Rapid Triage of Radiation Exposure: A 4-Gene PCR Test
Rapid Triage of Radiation Exposure Using a 4-Gene Real Time PCR Test
Robert H. Terbrueggen1, Samuel Dixon1, Timothy Guyon1, Aviva R. Jacobs1
- DxTerity Diagnostics Incorporated, Torrance, CA 90501 USA
OPEN ACCESS
PUBLISHED: 30 November 2024
CITATION: Terbrueggen, RH., Dixon, S., et al., 2024. Rapid Triage of Radiation Exposure Using a 4-Gene Real Time PCR Test. Medical Research Archives, [online] 12(11). https://doi.org/10.18103/mra.v12i11.6114
COPYRIGHT: © 2024 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI: https://doi.org/10.18103/mra.v12i11.6114
ISSN: 2375-1924
Abstract
The ability to distinguish the concerned public from the significantly exposed is of critical importance for mounting an effective response to a mass scale nuclear incident. Here we describe the development and performance of a radiation biodosimetry blood test with a sensitivity of 99.0% and specificity of 96.5% for the qualitative determination of irradiated vs non-irradiated. Similar performance is observed in human and non-human primate (rhesus macaque) blood samples for in vivo total body irradiation as well as ex vivo blood irradiation. The test is based on the relative expression of 3 radiation responsive mRNA biomarkers relative to a normalizer mRNA using a pre-established cut-off. Blood samples must be collected into an RNA stabilizing blood collection device and sample testing is compatible with the real time PCR infrastructure used worldwide for COVID-19 pandemic response.
Keywords
- Radiation Exposure
- Biodosimetry
- Real Time PCR
- mRNA Biomarkers
Introduction
Following a mass scale nuclear event caused by the detonation of a nuclear device or a nuclear reactor meltdown, hundreds to hundreds of thousands of individuals will require medical management. Emergency responders will start with treating individuals with obvious physical injury, however many with radiation injury will not display physical symptoms until several days to a week post-incident. Diagnostic tests that allow for the identification of those with radiation injury while calming the worried are key to providing an effective response.
Many radiation response scenarios envision using a qualitative triage test that initially separates the significantly exposed from the worried well, and then using a quantitative biodosimetry test to help guide treatment decisions. The REDI-Dx Biodosimetry Quantitative Test System recently became the first biodosimetry test to receive CE-IVD regulatory authorization. REDI-Dx Quantitative is an 18-plex gene expression assay that uses capillary electrophoresis for readout and is intended for use in a hospital setting. Other gene expression biodosimetry tests based on real time PCR or microarrays have been described.
Here we described the development of a high throughput, real time PCR biodosimetry test suitable for initial triage of a mass scale nuclear event. The test is designed for qualitative (irradiated versus non-irradiated) assessment using real time PCR.
Materials and Methods
Sample Collection and Stabilization
All human samples were collected from subjects providing appropriately documented, HIPAA compliant informed consent under clinical research protocols reviewed and approved by an accredited Institutional Review Board (IRB) and implemented in accordance with the ICH Harmonized Guidelines for Good Clinical Practice (GCP), applicable regulations (including CFR Title 21). Human TBI samples were obtained from Duke University and City of Hope Cancer Center. Ex Vivo Irradiated Blood Samples were obtained from the University of Arizona. Subject enrollment for human demographic samples occurred at various clinical sites, which were managed by Access Biologicals (Vista, California) and Cureline (Brisbane, California). Samples with potentially confounding medical conditions were obtained from Cureline (Brisbane, California and Serologix (New Hope, Pennsylvania). Burn and Trauma samples were obtained from Maricopa Integrated Health System (Phoenix, Arizona). For Burn victims, samples were assigned to Total Body Surface Area (TBSA) categories of <10%, 10-20% and >21%. Trauma patients were assigned to Injury Severity Scale (ISS) categories of 10-14, 15-24 and >25. Influenza samples were obtained from Duke University. Inclusion criteria for all conditions: ≥18 years of age; Negative testing for Hepatitis B/C, HIV-1 and HIV-2 in clinical history, with the exception of samples being collected due to HIV status. Donors were excluded if they had received chemotherapy or a blood transfusion within the last 6 months, had X-rays, CT scans, mammography, or radioactive dyes for scans or heart evaluations within the last 3 months.
