Helicase-Like Transcription Factor in Colorectal Cancer
Helicase-like transcription factor at the intersection of replication independent non-coding RNAs and histone subtype in a cell-line derived xenograft model of colorectal cancer
Rebecca A Helmer1,2 and Beverly S. Chilton1*
- Department of Cell Biology & Biochemistry, Texas Tech University Health Sciences Center, Lubbock, Texas 79430
- Current address: Garrison Independent School District, Garrison, Texas 75946
OPEN ACCESS
PUBLISHED 31 October 2025
CITATION Helmer, R.A., Chilton, B.S., 2025. Helicase-like transcription factor at the intersection of replication independent non-coding RNAs and histone subtype in a cell-line derived xenograft model of colorectal cancer. Medical Research Archives, [online] 13(10). https://doi.org/10.18103/mra.v13i10.7004
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v13i10.7004
ISSN 2375-1924
ABSTRACT
New strategies are desperately needed for the treatment of metastatic colorectal cancer. Early diagnostic techniques have improved the survival rate for individuals with early stage disease; however, the five-year survival rate for patients with metastatic disease is less than 10%. Understanding the mechanisms responsible for disease progression and metastasis is a critical need for these patients. Human helicase-like transcription factor is a tumor suppressor epigenetically silenced during the progression of colorectal cancer. To examine how exactly does silencing HLTF contribute to tumorigenesis, we developed a cell-line derived xenograft model to compare the extremes of disease development. We compared the initial phase in which cancer cells that express helicase-like transcription factor are located in a tumor microenvironment where helicase-like transcription factor is absent vs the final metastatic phase in which neither the cancer cells nor the cells of the tumor microenvironment express helicase-like transcription factor. Combinatorial use of RNA sequencing technology with species-specific mapping, spatial transcriptomics, immunohistochemistry and mass spectrometry provided a holistic understanding of glutathione-based antioxidant defense in the model. The collective re-evaluation of the data revealed changes in replication independent non-coding RNAs and a histone subtype previously unreported.
Keywords: Colorectal cancer (CRC); helicase-like transcription factor (HLTF); SWI-SNF-related, Matrix-Associated, Actin-dependent regulator of chromatin, Subfamily A, Member 3 (SMARCA3); tumor microenvironment (TME); cell-line derived xenograft (CDX), non-coding RNA U2, snoRNA U3, SNORD3A
INTRODUCTION
The increased incidence for six of the top 10 cancers (breast, prostate, melanoma, endometrial, pancreas and colorectal) shows the burden of cancer is shifting from older to younger adults(1). The fact that metastatic colorectal cancer (CRC) has increased in people younger than 55 years of age(2) underscores the need for better prevention based on a better understanding of the mechanism(s) of disease progression.
Genome integrity is protected by the tumor suppressor helicase-like transcription factor (HLTF) also known as SMARCA3. HLTF — a yeast RAD5 homolog with E3 ubiquitin ligase and DNA helicase activities — is a member of the SWI/SNF family of ATP-dependent chromatin remodelers(3,4) inactivated by promoter hypermethylation in 43% of primary colorectal tumors(5). The methylation status of HLTF DNA in serum and stool samples is significantly correlated with tumor size and advanced stage (III or IV) metastatic disease and shorter survival(6-8).
The HLTF gene is alternatively spliced. Its full-length mRNA encodes a multifunctional protein composed of distinct domains(9), i.e. a nuclear localization signal containing an embedded HIRAN (HIP116 and RAD5 N-terminal) domain, seven (I, Ia, II-VI) motifs split by the insertion of a C3HC4-type zinc finger or RING (Really interesting new gene) finger motif characteristic of E3 ubiquitin ligases into an N-terminal DExx module (I, Ia, II and III) for ATP binding/hydrolysis, and a C-terminal HELICc module (IV, V, and VI) for dsDNA translocase, and a DNA repair domain.
HLTF was first identified as a transcription factor by recognition site screenings. The sequence motif (C/A-C-T/A-T-A/T/G-T/G) was identified by cyclic amplification and selection of targets (CASTing) and authenticated with gel shift assays(13). Structure/function studies showed HLTF mediates genome stability and repair pathway choice via its RING-domain(14). At stalled replication forks, DNA damage is mitigated(15) via translesion DNA synthesis (TLS), template switching (TS) or replication fork repriming (RFR). Proliferating-cell nuclear antigen (PCNA), a sliding clamp for DNA polymerases maintains genomic stability during DNA replication.
