Genomics of SOX13: Insights into Type 1 Diabetes
Genomics of SOX13 gene is dynamic in Type 1 Diabetes
Shouhartha Choudhury1,2,3,
- Har Gobind Khorana School of Life Sciences, Assam University, Silchar-788011, Assam, India
- Department of Biotechnology, Assam University, Silchar-788011, Assam, India
- Department of Life Science and Bioinformatics, Assam University, Silchar-788011, Assam, India
OPEN ACCESS
PUBLISHED:31 July 2025
CITATION: Choudhury, S., 2025. Genomics of SOX13 gene is dynamic in Type 1 Diabetes. Medical Research Archives, [online] 13(7). https://doi.org/10.18103/mra.v13i7.6716
COPYRIGHT © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v13i7.6716
ISSN 2375-1924
ABSTRACT
Background: The SOX13 (SRY-box 13) genes belong to the high mobility group box (HMG-box) family and are primarily associated with testicular concern. This gene has an HMG-box domain that binds with DNA and plays a crucial role in growth and survival. Specifically, SOX13 is located on chromosome 1 and is an insulin-mediated gene. The presence of insulin is significant evidence of genomics variation in the SRY-box transcription factor 13, which is sensitive in type-1 diabetes (T1D). So, the SRY-box transcription factor 13 (ICA12) genes circulate in islets and the exocrine pancreas and are considered a powerful auto-antigen. Thus, beta-cell antigens trigger mechanisms of peripheral tolerance. The recognition of ICA12 as an auto-antigen is an effective marker for autoimmunity.
Aim: The study aims to in-silico analyze the SRY-box 13 gene from the family of SRY-related high mobility group box (HMG-box) genes (SOX gene family) in mammals. So, perform bioinformatics and computational pipelines and applications for experimentation and even upgradation of a particular gene and its family in two different organisms.
Results: The observation documented the total SOX13 and HMG-box domains in two mammalian genomes (i.e. Homo sapiens and Mus musculus). A computational and bioinformatics analysis demonstrated that SOX13 is an insulin-initiated gene. Also, the findings provided genomic variations of the SRY-box 13 gene response in T1D. Also, the study stated the molecular and immunologic mechanisms associated with autoimmune T1D.
Concluding Remarks: The series of data forwarded supreme future of autoimmune T1D are linked with SRY-related HMG-box 13 genes during the outgrowth of organisms. Also, the study justified that HMG-box genes reveal a significant nature during the growth in mammals. Thus, the high mobility group of ICA12 is an insulin-mediated gene that shows a preface in T1D. Also, auto-reactive CD4+ T cells and killer T cells play a fundamental role in autoimmune T1D.
Keywords:
SOX13; SOX family; T1D; Gene therapy; Immunotherapy
Highlights:
- This study is associated with diabetes research and development.
- The present finding illustrated that SOX13 (ICA12) is an insulin-dependent gene.
- The SOX13 gene observed as an auto-antigen is a vital marker for autoimmunity.
- Also, auto-reactive Th cells and TC cells play an immunologic role in autoimmune T1D.
- Therefore, the study leads to the improvement of T1D.
Abbreviation:
WHO: World health organization
IDDM: Insulin-dependent diabetes mellitus (T1D)
NIDDM: Non-insulin-dependent diabetes mellitus (T2D)
T1D: Type 1 Diabetes
T2D: Type 2 Diabetes
CD8+ T-cell: Cytotoxic T-cell
CD4+ Cells: T helper cells
DNA: Deoxyribonucleic acid
LADA: Latent autoimmune diabetes in adults
GAD: Glutamic acid decarboxylase
SOX13: Transcription factor SOX-13
SOX: SRY-related HMG-box
NCBI: National Center for Biotechnology Information
KEGG: Kyoto Encyclopedia of Genes and Genome
SMART: Simple Modular Architecture Research Tool
EMBL: European Molecular Biology Laboratory
NJM: Neighbor-Joining Methods
BLAST: Basic Local Alignment Search Tool
HMM: Hidden Markov Model
GO: Gene Ontology
MEGA: Molecular Evolutionary Genetics Analysis
MEME: Multiple EM for Motif Elicitation
SOX5: Transcription factor SOX-5
CEP85: Centrosomal protein 85
DMRT1: Doublesex and mab-3 related transcription factor 1
PTPRN: Receptor-type tyrosine-protein phosphatase-like N
TCF7L2: Transcription factor 7-like 2
TCF7: Transcription factor 7
TCF7L1: Transcription factor 7-like 1
LEF1: Lymphoid enhancer-binding factor 1
CTNNB1: Catenin beta-1
NOTCH1: Notch homolog 1, translocation-associated
SD: Standard deviation
TF’s: Transcription factors
HMG-box: High mobility group box
HSP60: Heat shock protein 60
IA-2: Islet antigen 2
IGRP: Islet-specific glucose-6-phosphatase catalytic subunit-related protein
β-cell: Beta Cell
CMV: Cytomegalovirus
GAD65: Glutamic acid decarboxylase 65
IFNG: Interferon gamma
TNF-α: Tumor necrosis factor alpha
CD28: Cluster Differentiation 28
Treg: Regulatory T cell
CD25: Cluster Differentiation 25
FOXP3: Forkhead box P3
NF-κB: Nuclear factor kappa-light-chain-enhancer of activated B cells
IL12B: Subunit beta of interleukin 12
IκBα: Nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha
IL-12: Interleukin 12
Introduction:
The primary challenges for computational biologists in the genomic era have been the question of how phenotypic diversity arises. The cutting-edge technology is a feature of the rapid observation of gene products such as gene fragments, homologs, paralogs, and orthologs in particular species. Novel genes may be integrated into living organisms and propose the adaptation of phenotypes. The target is not only the genes, but how, why, and what combinations of a particular gene are expressed at the cellular level is a primordial query in biology. The accessibility of complete draft genome sequences allowed the comparative study between different organisms. So, perform a genome-wide examination of the HMG box-mediated genes in two mammals. The HMG box-mediated genes govern essential functions in determining cell fate during the growth of organisms. The SRY-related HMG-box genes regulate different components of growth (i.e. testis determination, neural induction and determination, lens induction, heart, lymphocyte and thymocyte development, interneuron specification and limp development, cardiac myogenesis, chondrogenesis, vascular development, tissue development, neural crest function, glial maturation, arterial walls development, pancreatic islet development, myogenesis, endoderm specification, hair follicle & vascular growth, CNS patterning and male germ cell maturation). In this outlook, the study aimed to examine the susceptibility of the SOX13 gene from the SRY-related HMG-box family in two mammalian genomes. SRY-box 13 genes integrated into the enormous family of nuclear transcription factors (TFs) are testis determining factor encoded proteins with HMG-box domain bound to DNA. The SOX-13 (ICA12) transcription factor is an auto-antigen in T1D. Autoimmunity is a critical impairment of insulin-mediated beta cells in islets. Specifically, the SRY-box 13 gene commonly produces auto-antibodies in various islet cell components. However, the key auto-antigens with antibodies observed by GAD antibodies are an enzyme that controls the biosynthesis of inhibitory neurotransmitter aminobutyric acid. The GAD antibody is present in 60-70% of diabetes cohorts. Also, other auto-antigens like ICA512-tyrosine hydroxylase, insulin, and carboxypeptidase H circulate a vibration in T1D. The beta-cell antigens reveal a role in type-1 diabetes (T1D) is unclear. Further, auto-antibodies are essential as predictive markers and often a detectable signature of diabetes. However, insulin is substantial evidence of the genetic variation of the nuclear SOX13 transcription factor gene defined as a protective nature in type-1 diabetes: a well-known auto-antibody target in type-1 diabetes. Characteristically, a differentiation in polymorphism situated in the pioneer region on the insulin-dependent SOX13 gene coordinated with type-1 diabetes (T1D). The quantity of repeated units of polymorphism controls the ratio of the insulin-dependent SOX13 gene in a tissue-specific manner. The SOX13 is observed through auto-antigens by a screen using the blood of individuals diagnosed with type-1 diabetes. Therefore, a genome-wide study of the SRY-related HMG-box of TFs is imperative to probe the molecular function and mechanisms involved with SOX13 in T1D. So, the work aimed to analyze the auto-antigen-dependent HMG-box domain of the SOX13 gene in mammals.
