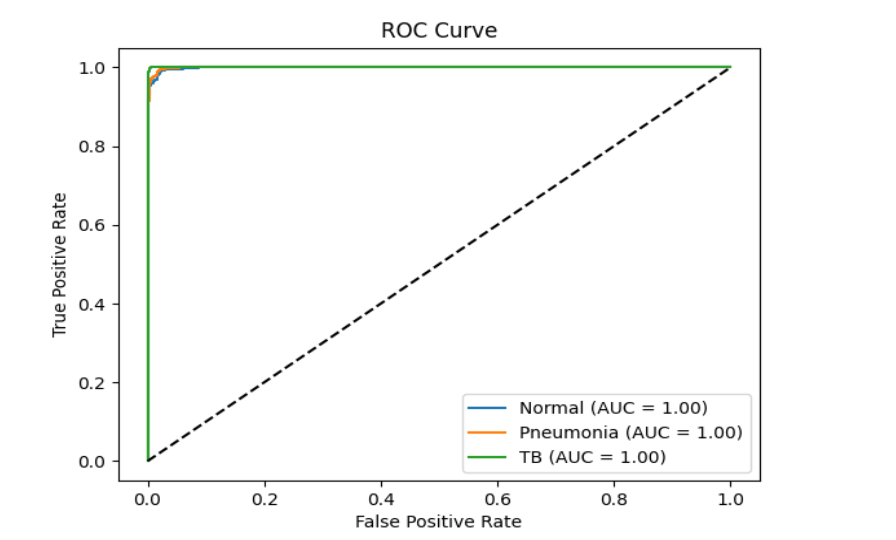
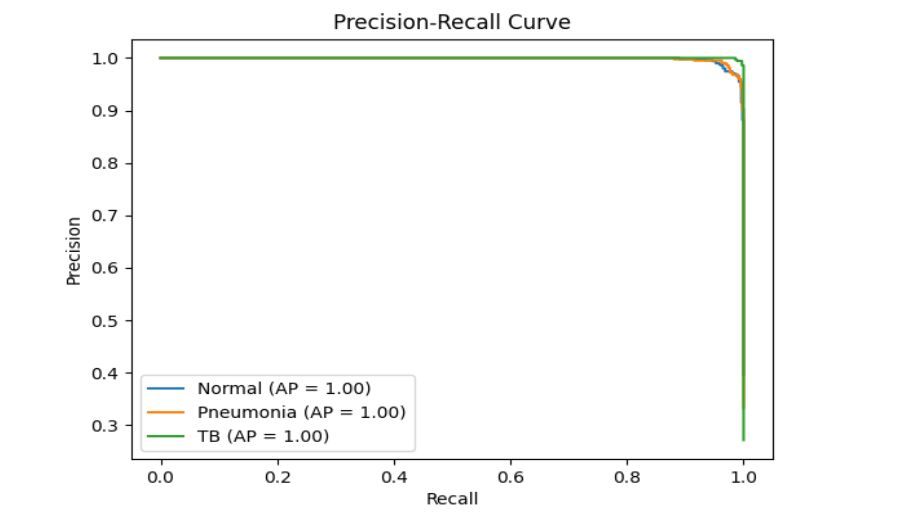
Note: Plots for individual models (DenseNet121, ResNet50, EfficientNet-B0, EfficientNet-B4, ResNet101) are available in their respective model_results/model_name/ directories and in the project repository for detailed comparison.
4. Gradient-weighted Class Activation Visualizations
Gradient-weighted Class Activation Mapping (Grad-CAM) provides an intuitive way to understand what a deep learning model “sees” when making a prediction. By tracing the gradients of the target class back through the final convolutional layer, this method produces a heatmap that visually highlights the regions of an image most influential to the model’s decision. Such visualization helps demystify model behavior, allowing researchers and clinicians to interpret whether the algorithm is attending to clinically meaningful features rather than irrelevant patterns. Below are example Grad-CAM visualizations for chest X-ray images, illustrating the regions that contributed most to the model’s classification:
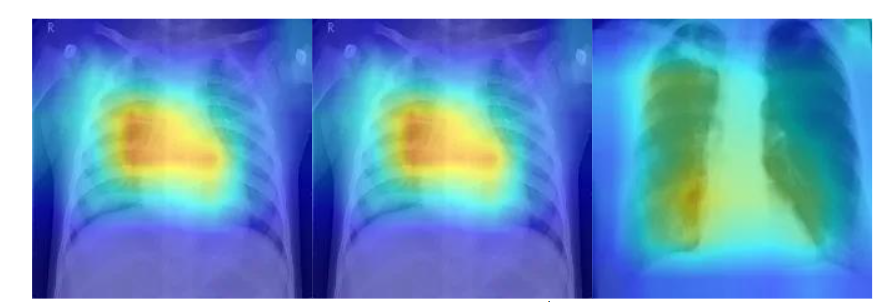
5. Discussion
The ensemble deep learning framework proposed in this study exhibited remarkable performance in classifying chest radiographs into normal, pneumonia, and tuberculosis categories. By merging the strengths of multiple convolutional neural network architectures, the ensemble achieved improved generalization and stability compared to any individual model. Such robustness is essential in medical imaging, where diagnostic consistency can be affected by variations in imaging quality, patient anatomy, or disease presentation.