PCNA recruits proteins repair proteins and can be functionally modified by monoubiquitination, polyubiquitination and SUMOylation(15). TLS and TS are stimulated by mono- and polyubiquitination of PCNA, respectively. In the presence of HLTF, PCNA monoubiquitination is followed by polyubiquitination that is obligatory for TS. In the absence of HLTF, monoubiquitinated PCNA activates the TLS pathway. TS is initiated by a strand invasion-dependent mechanism that requires HLTF’s ATPase domain. RFR and nucleotide excision repair (NER) are coordinated via HLTF’s ATP-dependent dsDNA translocase activity and its HIRAN domain(16-18).
HLTF is a histone H3K23-specific E3 ubiquitin ligase(19) that stabilizes H3K9me3 (transcriptional repression). Thus silencing HLTF resulted in upregulation of cancer-promoting genes. Employing a panel of mutants targeting key functional domains allowed Akano et al. (19) to show the HIRAN, ATPase and RING domains were essential for E3 ligase function in this epigenetic mechanism. In CRISPR-Cas9-mediated genome editing(20), HLTF engages its HIRAN domain and its translocase activity to enable downstream processing and repair. HLTF regulates the residency time of Cas9 at the DNA break site potentially affecting repair pathway choice.
Elevated transcription is associated with guanine-rich DNA sequences that can fold into four-stranded, noncanonical secondary structures known as G-quadruplexes(21). DNA methyltransferase enzymes (DNMTs) bind to G4 structures whose accumulation can cause DNA replication stress resulting in DNA double-strand breaks(21). HLTF suppresses G4 accumulation in an ATPase-dependent manner(22).
DNA trinucleotide repeat (TNR) disorders are mainly neuropsychiatric conditions caused by abnormal expansion of repetitive sequences leading to replicative stress. The HIRAN domain of HLTF preferentially recognizes the (T/C)TG-3′-OH nucleotide sequence in CTG TNR regions that are susceptible to replication errors(23). Thus HLTF helps to maintain the stability of difficult to replicate TNR microsatellite regions.
In early stage CRC, HLTF is expressed in tumor cells but not in the tumor microenvironment (TME). With disease progression, an ecosystem is established in which non-malignant cells of the TME devoid of HLTF expression interact with CRC cells in which HLTF hypermethylation causes HLTF loss of function. We developed a cell line-derived xenograft (CDX) model to show for the first time that HLTF-deletion in cancer cells and the TME results in altered expression of replication independent non-coding RNAs commensurate with a histone variant capable of reshaping the gene expression landscape.
MATERIALS AND METHODS
Detailed methodology for this work was published in the materials and methods section of Martinez-Marin et al.(24) and independently in protocols.io (doi.org/10.17504/protocols.io.ewov1oe37lr2/v1).
Hltf-deleted mice
Global Hltf knockout (KO) mice were developed in collaboration with genOway (Lyon, France) and bred (IACUC# 02007) into the recombinase activating gene 2 (Rag2)/common gamma (IL2rg) double knockout background to generate immunodeficient (ID) Hltf KO mice >99% C57BL/6 background genome. For the CDX model, primary tumors were established by direct orthotopic microinjection of HLTF+/+HCT116-Red-FLuc or HLTF-/-HCT116-Red-FLuc cells between the mucosa and the muscularis layers of the cecal wall of ID male Rag2-/-IL2rg-/- Hltf KO mice.
RNAseq and Spatial transcriptomics
RNAseq data for HLTF-/-CDX in ID Hltf KO are accessible through NCBI’s Gene Expression Omnibus (GEO) SuperSeries accession number GSE234728, SubSeries GSE234725 (RNA-Seq). RNAseq data for HLTF+/+CDX in ID Hltf KO are accessible through GSE161961. Spatial transcriptomics data for HLTF-/- and HLTF+/+CDX tumors in ID Hltf KO mice are accessible through NCBI’s Gene Expression Omnibus (GEO) SuperSeries accession number GSE234728, SubSeries GSE234517 (Spatial).