Results:
STRUCTURAL ANALYSIS:
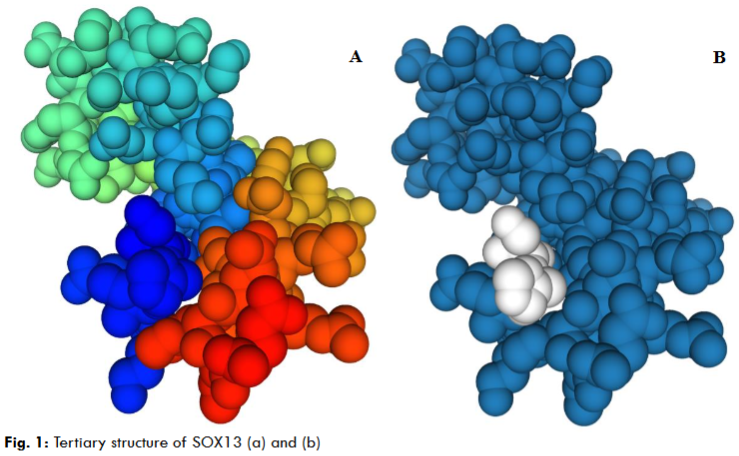
The primary sequence revealed the unique formation of the nucleotides and peptides. So, the sequence composed of 1869 nucleotides and 622 peptides with 71 peptides bound to the DNA sequence is well-known as an HMG-box domain (Table 1). The 3D structure illustrated that the SRY-related HMG-box domain is associated with DNA binding and protein-protein interaction. The formations of the HMG-box residue comprise triple helices in an asymmetrical array.
| (A) Nucleotide |
|---|
| >SOX13 CGCCTCGCAGCAGCGAGCCGCGAGCGCCCTTCTCCAGTCCCGGCTTGGAACTGAACTGTGTGAGCACGGGTCCTGGAACCCGGGCCCAGAACCGGCGAGCCCAGGTCTGAGCCCAGAGCTCAGCGGTCAGCCTCGTAGGCCCTGACTCGGAATCGAGCCGAGGCGCTGAGGTTGGAGCCGGAGAGCGTGAGAGCCGAAGAGCAGGGAGGGCGGGCCGGC TGCGCGTCCGACGAGTCGCAGAGCAGGACCGCGGAAGGCAGGGAGACGGCCGCAAGCCCAGGGCAGAGGG CAGAGGGCAGAGAGCGGCCTGGCTCGGCGGAGAGGGCGCCGCCCGGCCGGAACCAAGCTCGCCGCCCGGG ACGGCGGGCCCCGTGGGGCGCGGACCCAGGGTGGCCGTGGGTCCGCAGCGACTCCCCGGCCGACGGCGGG GGGCGTGCCCCCTCCCAGCCCAGCCTCCCCAACCCGGCCCGCCCGCCGCGTCGCGGGGGCATGTGAGCGGGA AGCCTAGGCTGCCAGCCGCGAGGACCGCACGGAGGAGGAGCAGGAGCGCGGAGCCGCGAGCCCCGAGCCCC GAGCCCGGCGCCTGGCTGAGTAGATGTCCATGAGGAGCCCCATCTCTGCCCAGCTGGCCCTGGATGGCGTTGGCA CCATGGTGAACTGCACCATCAAGTCAGAGGAGAAGAAAGAGCCTTGCCACGAGGCCCCCCAGGGCTCAGCCACTG CC GCTGAACCTCAGCCTGGAGACCCAGCCCGGGCCTCCCAGGATAGTGCTGACCCCCAAGCTCCAGCCCAGGGGA ATTTCAGGGGCTCCTGGGACTGTAGCTCTCCAGAGGGTAATGGGTCCCCAGAACCCAAGAGACCAGGAGTGTCGG AGGCTGCCTCTGGAAGCCAGGAGAAGCTGGACTTCAACCGAAATTTGAAAGAAGTGGTGCCAGCCATAGAGAAGCT GTTGTCCAGTGACTGGAAGGAGAGGTTTCTAGGAAGGAACTCTATGGAAGCCAAAGATGTCAAAGGGACCCAAGA GAGCCTAGCAGAGAAGGAGCTCCAGCTTCTGGTCATGATTCACCAGCTGTCCACCCTGCGGGACCAGCTCCTGACAG CCCACTCGGAGCAGAAGAACATGGCTGCCATGCTGTTTGAGAAGCAGCAGCAGCAGATGGAGCTTGCCCGGCAGC AGCAGGAGCAGATTGCAAAGCAGCAGCAGCAGCTGATTCAGCAGCAGCATAAGATCAACCTCCTTCAGCAGCAGATC CAGCAGGTTAACATGCCTTATGTCATGATCCCAGCCTTCCCCCCAAGCCACCAACCTCTGCCTGTCACCCCTGACTCCCA GCTGGCCTTACCCATTCAGCCCATTCCCTGCAAACCAGTGGAGTATCCGCTGCAGCTGCTGCACAGCCCCCCTGCCCC AGTGGTGAAGAGGCCTGGGGCCATGGCCACCCACCACCCCCTGCAGGAGCCCTCCCAGCCCCTGAACCTCACAGCC AAGCCCAAGGCCCCCGAGCTGCCCAACACCTCCAGCTCCCCAAGCCTGAAGATGAGCAGCTGTGTGCCCCGCCCC CCCAGCCATGGAGGCCCCACGCGGGACCTGCAGTCCAGCCCCCCGAGCCTGCCTCTGGGCTTCCTTGGTGAAGG GGACGCTGTCACCAAAGCCATCCAGGATGCTCGGCAGCTGCTGCACAGCCACAGTGGGGCCTTGGATGGCTCCCC CAACACCCCCTTCCGTAAGGACCTCATCAGCCTGGACTCATCCCCAGCCAAGGAGCGGCTGGAGGACGGCTGTGTG CACCCACTGGAGGAAGCCATGCTGAGCTGCGACATGGATGGCTCCCGCCACTTCCCCGAGTCCCGAAACAGCAGCC ACATCAAGAGGCCCATGAACGCCTTCATGGTGTGGGCCAAGGATGAGCGGAGGAAGATCCTGCAAGCCTTCCCAG ACATGCACAACTCCAGCATCAGCAAGATCCTTGGATCTCGCTGGAAGTCCATGACCAACCAGGAGAAGCAGCCCTACT ATGAGGAACAGGCGCGGCTGAGCCGGCAGCACCTGGAGAAGTATCCTGACTACAAGTACAAGCCGCGGCCCAAG CGCACCTGCATCGTGGAGGGCAAGCGGCTGCGCGTGGGAGAGTACAAGGCCCTGATGAGGACCCGGCGTCAG GATGCCCGC |
| SOX13 |
|---|
| MSMRSPISAQLALDGVGTMVNCTIKSEEKKEPCHEAPQGSATAAEPQPGDPARASQDSAD PQAPAQGNFRGSWDCSSPEGNGSPEPKRPGVSEAASGSQEKLDFNRNLKEVVPAIEKLLS SDWKERFLGRNSMEAKDVKGTQESLAEKELQLLVMIHQLSTLRDQLLTAHSEQKNMAAML FEKQQQQMELARQQQEQIAKQQQQLIQQQHKINLLQQQIQQVNMPYVMIPAFPPSHQPLP VTPDSQLALPIQPIPCKPVEYPLQLLHSPPAPVVKRPGAMATHHPLQEPSQPLNLTAKPK APELPNTSSSPSLKMSSCVPRPPSHGGPTRDLQSSPPSLPLGFLGEGDAVTKAIQDARQL LHSHSGALDGSPNTPFRKDLISLDSSPAKERLEDGCVHPLEEAMLSCDMDGSRHFPESRN SSHIKRPMNAFMVWAKDERRKILQAFPDMHNSSISKILGSRWKSMTNQEKQPYYEEQARL SRQHLEKYPDYKYKPRPKRTCIVEGKRLRVGEYKALMRTRRQDARQSYVIPPQAGQVQMS SSDVLYPRAAGMPLAQPLVEHYVPRSLDPNMPVIVNTCSLREEGEGTDDRHSVADGEMYR YSEDEDSEGEEKSDGELVVLTD |