Overall, the ensemble outperformed all standalone models, yielding the highest test accuracy and a well-balanced performance across disease classes. This outcome aligns with findings from Baltruschat et al., who demonstrated that the integration of multiple CNNs enhances abnormality detection in chest radiographs. The use of a soft voting strategy further refined the results by integrating probabilistic confidence scores from each model, thereby reducing the likelihood of overconfident or erroneous predictions.
A particularly noteworthy finding is the system’s performance in identifying tuberculosis. The ensemble achieved an F1-score of 0.99 for TB detection, indicating exceptional sensitivity and precision. This suggests that the combined model can discern subtle radiographic features that may be difficult to detect even for experienced radiologists, a finding consistent with Murphy et al., who observed radiologist-level accuracy from deep learning models in TB screening.
The success of this approach can be attributed to several factors. The incorporation of transfer learning from ImageNet-pretrained networks provided a strong initialization for feature extraction, while rigorous preprocessing and data augmentation improved resilience to variations in image quality and acquisition conditions. Such design choices are vital for clinical deployment, particularly in settings where data quality and hardware resources may vary significantly.
From a healthcare implementation standpoint, the proposed ensemble system offers a scalable, cost-effective diagnostic support tool. Its ability to identify multiple diseases simultaneously could prove highly beneficial for use in triage systems, mobile diagnostic units, and telemedicine applications, where rapid and reliable interpretation of chest radiographs is often critical.
That said, the model’s complexity introduces practical challenges. While ensemble architectures deliver high accuracy, they also demand greater computational resources and careful integration into existing clinical workflows. Single-model systems are lighter and easier to deploy, though they often sacrifice robustness. Therefore, a balanced approach, prioritizing accuracy without compromising usability, remains essential for successful real-world deployment. The implementation of AI in clinical radiology must continue to account for both technical feasibility and human factors, as discussed in recent frameworks for AI integration.
LIMITATIONS
Despite its promising performance, this study has several limitations. The dataset, although publicly available and diverse, may not fully capture the heterogeneity of real-world clinical environments. Variations in imaging equipment, patient demographics, and annotation quality can all influence generalization. As a result, external validation using data from independent healthcare institutions remains necessary.
Another limitation lies in the absence of clinical trial validation. Although internal test accuracy is high, model behaviour may differ in uncontrolled, real-world conditions. Future work should therefore include prospective studies within hospital workflows to assess the system’s clinical reliability and impact. External validation will be crucial for confirming its generalizability and safety.
Additionally, ensemble models are inherently more computationally demanding than single-model solutions. This may restrict their use in extremely resource-constrained environments such as rural or low-income healthcare settings. Techniques such as model pruning, quantization, or knowledge distillation could mitigate these issues by reducing inference time and memory usage without major losses in accuracy.
Finally, the potential presence of biases, stemming from uneven demographic representation or disease prevalence, poses an ethical challenge. Addressing these biases through balanced data curation and fairness-aware model training will be fundamental for equitable clinical deployment.
FUTURE WORK
Future research should prioritize external validation across diverse populations and healthcare systems to ensure the model’s reliability under varying conditions. Collaborating directly with clinical institutions will help evaluate usability, interpretability, and performance in routine diagnostic workflows. Such real-world studies can also generate valuable feedback from radiologists and practitioners, informing refinements in model design and user interface.
In parallel, further optimization of computational efficiency is warranted. Exploring lightweight ensemble techniques, such as weighted averaging or stacking, could enhance predictive power without excessive computational cost. Addressing potential data biases will also remain a central focus, ensuring that the system provides consistent diagnostic quality across all patient groups and imaging environments.
6. Conclusion
This study underscores the potential of ensemble deep learning for multi-disease chest X-ray classification, demonstrating that combining multiple CNN architectures can significantly enhance diagnostic reliability. The ensemble system achieved superior accuracy compared to individual models and maintained robust performance across varying imaging conditions.
The model’s ability to concurrently detect pneumonia and tuberculosis highlights its clinical relevance, particularly in low-resource settings where expert radiologists are scarce. Through transfer learning, meticulous preprocessing, and a soft voting strategy, the system demonstrated resilience against data variability and class imbalance, key challenges in medical image analysis.
In summary, the ensemble approach presented here represents a practical step toward deploying AI-based radiographic screening tools in real-world healthcare environments. Future directions should focus on external clinical validation, computational optimization, and fairness across diverse populations to ensure safe, effective, and equitable use in global health contexts.
7. Ethical Considerations
Integrating artificial intelligence into clinical imaging requires careful attention to ethical and regulatory issues. Foremost among these is the protection of patient privacy. All datasets must be fully anonymized and handled according to established privacy standards such as GDPR or HIPAA to safeguard sensitive health information.
Bias within training datasets remains another major concern. If data do not represent diverse populations, the resulting models risk producing inequitable outcomes. Ensuring demographic diversity and employing fairness-aware algorithms are therefore essential to avoid diagnostic disparities.