Statistical analysis
For this study. Bioconductor – DESeq2 software was used for a comparison of gene expression between the HLTF-/- and HLTF+/+CDX tumors and TME in ID Hltf KO mice. The Wald test was used to generate p-values and log2 fold changes. Genes with the Benjamini-Hochberg adjusted p-value (padj) < 0.05 and absolute log2 fold change > 1 were called as differentially expressed (DE) genes and clustered based on functional annotation by their gene ontology. Enrichment of gene ontology terms was tested using Fisher’s exact test (P-value < 0.05 in the DE gene sets).
RESULTS
HLTF expression is negatively associated with survival in human CRC
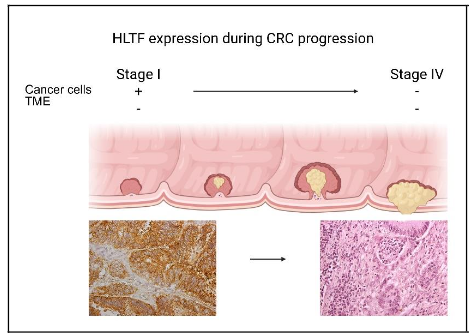
The negative correlation between HLTF expression and the progression of CRC is well documented(5-8,25,26) in humans (Figure 1). Our experimental model differentiated the early stages of CRC from end stage disease in which the complete absence of HLTF from tumor cells and the TME produced the most aggressive metastatic phenotype with the shortest life expectancy(24).
Comparative transcriptomics of TME
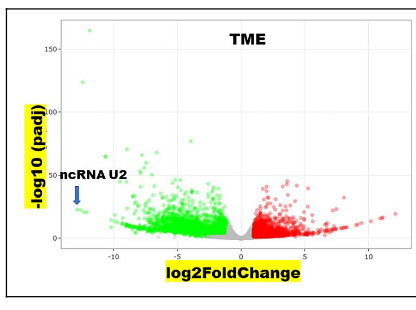
A total of 6,312 genes were DE in the TME of HLTF+/+ vs HLTF-/- ID Hltf KO mice. Non-coding (nc) RNA U2 (Gm25939) was the most transcriptionally downregulated (log2FC -12.86) gene (Figure 2). U2 is one of the ncRNA components of the replication-dependent spliceosomal pathway. Nucleosome assembly (GO:0006334; P value=0.002584) was the pathway in the TME most affected by the presence/absence of HLTF in the primary tumor. Transcription profiles of all major histones were reduced except for the replication-independent histone variant H3.3(27,28). The two genes that encode H3.3 (H3f3a, log2FC +1.52969, padj 1.09E-20; H3f3b, log2FC +0.970875, padj 9.78E-6) were transcriptionally activated.
Comparative transcriptomics primary tumor (CDX)
A total of 5,913 genes were DE in the CDX of HLTF+/+ vs HLTF-/- ID Hltf KO mice. Human box C/D SnoRNA U3 (SNORD3A) is an abundant non-coding RNA with an estimated 100-200 copies per cell. SNORD3A has a non-canonical role in ribosome biogenesis as a chaperone for pre-18S RNA folding(29). It functions as a scaffold during early ribosome assembly, and as a miR-U3 source in HCT116 cells(30). In the CDX, SNORD3A was the most significantly (log2FC -12.758) down regulated gene. A putative HLTF-binding site was found upstream of the SNORD3A promoter (Figure 3).

Protein S-glutathionylation (Pr-SSG)
Pr-SSG of key components of the glycolysis pathway promoted antioxidant defense in the HLTF-deleted CDX/TME compared to the control (24). Human 60 kDa heat shock protein (HSP60) also known as chaperonin 60 (Cpn60) is typically located inside mitochondria(31). Here it preserves proteome integrity via protein folding. HSP60 has three cysteine residues (Cys237, Cys442, and Cys447); however, only Cys447 was authenticated by MALDI-TOF/TOF as site-specific Pr-SSG in the HLTF-deleted CDX/TME model. Further use of MALDI-TOF/TOF followed by species-specific data analysis identified highly conserved cysteines in identical peptide fragment ion spectra from human and mouse.