GENOME-WIDE ANALYSIS:
The genome-wide analysis by HMMER results showed the multiple hits of 163 and 143 of SRY-related HMG-box domain in Homo sapiens and Mus musculus, respectively (Table 2). The standalone BLAST2 results represent 116 and 96 of the SOX13 homologs in Homo sapiens (Humans) and Mus musculus (Mice), respectively (Table 2). The HMMER hits enumerate from both organisms for GO annotation. So, the annotation demonstrated the number of SOX13 genes and SRY-related HMG-box domain in the SOX gene family in Humans and Mice (Table 3). The GO (gene ontology) analysis in Humans suggested the annotation 2291, mean level 8.247, and SD of 3.127. In Mice, annotation 2502, mean level 8.339 and SD of 3.093.
| (A) Summary of the HMG-box domain | Organism | HMMER Hits | BLAST Hits |
|---|---|---|---|
| Homo sapiens | 163 | 116 | |
| Mus musculus | 143 | 96 | |
| Total | 306 | 121 |
| Gene | Homo sapiens | Mus musculus |
|---|---|---|
| SOX-13 | 2 | 3 |
| SOX-5 | 7 | 5 |
| SOX-6 | 5 | 8 |
| SOX-11 | 1 | 1 |
| SOX-14 | 1 | 2 |
| SOX-12 | 1 | 2 |
| SOX-17 | 1 | 6 |
| SOX-4 | 1 | 1 |
| SOX-18 | 1 | 1 |
| SOX-7 | 2 | 1 |
| SOX-15 | 3 | 1 |
| SOX-21 | 1 | 1 |
| SOX-1 | 1 | 1 |
| SOX-3 | 1 | 1 |
| SOX-2 | 1 | 1 |
| SOX-8 | 1 | 3 |
| SOX-10 | 4 | 1 |
| SRY | 1 | 1 |
| SOX-9 | 1 | 1 |
| SOX-30 | 3 | 1 |
| Total | 39 | 42 |
| Group | Group A | Group B | Group C |
|---|---|---|---|
| SoxA | SRY | ||
| SoxB1 | SOX1 | SOX2 | SOX3 |
| SoxB2 | SOX14 | SOX21 | |
| SoxC | SOX4 | SOX11 | SOX12 |
| SoxD | SOX5 | SOX6 | SOX13 |
| SoxE | SOX8 | SOX9 | SOX10 |
| SoxF | SOX7 | SOX17 | SOX18 |
| SoxG | SOX15 | ||
| SoxH | SOX30 |
Table 3: Summary of the Gene Ontology annotation (a) Homo sapiens and (b) Mus musculus
(A) Homo sapiens
| Gene Id | Gene Protein |
|---|---|
| ENSP00000356172.1 | SOX13 transcription factor SOX-13 isoform X1 |
| ENSP00000478239.1 | SOX13 transcription factor SOX-13 isoform X1 |
| ENSP00000379328.2 | SOX5 transcription factor SOX-5 isoform X5 |
| ENSP00000370788.2 | SOX5 transcription factor SOX-5 isoform X5 |
| ENSP00000441973.1 | SOX5 transcription factor SOX-5 isoform X5 |
| ENSP00000439832.1 | SOX5 transcription factor SOX-5 isoform X2 |
| ENSP00000437487.1 | SOX5 transcription factor SOX-5 isoform X3 |
| ENSP00000443520.1 | SOX5 transcription factor SOX-5 isoform X3 |
| ENSP00000398273.2 | SOX5 transcription factor SOX-5 isoform X5 |
| ENSP00000432134.1 | SOX6 transcription factor SOX-6 isoform X12 |
| ENSP00000434455.1 | SOX6 transcription factor SOX-6 isoform X12 |
| ENSP00000324948.6 | SOX6 transcription factor SOX-6 isoform X10 |
| ENSP00000379644.3 | SOX6 transcription factor SOX-6 isoform X10 |
| ENSP00000433233.1 | SOX6 transcription factor SOX-6 isoform X5 |
| ENSP00000322568.3 | SOX11 transcription factor SOX-11 |
| ENSP00000347646.1 | SOX12 transcription factor SOX-12 |
| ENSP00000305343.1 | SOX14 transcription factor SOX-14 |
| ENSP00000297316.4 | SOX17 transcription factor SOX-17 |
| ENSP00000244745.1 | SOX4 transcription factor SOX-4 |
| ENSP00000341815.7 | SOX18 transcription factor SOX-18 |