Transparency and interpretability are also critical for fostering clinician trust. Tools such as Grad-CAM, used in this study, can illuminate the decision-making process of deep models, confirming that predictions align with clinically meaningful image regions. Nonetheless, AI should function as a supportive diagnostic assistant rather than a replacement for human expertise. Proper training for medical professionals on how to interpret AI outputs will be key to preventing overreliance or misapplication.
Finally, continued oversight after deployment is vital. Real-world monitoring, feedback mechanisms, and regular performance audits should form part of every AI implementation framework. Ethical use of AI in radiology demands adherence to transparency, accountability, and inclusivity across all stages, from data collection to clinical integration.
8. Data and Code Availability
All code used in this research, including model training scripts, evaluation notebooks, and visualizations, is available in the accompanying project repository. Trained model weights, per-architecture performance metrics, and evaluation plots can be found in the model_results directory. The dataset utilized for this study is derived from publicly accessible chest X-ray sources, with preprocessing and data management procedures documented in detail within the project files.
Conflict of Interest
The authors declare there are no conflicts of interest related to this study.
Funding
This research received no external funding.
References
- World Health Organization. Global Tuberculosis Report 2023. World Health Organization; 2023. Accessed October 25, 2025.
- Litjens G, Kooi T, Bejnordi BE, et al. A survey on deep learning in medical image analysis. Med Image Anal. 2017;42:60–88.
- Rajpurkar P, Irvin J, Ball RL, et al. CheXNet: Radiologist-level pneumonia detection on chest X-rays with deep learning. arXiv preprint arXiv:1711.05225. 2017.
- Baltruschat IM, Nickisch H, Grass M, Knopp T, Saalbach A. Comparison of deep learning approaches for multi-label chest X-ray classification. Sci Rep. 2019;9(1):6381.
- Irvin J, Rajpurkar P, Ko M, et al. CheXpert: A large chest radiograph dataset with uncertainty labels and expert comparison. Proc AAAI Conf Artif Intell. 2019;33:590–597.
- Lakhani P, Sundaram B. Deep learning at chest radiography: Automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology. 2017;284(2):574–582.
- Pasa F, Golkov V, Pfeiffer F, Cremers D, Pfeiffer D. Efficient deep network architectures for fast chest X-ray tuberculosis screening and visualization. Sci Rep. 2019;9(1):6268.
- Litjens G, Kooi T, Bejnordi BE, et al. Deep learning for chest X-ray analysis: A survey. Med Image Anal. 2021;72:102098.
- Ahsan MM, Nazim R, Siddique Z, Huebner P. Explanatory classification of CXR images into COVID-19, pneumonia and tuberculosis using deep learning and XAI. Comput Biol Med. 2022;150:106156.
- Murphy K, Habib SS, Zaidi SMA, et al. Deep learning detection of active pulmonary tuberculosis at chest radiography matched the clinical performance of radiologists. Radiology. 2023;306(1):124–137.
- Hooda R, Mittal A, Sofat S. Advances in deep learning for tuberculosis screening using chest X-rays: The last 5 years review. J Med Syst. 2022;46(12):88.
- Khan SH, Zaidi SMA, Naseer MM. A spatial analysis of TB cases and abnormal X-rays detected through active case-finding in Karachi, Pakistan. Sci Rep. 2023;13(1):1238.
- Khan FA, Majidulla A, Tavaziva G, et al. Evaluation of the diagnostic accuracy of computer-aided detection of tuberculosis on chest radiography among private sector patients in Pakistan. Sci Rep. 2018;8(1):12339.
- Qin ZZ, Ahmed S, Sarker MS, et al. Chest X-ray analysis with deep learning–based software as a triage test for pulmonary tuberculosis: A prospective study of diagnostic accuracy in Karachi, Pakistan. Lancet Digit Health. 2021;3(1):e36–e44.
- Qin ZZ, Naheyan T, Ruhwald M, et al. Machine and deep learning for tuberculosis detection on chest X-rays: Systematic literature review. J Med Internet Res. 2023;25:e43154.
- Hanson C, Jose J, McKinney A, et al. Integrating AI into radiology: A framework for safer, smarter imaging. Am Hosp Healthc Manag. 2025.
- Linguraru MG, et al. Deploying AI in radiology: Expert perspectives on clinical, cultural, and regulatory considerations. Radiol Artif Intell. 2024.
- Aldhafeeri FM. Governing artificial intelligence in radiology: A systematic review of ethical, legal, and regulatory frameworks. Diagnostics. 2025;15(18):2300.
- Yu AC, Mohajer B, Eng J. External validation of deep learning algorithms for radiologic diagnosis: A systematic review. Radiol Artif Intell. 2022;4(3):e210064.
- Acharya V. Ethical frameworks for deploying AI agents in patient care: Balancing innovation and responsibility. Glob Bus Econ J. 2025.
- Na S, Sung YS, Ko Y, et al. Development and validation of an ensemble artificial intelligence model for comprehensive imaging quality check. BMC Med Imaging. 2022;22:815.