Nucleosomes, the fundamental units of chromatin, are composed of 145-147-bp of DNA wound around a histone octamer consisting of 2 copies of each of four core histones: H2A, H2B, H3, and H4(32). All histone proteins with the exception of H4 have variant counterparts. Three main histone H3 variants: H3.1 and H3.2 are known as canonical histone H3, and the replication independent variant is H3.3. The centromere-specific variant is CENP-A. Histone H3 variants (H3.1, H3.2 and H3.3) detect cellular redox changes via glutathionylation of Cys110 residues(33) that increases during proliferation and promotes chromatin opening(34).
Importantly, Pr-SSG at H3Cys110 was not detected in either the TME or the primary tumor of the HLTF-deleted model. In addition, consistent with findings for the TME, transcriptional down regulation of histone genes was identified by RNAseq in the primary tumor. The primary gene encoding canonical Histone H3 is HIST1H3A (log2FC -8.0735, padj = 9.33E-15). Region-specific down regulation of Hist1H3A was confirmed by spatial transcriptomics (Figure 4).
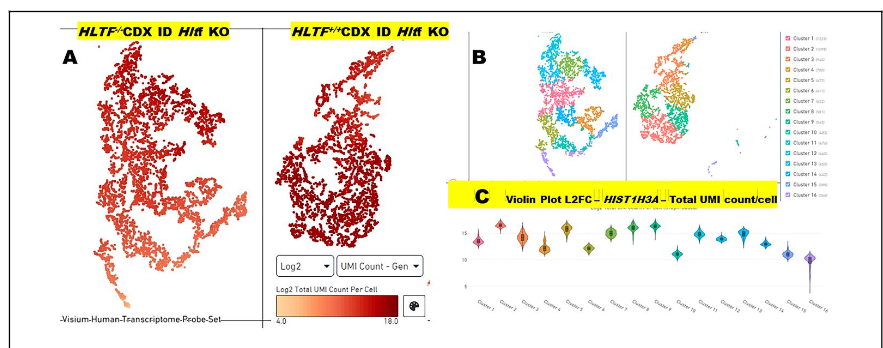
In contrast, the two distinct genes that encode the replication-independent histone variant H3.3 in human and mouse are transcriptionally activated by HLTF-deletion: H3F3A (log2FC +1.640175; padj = 5.73E-6) and H3F3B (log2FC +0.869786, padj = 4.40E-08). As in the TME, specific chaperones used by the H3.3 variant (HIRA, DAXX/ATRX, and DEK) are all transcriptionally available.
DISCUSSION
Orthotopic xenograft transplantation studies with HCT116 cells recapitulate TME-specific interactions including local tumor invasion, angiogenesis and metastasis. HCT116-Red-FLuc cells have wild-type p53 expression and a mutation in codon 13 (G13D) of the KRAS gene component of the RAS/RAF/MED/ERK signaling pathway. Their oncogenic aggressiveness and limited differentiation potential made them an ideal model for the introduction of an HLTF-deletion (24).
To better understand tumor-stroma interactions in vivo, we manipulated HLTF-expression in the primary tumor (CDX) and the TME (mouse) and performed RNAseq with species-specific sequencing. Spatial transcriptomics validated DE in the human transcriptome. Immunohistochemistry and mass spectrometry provided insights to glutathione-based antioxidant defense in the model. Collective re-evaluation of the data allows us to report decreased availability of ncRNA U2 (TME) and SNORD3A (CDX) correlates with a unified shift to replication-independent availability of H3.3 to chromatin in the HLTF-deleted CDX/TME compared to the control.
Decreased availability of ncRNA U2 would reduce its role in alternative splicing in CRC(35), and reduce its potential as a biomarker/therapeutic target in late stage metastatic disease(36-37). Mechanistically, SNORD3A promotes ferroptosis in acute kidney injury(38), chemosensitization (5-FU) in breast cancer(39), and maintenance of leukemia stem cells(40). In CRC, negligible SNORD3A expression could mitigate cell proliferation in favor of metastasis.