Human samples were collected with DxCollect® Blood Collection tubes (BCT- DxTerity Diagnostics) using standard phlebotomy techniques. These tubes contain RNA stabilization buffer, which lyses the cells and stabilizes the RNA.
Samples from single dose in-vivo irradiated Non-Human Primates (Rhesus Macaque) were provided by Duke University, which were obtained from studies performed in compliance with protocols approved by an accredited IACUC and under the Biomedical Advanced Research and Development Authority (BARDA), Office of the Assistant Secretary for Preparedness and Response, Office of the Secretary, Department of Health Human Services Contract No: HHSO100201000001C and HHSO100201600034C. For additional details on the irradiation protocol refer to Iversen, et al 2018.
Samples from Total Body Irradiated (TBI) Patients were obtained from City of Hope (Duarte, California), UCLA (Los Angeles, CA) and Duke University (Durham, NC) and collected per IRB approved protocols. Inclusion criteria were for planned fractionated TBI as part of clinical care, for malignant and non-malignant diseases treated with hematopoietic stem cell transplant, Subjects were excluded if they had received chemotherapy within 21 days, had concurrent chemotherapy with fludarabine, cytokine inhibitor or cytokine‐inducer therapy within 30 days either prior to or during the irradiation regimen. Patients receiving granulocyte colony‐stimulating factor within 30 days prior to TBI, were also excluded.
Human Ex Vivo Irradiation
Human whole blood was collected from consented healthy donor into 5ml sodium citrate vacutainers by venipuncture. After mixing by inversion, the blood was irradiated using a Varian 21EX linear accelerator within 15 minutes of collection. Samples were irradiated at 212 MU/min.
After irradiation, 2.5 mL of blood was mixed by inversion with 2.5 ml of pre-warmed to 37°C RPI-1640 media in a 15 ml falcon plastic tube. Samples were incubated at 37°C in a CO2 incubator for 24 hours. The culture samples were then mixed with an equal volume of DxCollect® RNA stabilization buffer.
Human Total Body Irradiation
Peripheral blood was collected from adult patients undergoing TBI-based myeloablative allogenic or autologous stem-cell transplantation. Samples were collected prior to irradiation at the beginning of each day from patients receiving either 150 cGy (Duke University) or 120 cGy (City of Hope). TBI patient blood samples with cumulative doses of 300 cGy, 600 cGy, 900 cGy and 1050 cGy (Duke University) or 360 cGy, 720 cGy and 1080 cGy (City of Hope).
Total RNA Isolation
Total RNA was isolated from DxCollect® stabilized blood samples by magnetic bead capture and elution using a proprietary automated method similar to automated processing using Beckman Coulter Agencourt RNAdvance Blood kits. Briefly, 350 microliters of stabilized blood were mixed with 265 microliters of lysis buffer and 20 microliters of protein kinase reagent in a deep well plate. After vortexing, the plate is incubated at 55°C for 15 minutes. Then 5ul of Sera-Mag Speedbeads (GE Life Sciences) in 350 microliters of ethanol were added. The RNA isolation and elution was then performed using an Auto-Pure 96 instrument (Hangzhou Allsheng) with final elution in 50 microliters of nuclease free water.
Reverse Transcriptase qPCR
In a 96-well plate, 6.25 microliters of TaqPath 1-Step Multiplex Master Mix (No Rx), 1.25 microliters of Primer mix, 7.5 microliters of nuclease-free water, and 10.0 microliters of isolated total RNA from bead isolation procedure were mixed together. RT-qPCR was performed using ABI ViiA7 real time PCR machine with reporter dyes of FAM, VIC, Cy5 and TEXAS RED. After 2 minutes at 25°C for UNG incubation, 10 min at 53°C for reverse transcriptase, 2 min at 95°C for Taq activation and 45 cycles of cycling between 95°C for 3 sec and 60°C for 30 seconds. Ct Values were determined using QuantStudio V1.3 software.