| ENSP00000301921.1 | SOX7 transcription factor SOX-7 |
| ENSP00000458286.1 | SOX15 protein SOX-15 |
| ENSP00000366144.2 | SOX21 transcription factor SOX-21 |
| ENSP00000355354.2 | SOX15 protein SOX-15 |
| ENSP00000439311.2 | SOX15 protein SOX-15 |
| ENSP00000330218.1 | SOX1 transcription factor SOX-1 |
| ENSP00000359567.2 | SOX3 transcription factor SOX-3 |
| ENSP00000323588.1 | SOX2 transcription factor SOX-2 |
(B) Mus musculus
| Gene Id | Gene Protein |
|---|---|
| ENSMUSP00000122980.1 | SOX13 transcription factor SOX-13 isoform X1 |
| ENSMUSP00000092130.4 | SOX13 transcription factor SOX-13 isoform X1 |
| ENSMUSP00000119729.2 | SOX13 transcription factor SOX-13 isoform X1 |
| ENSMUSP00000107377.1 | SOX5 transcription factor SOX-5 isoform X7 |
| ENSMUSP00000107378.1 | SOX5 transcription factor SOX-5 isoform X7 |
| ENSMUSP00000133041.2 | SOX5 transcription factor SOX-5 isoform X6 |
| ENSMUSP00000076403.5 | SOX5 transcription factor SOX-5 isoform X6 |
| ENSMUSP00000047567.7 | SOX5 transcription factor SOX-5 isoform X1 |
| ENSMUSP00000145919.1 | SOX6 transcription factor SOX-6 isoform X6 |
| ENSMUSP00000126404.1 | SOX6 transcription factor SOX-6 isoform X6 |
| ENSMUSP00000129512.1 | SOX6 transcription factor SOX-6 isoform X6 |
| ENSMUSP00000102223.1 | SOX6 transcription factor SOX-6 isoform X7 |
| ENSMUSP00000145732.1 | SOX6 transcription factor SOX-6 isoform X1 |
| ENSMUSP00000145931.1 | SOX6 transcription factor SOX-6 isoform X1 |
| ENSMUSP00000072583.4 | SOX6 transcription factor SOX-6 isoform X2 |
| ENSMUSP00000129027.1 | SOX6 transcription factor SOX-6 isoform X2 |
| ENSMUSP00000078070.5 | SOX11 transcription factor SOX-11 |
| ENSMUSP00000141674.1 | SOX17 transcription factor SOX-17 isoform X1 |
| ENSMUSP00000142154.1 | SOX17 transcription factor SOX-17 isoform X1 |
| ENSMUSP00000064250.7 | SOX12 transcription factor SOX-12 |
| ENSMUSP00000138293.1 | SOX12 transcription factor SOX-12 |
| ENSMUSP00000088717.1 | SRY sex-determining region Y protein |
| ENSMUSP00000091310.5 | SOX14 transcription factor SOX-14 |
| ENSMUSP00000027035.3 | SOX17 transcription factor SOX-17 isoform X1 |
| ENSMUSP00000112351.2 | SOX17 transcription factor SOX-17 isoform X1 |
| ENSMUSP00000000579.2 | SOX9 transcription factor SOX-9 |
| ENSMUSP00000039466.4 | SOX10 transcription factor SOX-10 |
| ENSMUSP00000133742.1 | SOX8 transcription factor SOX-8 |
| ENSMUSP00000037519.3 | SOX30 transcription factor SOX-30 |
| ENSMUSP00000142204.1 | SOX17 transcription factor SOX-17 isoform X1 |
| ENSMUSP00000142116.1 | SOX17 transcription factor SOX-17 isoform X1 |
DOMAIN, MOTIFS AND PHYLOGENY ANALYSIS:
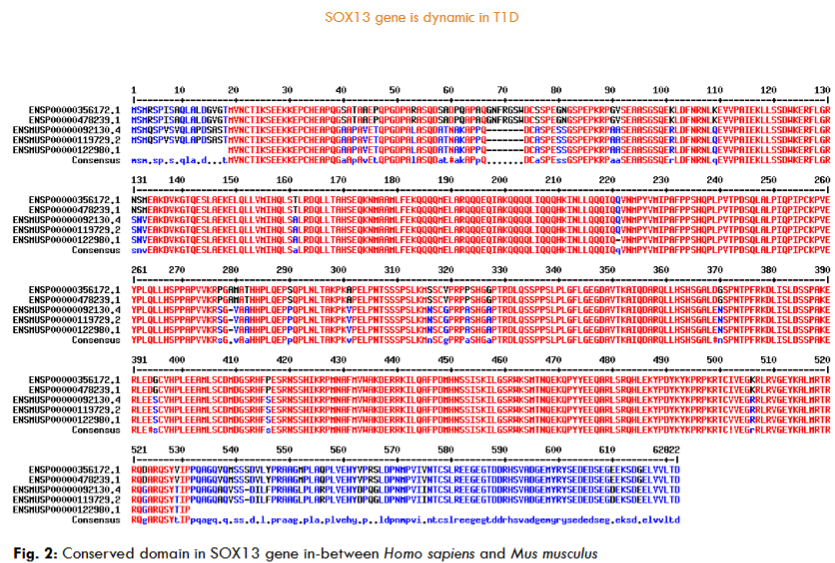
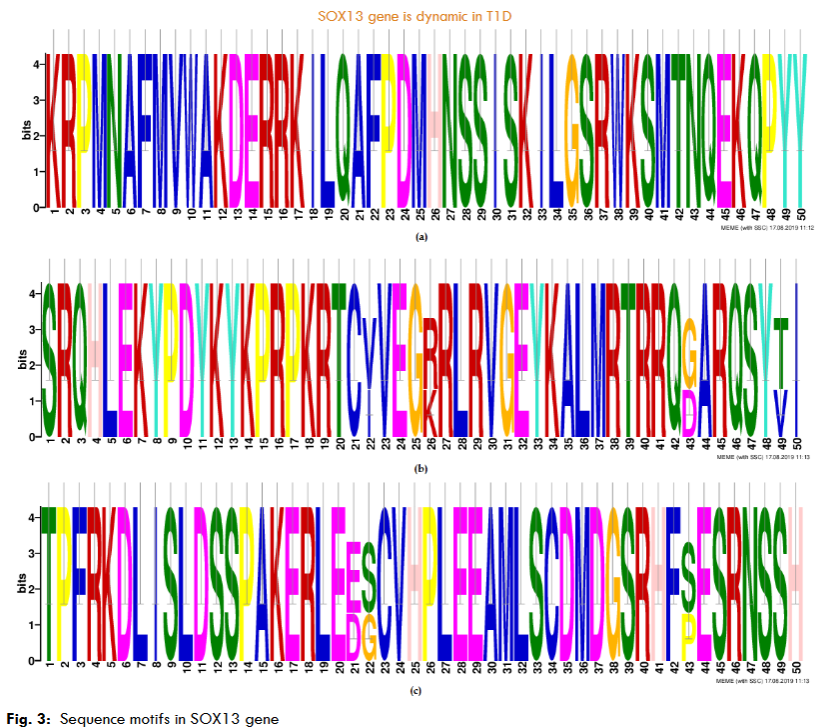
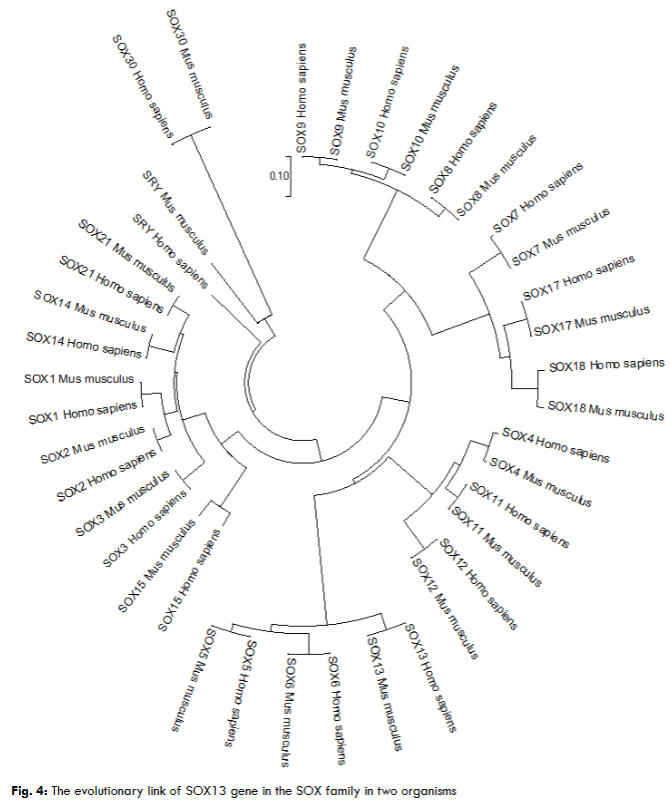
The highest hits of the SRY-box 13 (ICA12) genes selected from both organisms for sequence alignment, a multiple sequence alignment (MSA) determines the conserved domain. The high consensus (90%) indicates the extended HMG-box domain and specific motifs. The phylogenetic tree demonstrated the molecular evolutionary link of the ICA12 gene in Homo sapiens and Mus musculus. Also, a particular clade represents the multifunctional HMG-box domain-associated genes in the SOX family in both organisms’ genomes.