Histone H3C110 variants — the only nucleosomal proteins with cysteine residues that can be modified by glutathione — were not among the glutathionylated proteins we identified. Perhaps this is because cell proliferation and sensing cellular redox changes are connected through glutathionylation of Cys110(33,34), neither of which is consistent with the metastatic properties of our CDX model.
The loss of canonical histones at the mRNA level in both the TME and the CDX commensurate with increased transcriptional availability of H3.3 in the HLTF-deleted CDX/TME compared to the control supports Gomes et al. (41) who recently showed the incorporation of H3.3 into chromatin drives metastasis. H3.3 differs from canonical H3.1 by 4 amino acid (aa) residues A87, I89, G90 and S96 in the histone fold domain and aa residue S31 in the N-terminal tail. H3.3 differs from H3.2 by only 4 aa residues S31, A87, I89, G90. A87, I89 and G90 (AIG motif) are important for recognition by H3.3-specific chaperones. S31 is phosphorylated in the pericentromeric region during mitosis. The N-terminal tail (aa 1-50) of H3 and H3.3 have the same Lys23 residue that is not commonly mutated. HLTF targets H3 at Lys23 (H3K23) for ubiquitination in CRC pathogenesis (19). HLTF histone ubiquitination activity is stimulated by the repressive H3K9me3 mark. Loss of HLTF reduces H3K23ub and H3K9me, which increases chromatin accessibility at promoters and enhancers of cancer/metastasis promoting genes.
CONCLUSION
The elimination of helicase-like transcription factor (HLTF), also known as SMARCA3, supported redox homeostasis in a CDX/TME model compared to the control(24). Additionally, decreased transcriptional availability of SNORD3A (CDX) and ncRNA U2 (TME), and increased transcriptional availability of the replication-independent histone H3.3 variant (CDX and TME) advances epigenetic reprogramming for tumor progression and metastatic behavior.
Conflict of Interest Statement: None.
Funding Statement: Harry Weitlauf Endowment for Cancer Research.
REFERENCES
- Siegel RL, Kratzer TB, Giaquinto AN, Sung H, Jemal A. Cancer statistics, 2025. CA Cancer J Clin. 2025;75(1):10–45. doi: 10.3322/caac.21871.
- Maida M, Dahiya DS, Shah YR, et al. Screening and surveillance of colorectal cancer: A review of the literature. Cancers (Basel), 2024;16(15):2746. doi: 10.3390/cancers16152746.
- Waheed Y, Mojumdar A, Shafiq M, de Marco A, De March M. The fork remodeler helicase-like transcription factor in cancer development: all at once. Biochim Biophys Acta Mol Basis Dis. 2024;1870(7):167280. doi: 10.1016/j.bbadis.2024.167280.
- Dhont L, Mascaux C, Belayew A. The helicase-like transcription factor (HLTF) in cancer: loss of function or oncomorphic conversion of a tumor suppressor. Cell Mol Life Sci. 2016;73(1):129–145. doi: 10.1007/s00018-015-2060-6.
- Moinova HR, Chen WD, Shen L, et al. HLTF gene silencing in human colon cancer. Proc Natl Acad Sci USA. 2002;99(7):4562–4567. https://doi.org/10.1073/pnas.062459899. PMID:11904375.
- Leung WK, To KF, Man EP, et al. Detection of hypermethylated DNA or cyclooxygenase-2 messenger RNA in fecal samples of patients with colorectal cancer or polyps. Am J Gastroenterol. 2007;102(5):1070–1076.
- Philipp AB, Stieber P, Nagel D, et al. Prognostic role of methylated free circulating DNA in colorectal cancer. Int J Cancer 2012;131:2308-2319. doi: 10.1002/ijc.27505. PMID: 22362391.
- Jung G, Hernandez-Illan E, Moreira L, Balaguer F, Goel A. Epigenetics of colorectal cancer: biomarker and therapeutic potential. Nat Rev Gastroenterol Hepato. 2020;17(2):111-130. doi: 10.1038/s41575-019-0230-y. PMID:31900466.
- Dhont L, Belayew A. HLTF (helicase-like transcription factor). Atlas Genet Cytogenet Oncol Haematol. 2016-11-01. Online version: http://atlasgeneticsoncology.org/gene/42332/hltf.