Data Analysis
The Ct values observed for BAX, CDKN1A and DDB2 were normalized relative to MRPS5. A weighted averaged metagene was calculated and fold change relative to the averaged metagene for unirradiated was determined. The cutoff for exposed versus unexposed is based on 3 standard deviations of the metagene for 149 healthy human samples, corresponding to a delta Ct of -0.76, corresponding to a 1.7-fold change.
Results
Testing of Ex Vivo Irradiated Human Blood Samples
Total RNA isolated from human blood samples that had been ex vivo irradiated at 0, 0.05, 0.1, 0.25, 0.5, 1.0, 1.5, 2.0, 3.0, and 4.0 Gy. The samples were tested using the 4-gene real time PCR assay. A total of 138 samples were tested. The observed fold changes of the BAX, CDKN1A, DDB2 metagene as a function of dose are shown in table 1.
| Dose (Gy) | Samples | Metagene Fold Change |
|---|---|---|
| 0.0 | 18 | 1 |
| 0.05 | 4 | 1.2 |
| 0.1 | 4 | 1.4 |
| 0.25 | 4 | 1.9 |
| 0.5 | 18 | 3.4 |
| 1.0 | 18 | 3.6 |
| 1.5 | 18 | 4.1 |
| 2.0 | 18 | 4.7 |
| 3.0 | 18 | 5.5 |
| 4.0 | 18 | 5.7 |
Table 1. Summary of fold change observed for the metagene as a function of dose for ex vivo irradiated human blood samples.
Testing of In Vivo, Total Body Irradiated (TBI) Non-Human Primates (NHPs)
NHP samples irradiated from 0.5 to 10.8 Gy were tested. The blood samples were collected from 24 hours to 168 hours (7 Days) post-exposure following irradiation from 0.5 Gy to 10.8 Gy. The metagene fold changes as a function of dose and time are shown in Table 2. The results are based on a total of 461 NHP samples.
| Time (Hours) | 0.5 Gy | 1 Gy | 2 Gy | 4 Gy | 6 Gy | 8 Gy | 10.8 Gy |
|---|---|---|---|---|---|---|---|
| 24 | 2.6 | 5.3 | 5.2 | 5.7 | 7.9 | 5.9 | 5.1 |
| 72 | 2.4 | 4.1 | 8.3 | 9.4 | 11.3 | 8.5 | 12.4 |
| 120 | 2.0 | 3.0 | 6.5 | 7.9 | 11.8 | 13.0 | 16.2 |
| 168 | 1.9 | 1.9 | 3.4 | 6.7 | 8.8 | 8.1 | 9.5 |
Table 2. Calculated fold change for the metagene as a function of time post irradiation (hours) and absorbed dose for NHP single dose TBI samples.
Testing of Human TBI Fractionated Samples
Blood samples from Human Cancer Patients undergoing Total Body Irradiation were tested. The total fractionated dose received varied from 1.5 Gy to 13.2 Gy and a cumulative time post irradiation varied from 5.5 hour to 96 hours. Observed fold change as a function of time post irradiation and dose is shown in Table 3.
| Dose (Gy) | Time Post Irradiation (Hr) | Samples | Fold Change |
|---|---|---|---|
| 0 | 0 | 23 | 1.0 |
| 1.5 | 5.5 | 41 | 5.0 |
| 2.4 | 24 | 3 | 3.8 |
| 3 | 24 | 7 | 5.4 |
| 3.6 | 24 | 12 | 4.0 |
| 6 | 48 | 11 | 5.5 |
| 7.2 | 48 | 12 | 4.8 |
| 9 | 72 | 5 | 7.1 |
| 9.6 | 72 | 3 | 6.9 |
| 10.5 | 72 | 6 | 6.7 |
| 10.8 | 72 | 8 | 6.5 |
| 13.2 | 96 | 5 | 8.9 |
Table 3. Calculated fold change for the metagene as a function of time post irradiation (hours) and absorbed dose for human fractionated dose TBI samples.