GENE EXPRESSION, CHROMOSOME LOCATION AND GENE REGULATORY NETWORK ANALYSIS:
The expression analysis of the five anatomical parts from the dataset shown of one measurement demonstrated the SOX13 (ICA12) gene critically expressed in the tissue, nervous system, PNS, peripheral nerve, and sural nerve in Homo sapiens.
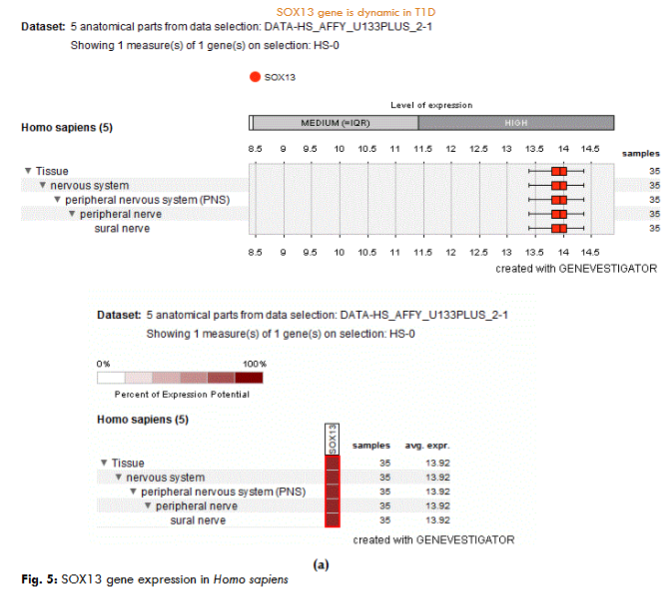
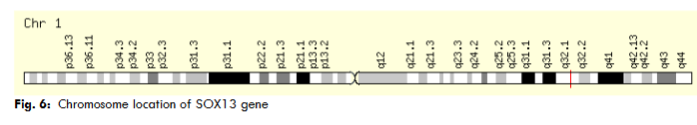
The chromosome localization study confirmed that the ICA12 gene located band 1q32.1 (start at 181,711,925 bp, end at 204,127,743 bp). The gene network study reveals that the SRY-box 13 (SOX13) gene interacts with other molecules, such as PTPRN, TCF7L2, TCF7, TCF7L1, LEF1, CTNNB1, NOTCH1, SOX5, CEP85, DMRT1, those molecular interactions govern the outcome of the ICA12 gene in the cells.
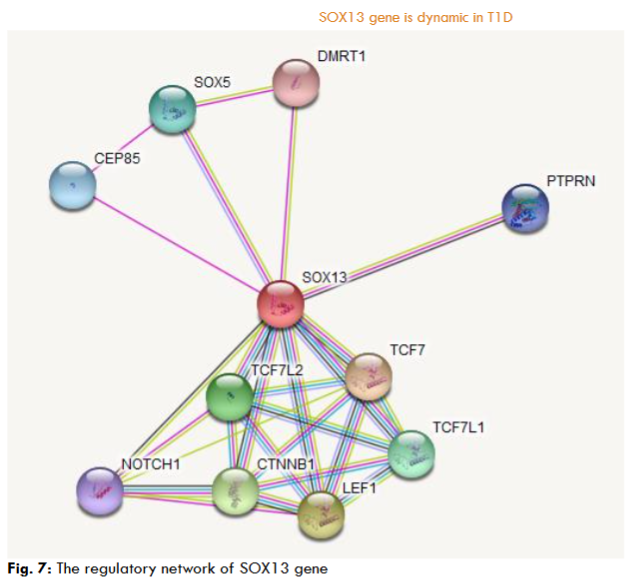
Discussion
Widespread data revealed that IDDM and autoimmunity are generalized by genetic diplomacy and environmental elements. Those factors attributed to autoimmune diseases included extensive variations of chemicals, pathogens, drugs, toxins, diet, stress, viral infection, organ phosphates, heavy metals, and solvents. Other factors contributing to T1D and autoimmunity are weight, puberty, increased linear growth, body mass index, and other parameters of body habits. Known studies suggested the nature of chemicals such as N-nitroso compounds, air pollutants, and persistent organic pollutants in the affected environment prevail over the thread of T1D. Those chemicals in the polluted environment reveal the gene functions and mechanisms linked with the immune system that lead to autoimmunity, and also improvement of T1D. Those factors are common for the enhancement of T1D. New findings will accelerate our understanding of IDDM. A reasonable understanding will interpret a new approach to predict the threat of T1D and validate a unique target to prevent the aggregation of the disease. The WHO classified diabetes into two categories: (a) IDDM (T1D) and (b) NIDDM (T2D) based on the clinical criteria. IDDM is a persistent chronic disease of human correlated autoimmune impairment of the beta-cells in islets. Autoimmune is a typically expended way of self-tolerance defined as a lack of immune reactivity. The feature of IDDM is knowledge of islet autoantigens by auto-reactivity (auto-reactive Th cells and cytotoxic T-cell), either humoral or cellular effectors of the immune system to islet autoantigens (insulin and GAD65). Those mechanisms suggested that Th cells (CD4+ cells) and TC unlock an immunologic role in T1D. Auto-reactive response initiation factors are unclear. However, APCs control specific autoantigens. APCs included DCs (dendritic cells), macrophages, and B lymphocytes in the islets. The autoantigens to naive T lymphocytes by diabetes-associated HLA (human leukocyte antigen) molecules consent the pathogenic T lymphocytes and autoreactive Th cells. These pioneer Th cells generate cytokines and initiate beta-cell-specific CTLs. The initial T cells control islets and stimulate macrophages, and T lymphocytes contribute to control islet-beta cells. So, the defensive IFNG (IFN-γ) and IL-10 manifest by the response of CD4+ T-cells to diabetes-associated antigens and frequencies of autoreactive CD8+ T-cells against HLA-A2-restricted antigenic epitopes. The maturity of diabetes elevated the ratio of autoreactive CD8+ T-cells in the T1D and T2D had analogous autoreactive CD4+ T-cell-derived cytokine response. Further, the HLA have triple residues recognized by (1) class I, (2) class II, and (3) class III molecules depending on the formation of genes. Primary CD8+ T-cells act with islet antigens and stipulated over-expression of class I molecule on beta-cells in T1D. Also, CD4+ autoreactivity exists in diabetes and suggests mere beta-cell stress leads to the autoantigens via endocytic and phagocytic pathways. The prime response of class I molecule-mediated gene products (i.e. HLA-A, HLA-B, and HLA-C) rewarded peptides (endogenous) to counter cytotoxic T cells. Also, class II molecules-initiated genes like HLA-DR, HLA-DP, and HLA-DQ have restricted function and exercise peptides (exogenous) for the account of CD4+ cells. Class III molecule sustains genes encoded for immune-regulated molecules like cachexin (TNF-α), HSPs, and C3, C4, and C5 factors in the robust complement system. Theoretically, class I molecule responds as single-chain peptides that exist in intracellular antigens by killer T cells. Also, class II molecules are heterodimers and expressed mainly on antigen-presenting cells (APCs). Both are intent by alpha and beta (α/β) chains and are subject to presenting extracellular antigens to helper T cells. The dynamic link with T1D in class II molecules contributes to T1D susceptibility. The precise mechanisms by preface of class II molecules pour to the break of islet beta cells are unclear, but the secure characteristics of superior peptides acquired from insulinoma-associated antigen 2 (IA-2), glutamic acid decarboxylase (GAD), proinsulin, and ZnT8 to antigen-presenting cells. Insulin precursor and pre-pro-insulin are targeted auto-antigens for beta cells. Decreased levels of proinsulin influence the response of T lymphocytes in the thymuses and explore the migration of CD4+ and pro-insulin-mediated T lymphocytes to the surface and reveal the T1D. In contrast, the ratio of proinsulin in the thymus enhances the negative (-ve) selection of insulin-specific autoreactive T lymphocytes that leads to immune tolerance and