- Sheridan PL, Schorpp M, Voz ML, Jones KA. Cloning of an SNF2/SWI2-related protein that binds specifically to the SPH motifs of the SV40 enhancer and to the HIV-1 promoter. J Biol Chem. 1995;270(9):4575–4587. 10.1074/jbc.270.9.4575;
- Hayward-Lester A, Hewetson A, Beale EG, Oefner PJ, Doris PA, Chilton BS. Cloning, characterization and steroid-dependent posttranslational processing of RUSH-1α and β, two uteroglobin promoter binding proteins. Mol Endocrinol 1996; 10:1335–1349. 10.1210/mend.10.11.8923460;
- Ding H, Descheemaeker K, Marynen P, et al. Characterization of helicase-like transcription factor involved in the suppression of the human plasminogen activator inhibitor-1 gene. DNA Cell Biol. 1996;15(6):429–442. 10.1089/dna.1996.15.429;
- Hewetson A, Hendrix EC, Mansharamani M, Lee VH, Chilton BS. Identification of the RUSH consensus-binding site by cyclic amplification and selection of targets: demonstration that RUSH mediates the ability of prolactin to augment progesterone-dependent gene expression. Mol Endocrinol. 2002;16(9):2101-12. doi: 10.1210/me.2002-0064.
- Unk I, Hajdu I, Fatyol K, et al. Human HLTF functions as a ubiquitin ligase for proliferating cell nuclear antigen polyubiquitination. Proc Natl Acad Sci USA 2008;105(10):3768-3773. doi: 10.1073/pnas.0800563105.
- Kang S, Yoo J, Myung K. PCNA cycling dynamics during DNA replication and repair in mammals. Trends Genet. 2024;40(6):526-539. doi: 10.1016/j.tig.2024.02.006.
- Kile AC, Chavez DA, Bacal J, et al. HLTF’s Ancient HIRAN domain binds 3′ DNA ends to drive replication fork reversal. Mol Cell 2015;58(6):1090-1100.
- Chavez DA, Greer BH, Eichman BF. The HIRAN domain of helicase-like transcription factor positions the DNA translocase motor to drive efficient DNA fork regression. J Biol Chem. 2018; 293(22):8484-8494. doi: 10.1074/jbc.RA118.002905.
- van Toorn M, Turkyilmaz Y, Han S, et al. Active DNA damage eviction by HLTF stimulates nucleotide excision repair. Mol Cell. 2022;82(7):1343–1358. e8. doi: 10.1016/j.molcel.2022.02.020.
- Akano I, Hebert JM, Tiedmann RL, et al. The SWI-SNF related protein SMARCA3 is a histone H3K23 ubiquitin ligase that regulates H3K9me3 in cancer. Mol Cell. 2025;85(15):2885-2899. e8. doi: 10.1016/j.molcel.2025.06.020.
- Reginato G, Dello Stritto MR, Wang Y, et al. HLTF disrupts Cas9-DNA post-cleavage complexes to allow DNA break processing. Nat Commun. 2024;15(1):5789. doi: 10.1038/s41467-024-50080-y.
- Spiegel J, Adhikari S, Balasubramanian S. The structure and function of DNA G-quadruplexes. Trends Chem. 2020;2(2):123-136. doi: 10.1016/j.trechm.2019.07.002.
- Bai G, Endres T, Kuhbacher U, et al. HLTF resolves G4s and promotes G4-induced replication fork slowing to maintain genome stability. Mol Cell. 2024;86(16): 3044-3060.e11. doi: 10.1016/j.molcel.2024.07.018.
- Dusek CO, Dash RC, McPherson KS, et al. DNA sequence specificity reveals a role of the HLTF HIRAN domain in the recognition of trinucleotide repeats. Biochemistry. 2022;24:10.1021/acs.biochem.2c00027. doi: 10.1021/acs.biochem.2c00027.
- Martinez-Marin D, Helmer RA, Kaur G, et al. Helicase-like transcription factor (HLTF)-deleted CDX/TME model of colorectal cancer increased transcription of oxidative phosphorylation genes and diverted glycolysis to boost S-glutathionylation in lymphatic intravascular metastatic niches. PLoS One. 2023;18(9):e0291023. doi: 10.1371/journal.pone.0291023.