controls the risk of T1D. Further, the immunologic theorem coheres towards autoimmune T1D and CTLA4 molecules. The CD152 is the co-stimulatory molecule on the helper T cell’s surface and binds with B7 ligands that activate CD28 molecules in T lymphocyte co-stimulation. CTLA4 reveals a signal to control T lymphocyte activation by the abundance of Treg. The CTLA4 interacts with a cluster of differentiation 3 (CD3) receptors to initiate the phosphorylation of molecules and directed enhancement of T lymphocytes after binding to HLA molecules via APCs. Further, IL2RA encodes the alpha chain associated with T1D. IL2RA response is observed in memory T lymphocytes, naive T lymphocytes, and activated monocytes. CD25 is key for binding with IL-2 and is manifested as a preface in the growth of regulatory T-cells (Treg). TAC (IL-2R) antigen is controlled by the activity of effector T cells and regulatory cells (Treg) via the response of FOXP3. So, CD25 (P55) may significantly lead to the therapy of autoimmunity. The response of IL2RA on the surface of the Treg is good for regulating T-cell proliferation by response to immunogenic stimulus. Nevertheless, lymphoid tyrosine phosphatase (PTPN22) is a protein tyrosine phosphatase associated with T1D. Lymphoid tyrosine phosphatase (LYP) mainly responds in T cells and is revealed by kinases in the TCR signalling. Lymphoid tyrosine phosphatase interacts with C-terminal Src tyrosine kinase (CSK) by T lymphocyte activation. Further, the small ubiquitin-like modifier 4 (SUMO4) is a major factor in T1D. The reciprocate of methionine to valine (M55V) in IDDM5 reveals an etiological variant linked with T1D. This phenomenon created a potential decrease in sumoylation capacity, higher NF-κB, and release of IL12B. Limited evidence suggests that IDDM5 (SUMO4) sumoylates IκBα and regulates NF-κB transcriptional dynamism. The NF-κB reveals a core regulatory nature in the immune system involved in autoimmune diabetes. Besides, polymorphisms in the classical STAT protein family are linked with T1D. STAT4 from the STAT protein family is found in peripheral blood monocyte cells (PBMCs), dendritic cells (DCs), and macrophages (MΦ) at places of inflammation. STAT4 directly interacts with the IL12RB (CD212) and promotes active roles in IL-12 signalling. IL-12 is an immunoregulatory cytokine in the phenomenon of killer T cells and Th1 cells, supporting the nature of proinflammatory cytokines. However, the TGF-beta, NF-κB, p38 MAPK, toll-like receptor, and interleukin 6 are also linked with autoimmune and inflammation related to T1D. So, the immunologic theorem indicated that the CD4+ T-cell auto-reactivity is linked to T1D/T2D and represents a combined mechanism of beta-cell activity. Therefore, autoreactive CD8+ T-cells appear to T1D and are subject to the accelerated beta-cell mass. Hence, estimating CD8+ autoreactivity in T1D reveals illness conditions and possibly a response to immunotherapy. Many years of investigation illustrated insulin or pro-insulin as the initial auto-antigen. The normal functions of insulin in islet-beta cells make a healthy candidate. We keep in mind that the prime auto-antigen in T1D is unclear. The T1D-associated auto-antigens propose immunologic mechanisms involved with islet beta cells controlled by the immune process. So, the auto-antigens are robust and control of antigen tolerance is associated with immunotherapy for symptom and prognostic markers of T1D. The characteristic of autoimmune T1D is known by the phenomenon of auto-reactive T lymphocytes and auto-antibodies. Antibodies target self-antigens and promote the production of auto-antibodies, which is a characteristic of autoimmunity. However, the autoimmunity does not correspond to autoimmune disorders. Autoimmunity can appear as a complex framework in the immune process and maintain immune homeostasis. However, the quantitative and qualitative variations reveal that natural antibodies and auto-antibodies prevail as pathogenetic autoimmunity. Thus, the observation of auto-antibodies is a condition to consider as an autoimmune nature. Besides, autoimmunity is not allowed as a pathogenetic role in NIDDM (T2D). Etiologically unclear, but environmental factors can contribute to obesity and poor physical activity. Characteristics of NIDDM can support antibodies to self-antigens in the formation of complex antigens. Also, non-insulin-dependent diabetes mellitus refers to LADA or hypothetically slow-onset autoimmune diabetes. So, the dynamism of autoimmune is linked with lymphocyte infiltration of islets and circulating serum antibodies. Also, the markers of autoimmune include auto-antibodies to insulin (IAAs), GAD65, and tyrosine phosphatases (IA2 and IA-2BETA). However, the notable finding in this work is that the beta-cell antigen-dependent genes reveal a major role in T1D. Beta-cell antigens enhance peripheral tolerance mechanisms and act proficiently in diagnosing autoimmune diabetes. The antigen-dependent SOX13 (ICA12) gene homologs of the SOX family play a sole function in T1D. SOX13 gene is not membrane-bound like GAD65 and IA-2 but binds to chromatin. However, the examination of the insulin-dependent SOX13 gene is essential for antigens to get a better understanding of T1D. So, identified beta-cell auto-antigens are necessary for the control of diabetes. The ubiquity of SOX13 observed as an auto-antigen is a molecular signature for autoimmunity. Many studies worldwide demonstrated that auto-antigens are etiopathogenesis of T1D.
Molecular Mimicry
Viral illness is the analytical factor influencing the risk of T1D. So, the characteristics of viruses associated with T1D. Those viruses include rotavirus, mumps virus, coxsackie A virus, coxsackie B virus, Poliovirus, Echovirus, CMV and other viruses. It is unclear whether viruses act as an accelerator in the dynamic immune process initiated by different factors or whether viruses only attack the entire immune process. Generally, the molecular mechanism describing the viral illness observed during the threat of IDDM is molecular complexity. The molecular blueprint of T1D depends on the structural similarity of peptide sequence (conformational structure) between auto-antigen and pathogen. The virus shares a uniform epitope on the islet cell component that processes similar auto-antigens and activates T lymphocytes (T-cells) to encourage a cross-reactive autoimmune response. However, the functional study revealed that the T cells with viral peptides assume the islet auto-antigen is vital to prove the preface of the molecular complexity and improvement of T1D. The major characteristics of IDDM are the knowledge of beta-cell-initiated proteins as auto-antigens by auto-reactive CD4+ and CD8+ T cells and auto-antibodies. These auto-antigens include pro and pre-insulin, GAD65, IA-2, ZnT8, ICAs, Imogen 38, PDX1, CHGA, IGRP, HSP60, and ICA69. So, the autoreactive T lymphocytes are sensitive to beta cell autoantigens like GAD, IA-2, and ZnT8. IGRP and Imogen 38 revealed β-cell autoantigens by the function of pancreatic islet autoreactive Th cells. The complementary DNA subtraction forwards the analytical response of β-cell-initiated proteins. So, the β-cell loss is caused by lymphocytic infiltration of the islet via dendritic cells, macrophages, and T lymphocytes. Empirical studies proposed that Th cells and CTLs reveal a characteristic towards pathogenesis and autoimmune diabetes (T1D). However, the factors initiating auto-reactive responses are unclear. Yet, the specific finding forwarded the auto-antigens recruited by APCs. Also, the APCs include DCs, macrophages, and B lymphocytes in the pancreatic islet cells. In contrast, the auto-antigens to naive T cells by diabetes-associated HLA molecules lead to the priming and activation of pathogenic T lymphocytes and the pre-activation of auto-reactive CD4+ T cells. These activated Th cells produce cytokines and beta-cell-specific cytotoxic T lymphocytes. Auto-antibodies play a unique role in auto-antigen processing by HLA molecules. Numerous studies observed that the HLA alleles are linked with auto-antibodies. The loop between HLA and auto-antibodies still needs further examination to interpret the two significant pathogenic mechanisms. Therefore, it is hard to pinpoint which immunological mechanisms initiate viral illness and autoimmunity.
Methods:
DATABASE AND PRIMARY SEQUENCE:
The objective (query) sequence is retrieved from various specific databases such as Universal Protein Resource (UniProt), National Center for Biotechnology Information (NCBI), GenBank, European Molecular Biology Laboratory (EMBL), Kyoto Encyclopedia of Genes and Genomes (KEGG), and DNA Data Bank of Japan (DDBJ). Web-based application of Simple Modular Architecture Research Tool (SMART) performs the examinations of the particular peptide residues in objective sequence (query or suspected sequence). The SWISS-MODEL database retrieves for prediction of amino acids (peptides) structure. The above database is a bioinformatics web server for comparative modelling of the structure of protein molecules. This database generates a 3D structure utilized in different effective research applications. The Swiss model is a regularly revised database of remodelling of organism proteome for biological research.
GENOME SEQUENCE
The organism’s genome sequences are downloaded from various specialized databases:
1. Ensembl Genomes.
2. NCBI.
ORGANISMS:
1. Homo sapiens: Genome assembly: GRCh38.p13 (GCA_000001405.28)
2. Mus musculus: Genome assembly: GRCm39 (GCA_000001635.9)
STANDALONE TOOLS
The HMM ER software packages execute through MSA methods for the objective domain as a profile search (parameters: 1.0e-3). The above mathematical and statistical algorithms are assembled by an MSA of the query sequence residue for profile search. The standalone basic local alignment search tool 2 (BLAST2) executes for homolog genes in both organisms.
GENE ONTOLOGY ANNOTATION
Gene ontology annotation is a method of the functional analysis of the genes in the genome across species for biological variance. So, the Omics box initializes using parameters 1.0e-3 for GO (gene ontology) annotation. The functional property of genes rectified via gene ontology annotation is a popular tool for practical work.
DOMAIN
Sequence domain analysis is a core component for the upgradation of conserved residue in two different organisms’ genomes. So, MSA methods are performed using a web-based application of MultAlin for analysis of the conserved region in sequences between both organisms. The MSA method calculates the specific pair of the homologous sequences and steak them up. Then observe the identical, differences, and similarities in sequences. The highest hits sequences are applied for MSA to upgrade the sustain domain.
MOTIFS
The motif-based sequence analysis tools retrieve for resolution of sequence-specific motifs. MEME suite is a bioinformatics and computational web-based tool to analyze and validate particular motifs in the sequence.
PHYLOGENY
An analysis of phylogeny in genomes is necessary to explore the molecular Darwinism link between genes in both organisms. Therefore, MEGAX practice for constructing an evolutionary tree using Neighbor-Joining Methods.
GENE EXPRESSION
The gene expression analysis is elicited by the genevestigator application. The genevestigator is a high-performance research engine for expression analysis of an organism’s gene contents. The genevestigator initializes to identify and validate specific targets.
CHROMOSOMAL LOCATION
The chromosomal location is retrieved through a gene card web-based application. The gene card is a database of an organism’s genes that provides knowledge on all recognized and predicted genes. The above database is updated and available for biomedical research.
GENE REGULATORY NETWORK
The gene regulatory matrix reveals a molecular interaction in the cellular process to dominate the volume of mRNA or proteins. Some proteins act to activate genes as the TFs bind to the promoter area and initiate the response of different proteins known as regulatory cascades.
Declaration:
CONSENT FOR PUBLICATION:
The work furnished in this paper is original and communicated by the correspondent addressed in the manuscript. The author disclosed that the documents are not concerned elsewhere and have not been received for evaluation by other journals.
ETHICAL APPROVAL:
The study contains an in-silico analysis of the mammalian genome examination and upgradation of the particular gene in different organisms.
AVAILABILITY OF DATA AND MATERIALS:
The data and materials are available on reasonable request.
COMPETING OF INTERESTS:
The author declared that the work has no conflict of interest.
FUNDING:
The author did not avail of financial assistance from any source in undertaking the present study.
AUTHOR CONTRIBUTIONS:
This research paper contains a single author. SC proposed the idea, experimented, analyzed data and prepared the manuscript.
AUTHOR DETAILS:
The author is a PhD scholar in the Har Gobind Khorana School of Life Sciences, Department of Biotechnology, Assam University, Silchar-788011, Assam, India.
ACKNOWLEDGEMENT:
The author was grateful to the editors and reviewers from the European Society of Medicine for valuable comments and suggestions during the evaluation. Also, the author is thankful to Assam University, Silchar, Assam, India, for providing the lab facilities during the research work.
References:
- Roose, J. et al. High expression of the HMG box factor sox-13 in arterial walls during embryonic development. Nucleic acids research. 1998, 26(2):469-476.
- Roose, J. et al. The Sox-13 Gene: Structure, Promoter Characterization, and Chromosomal Localization. Genomics. 1999, 57(2):301-305.
- Kasimiotis H. et al. Sex-determining region Y-related protein SOX13 is a diabetes autoantigen expressed in pancreatic islets. Diabetes. 2000, 49(4): 555-561.
- Schepers, G.E., R.D. Teasdale, and P. Koopman. Twenty pairs of sox: extent, homology, and nomenclature of the mouse and human sox transcription factor gene families. Developmental cell, 2002. 3(2): p. 167-170.
- Graves, J.A.M. Two uses for old SOX. Nature genetics. 1997, 16(2):114.
- Fida S et al. Autoantibodies to the transcriptional factor SOX13 in primary biliary cirrhosis compared with other diseases. Journal of autoimmunity. 2002, 19(4):251-257.
- Kasimiotis H et al. Antibodies to SOX13 (ICA12) are associated with type 1 diabetes. Autoimmunity. 2001, 33(2):95-101.
- Argentaro A et al. Linkage studies of SOX13, the ICA12 autoantigen gene, in families with type 1 diabetes. Molecular genetics and metabolism. 2001, 72(4):356-359.
- Bieg S, J. Seissler, and W.A. Scherbaum. Cytoplasmic islet cell antibodies and GAD antibodies in diagnosis of diabetes. Medizinische Klinik (Munich, Germany: 1983). 1995, 90(10):594-598.