- Wallner M, Herbst A, Behrens A, et al. Methylation of serum DNA is an independent prognostic marker in colorectal cancer. Clin Cancer Res. 2006;12(24):7347–7352. doi: 10.1158/1078-0432.CCR-06-1264.
- Herbst A, Wallner M, Rahmig K, et al. Methylation of helicase-like transcription factor in serum of patients with colorectal cancer is an independent predictor of disease recurrence. Eur J Gastroenterol Hepatol. 2009; 21(5):565–569. doi: 10.1097/MEG.0b013e328318ecf2.
- Ahmad, K. & Henikoff, S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell 9, 1191–1200 (2002).
- Goldberg AD, Banaszynski LA, Noh K-M et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell. 2010;140(5): 678–691. doi: 10.1016/j.cell.2010.01.003
- Dutca LM, Gallagher JEG, Saserga SJ. The initial U3 snoRNA:pre-rRNA base pairing interaction required for pre-18S rRNA folding revealed by in vivo chemical probing. Nucleic Acids Res. 2011;39(12):5164-80. doi: 10.1093/nar/gkr044.
- Lemus-Diaz N, Ferreira RR, Bohnsack KE, Gruber J, Bohnsack MT. The human box C/D snoRNA U3 regulates expression of sortin nexin 27. Nucleic Acids Res. 2020;48(14):8074-8089. doi: 10.1093/nar/gkaa549.
- Singh MK, Shin Y, Han S, et al. Molecular chaperonin HSP60: Current understanding and future prospects. Int J Mol Sci. 2024;25(10): 5483; https://doi.org/10.3390/ijms25105483.
- Feng Y, Endo M, Sugiyama H. Nucleosomes and epigenetics from a chemical perspective. Chembiochem. 2021;22(4):595-612. doi.org/10.1002/cbic.202000332.
- Garcia-Gimenez JL, Olaso G, Hake SB, et al. Histone H3 glutathionylation in proliferating mammalian cells destabilizes nucleosomal structure. Antioxid. Redox Signal. 2013;19(12):1305–1320. doi: 10.1089/ars.2012.5021.
- Garcia-Gimenez JL, Pallardo FV. Maintenance of glutathione levels and its importance in epigenetic regulation. Front Pharmacol. 2014;5:88. doi: 10.3389/fphar.2014.00088.
- Guiro J, Fagbemi M, Tellier M, et al. CAPTURE of the human U2 snRNA genes expands the repertoire of associated factors. Biomolecules 2022;12(5):704. doi: 10.3390/biom12050704.
- Song Z, Bae B, Schnabl S, et al., Mapping snoRNA-target RNA interactions in an RNA-binding protein-dependent manner with chimeric eCLIP. Genome Biol. 2025; 26:39. doi: 10.1186/s13059-025-03508-7
- Maqbool M, Khan A, Shahzad. Predictive biomarkers for colorectal cancer: a state-of-the-art systematic review. Biomarkers 2023;28(6):562-598. doi: 10.1080/1354750X.2023.2247185.
- Zhu H, Wang J, Miao J, et al. SNORD3A regulates STING transcription to promote ferroptosis in acute kidney injury. Adv Sci (Weinh). 2024 Sep;11(33):e2400305. doi: 10.1002/advs.202400305.
- Luo, L, Zhang, J, Tang, H, et al. LncRNA SNORD3A specifically sensitizes breast cancer cells to 5-FU by sponging miR-185-5p to enhance UMPS expression. Cell death & disease, 2020;11,329. doi:10.1038/s41419-020-2557-2.
- Yun, H, Zoller, J, Zhou, F, et al. The landscape of RNA-chromatin interaction reveals small non-coding RNAs as essential mediators of leukemia maintenance. Leukemia. 2024; 38, 1688-1698. doi:10.1038/s41375-024-02322-7.
- Gomes AP, Llter D, Low V, et al. Dynamic incorporation of Histone H3 variants into chromatin is essential for acquisition of aggressive traits and metastatic colonization. Cancer Cell. 2019; 36(4): 402–417.e13. doi:10.1016/j.ccell.2019.08.006.