- Wilkin T.J. Autoantibodies as mechanisms, markers, and mediators of B‐cell disease. Diabetes/metabolism reviews. 1991, 7(2):105-120.
- Lucassen A.M. et al. Susceptibility to insulin dependent diabetes mellitus maps to a 4.1 kb segment of DNA spanning the insulin gene and associated VNTR. Nature genetics. 1993, 4(3):305.
- Rabin D.U. et al. Cloning and expression of IDDM-specific human autoantigens. Diabetes. 1992, 41(2):183-186.
- Wildner G, Kaufmann U. What causes relapses of autoimmune diseases? The etiological role of autoreactive T cells. Autoimmun Rev. 2013, 12 (11):1070-1075
- Sarikonda G, Pettus J, Phatak S, Sachithanantham S, Miller JF, Wesley JD et al. CD8 T-cell reactivity to islet antigens is unique to type 1 while CD4 T-cell reactivity exists in both type 1 and type 2 diabetes. J Autoimmun. 2004, 50:77-82
- Han S, Donelan W, Wang H, Reeves W, Yang LJ. Novel autoantigens in type 1 diabetes. AmJ Transl Res. 2013, 5(4):379-392
- Cervin C, Lyssenko V, Bakhtadze E, Lindholm E, Nilsson P, Tuomi T, Cilio M C, Groop. Genetic similarities between latent autoimmune diabetes in adults, type 1 diabetes, and type 2 diabetes. Diabetes. 2008. 57(5):1433-1437
- Ounissi-Benkalha H, Polychronakos C. The molecular genetics of type 1 diabetes: new genes and emerging mechanisms. Trends Mol Med. 2008. 14(6):268-275
- van Belle TL, Coppieters K T, von Herrath MG. Type 1 diabetes: etiology, immunology, and therapeutic strategies. Physiol Rev. 2011. 91(1):79-118
- Pociot F, Akolkar B, Concannon P, Erlich HA, Julier C, Morahan G. Nierras RC, Todd AJ, Rich SS, and Nerup J. Genetics of type 1 diabetes: what’s next? Diabetes. 2010. 59(7):1561-1571
- Vafiadis P, Ounissi-BenkalhaH, PalumboM,Grabs R, Rousseau M, Goodyer CG. Polychronakos C. Class III alleles of the variable number of tandem repeat insulin polymorphism associated with silencing of thymic insulin predispose to type 1 diabetes. J Clin Endocrinol Metab. 2001. 86(8):3705-3710
- Nistico L, Buzzetti R, Pritchard LE, Van der Auwera B, Giovannini C, Bosi E. Larrad TM, Rios SM, Chow CC, Cockram SC, Jacobs K, Mijovic C, Bain CS, Barnett HA, Vandewalle LS, Schuit F, Gorus KF, Tosi R, Pozzilli P, and Todd AJ. The CTLA-4 gene region of chromosome 2q33 is linked to, and associated with, type 1 diabetes. Belgian Diabetes Registry. Hum Mol Genet. 1996. 5(7):1075-1080
- Ueda H, Howson JM, Esposito L, Heward J, Snook H, Chamberlain G et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature. 2003. 423(6939):506-511
- Wing K, Onishi Y, Prieto-Martin P, Yamaguchi T, Miyara M, Fehervari Z. Nomura T, Sakaguchi S. CTLA-4 control over Foxp3+ regulatory T cell function. Science. 2008. 322(5899):271-275
- Corthay A. How do regulatory T cells work? Scand J Immunol. 2009. 70(4):326-336
- Mehers KL, Gillespie KM. The genetic basis for type 1 diabetes. Br Med Bull. 2008. 88(1):115-129
- Askenasy N. Enhanced killing activity of regulatory T cells ameliorates inflammation and autoimmunity. Autoimmun Rev. 2013. 12(10): 972-975
- Smyth D, Cooper JD, Collins JE, Heward JM, Franklyn JA, and Howson JM. Replication of an association between the lymphoid tyrosine phosphatase locus (LYP/PTPN22) with type 1 diabetes, and evidence for its role as a general autoimmunity locus. Diabetes. 2004. 53(11):3020-3023
- Gregersen PK and Behrens TW. Genetics of autoimmune diseases-disorders of immune homeostasis. Nat Rev Genet. 2006. 7(12):917-928
- Guo D, Li M, Zhang Y, Yang P, Eckenrode S, and Hopkins D. A functional variant of SUMO4, a new I kappa B alpha modifier, is associated with type 1 diabetes. Nat Genet. 2004. 36(8):837-841
- Guo D, Han J, Adam BL, Colburn NH, Wang MH, and Dong Z. Proteomic analysis of SUMO4 substrates in HEK293 cells under serum starvation-induced stress. Biochem Biophys Res Commun. 2005. 337(4):1308-1318
- Caamano J and Hunter CA. NF-kappaB family of transcription factors: central regulators of innate and adaptive immune functions. Clin Microbiol Rev. 2002. 15(3):414-429
- Lee HS, Park H, Yang S, Kim D, and Park Y. STAT4 polymorphism is associated with early-onset type 1 diabetes, but not with late-onset type 1 diabetes. Ann N Y Acad Sci. 2008. 1150:93-98
- Frucht DM, Aringer M, Galon J, Danning C, Brown M, Fan S. Stat4 is expressed in activated peripheral blood monocytes, dendritic cells, and macrophages at sites of Th1-mediated inflammation. J Immunol. 2000. 164(9):4659-4664
- Yang Z, Chen M, Ellett JD, Fialkow LB, Carter JD, and McDuffie M. Autoimmune diabetes is blocked in Stat4-deficient mice. J Autoimmun. 2004. 22(3):191-200
- Trinchieri G. Interleukin-12: a cytokine produced by antigen-presenting cells with immunoregulatory functions in the generation of T-helper cells type 1 and cytotoxic lymphocytes. Blood. 1994. 84(12):4008-4027
- Lin J, Zhou ZG,Wang JP, Zhang C, Huang G. From Type 1, through LADA, to type 2 diabetes: a continuous spectrum? Ann N YAcad Sci. 2008. 1150:99-102
- Organization, W.H., Definition, diagnosis and classification of diabetes mellitus and its complications: report of a WHO consultation. Part 1, Diagnosis and classification of diabetes mellitus. 1999, Geneva: World health organization.
- SHTAUVERE‐BRAMEUS, A., et al., Antibodies to new beta cell antigen ICA12 in Latvian diabetes patients. Annals of the New York Academy of Sciences. 2002, 958(1):297-304.
- Elkon, K. and P. Casali, Nature and functions of autoantibodies. Nature Reviews Rheumatology.2008, 4(9):491.
- Bosi, E. and E. Bonifacio. Autoantibodies in insulin-dependent diabetes mellitus. Journal of endocrinological investigation. 1994, 17(7):521-531.
- Pi-Sunyer, F.X. Medical hazards of obesity. Annals of internal medicine, 1993, 119:655-660.
- Willis J.A. et al. Islet cell antibodies and antibodies against glutamic acid decarboxylase in newly diagnosed adult-onset diabetes mellitus. Diabetes research and clinical practice. 1996,33(2):89-97.
- Lernmark Å. Glutamic acid decarboxylase–gene to antigen to disease. Journal of internal medicine. 1996, 240(5):259-277.
- Bonifacio E. et al. Identification of protein tyrosine phosphatase-like IA2 (islet cell antigen 512) as the