Potentially Confounding Human Samples
Isolated total RNA from 22 different potentially confounding human conditions were tested including burn, trauma, flu, bacterial infection, and post-treatment with granulocyte colony stimulating factor (G-CSF). 355 of the 360 (98.6%) of the samples tested were correctly called unexposed.
| Confounding Condition | Samples | Accuracy (percent called unirradiated) |
|---|---|---|
| Burn | 36 | 97.2% |
| Trauma | 19 | 100% |
| Influenza | 50 | 96% |
| Granulocyte Colony-Stimulating Factor (G-CSF) | 13 | 100% |
| Moderate to Severe Infection | 36 | 94.4% |
| Anti-Emetic | 25 | 100% |
| Anti-Diarrheal | 10 | 100% |
| Aspirin | 15 | 100% |
| Ibuprofen | 10 | 100% |
| Methamphetamines | 14 | 100% |
| Steroidal anti-inflammatories | 15 | 100% |
| Inflammatory Bowel Disease (IBD) | 13 | 100% |
| Arthritis | 11 | 100% |
| Cardiovascular Disease | 14 | 100% |
| Immunocompromised | 15 | 100% |
| Lipemia | 12 | 100% |
| Obesity/High BMI | 11 | 100% |
| Pregnancy | 11 | 100% |
| Lactation | 10 | 100% |
| Smoker | 10 | 100% |
| Type II Diabetes | 10 | 100% |
| Total | 360 | 98.6% |
Table 4. Accuracy of prediction of blood samples from humans suffering from conditions likely to be present post-nuclear incident (burn, trauma, influenza and infection), likely treatments (G-CSF, anti-emetic, anti-diarrheal, aspirin, ibuprofen, steroidal anti-inflammatories) and chronic human conditions.
Calculation of Assay Performance: Sensitivity and Specificity
Assay Performance was assessed based on a delta Ct cutoff of -0.77Ct for the normalized metagene. A total of 1486 in vivo irradiated, ex vivo irradiated and non-irradiated human samples were used to assess performance.
| Species | Irradiation Type | N Rows |
|---|---|---|
| Human | Ex Vivo | 150 |
| Human | In Vivo Fractionated Dose | 142 |
| Human | Healthy-N/A | 304 |
| Human | Confounders-N/A | 360 |
| NHP | Ex Vivo | 19 |
| NHP | In Vivo Fractionated Dose | 50 |
| NHP | In Vivo Single Dose | 461 |
| Total | 1486 |
Table 5. List of samples used to assess performance of radiation biodosimetry for qualitative assessment of irradiated vs non-irradiated.
| Irradiation Status | Predicted | Actual Irradiated | Non-Irradiated |
|---|---|---|---|
| Irradiated | 613 | 6 | |
| Non-Irradiated | 30 | 837 |
Table 6. Actual vs Predicted Radiation Status for 1486 blood samples based on a metagene cutoff of -0.77 (1.7-fold increase) in gene expression.
Discussion
Genomic and proteomic technologies and diagnostic tests offer great promise for improving preparedness and response to a nuclear incident, however the lack of human samples for new test development is a major barrier to promise fulfillment. While Non-Human Primates (NHPs) are the preferred model system for test development, the expense and humane considerations limit their availability. Many test developers are forced to rely upon less reliable surrogates such as mouse models and ex vivo irradiated human blood.
Here we used upon a combination of total body irradiated (TBI) single dose NHPs, TBI fractionated dose NHPs, ex vivo irradiated human blood, human fractionated dose TBI cancer patients, and ex vivo irradiated human blood to demonstrate test performance. The NHP samples enable evaluation for up to 7 days post-exposure, the ex vivo irradiated samples provide evaluation at low dose without unnecessary use of NHPs, and the Human TBI samples provide verification of the NHP results.
The mRNA biomarkers used in the biodosimetry test were selected based on being highly responsive to radiation and showing comparable response in all model systems. The highly radiation responsive gene FXDR was excluded because of very different response in humans and NHPs.
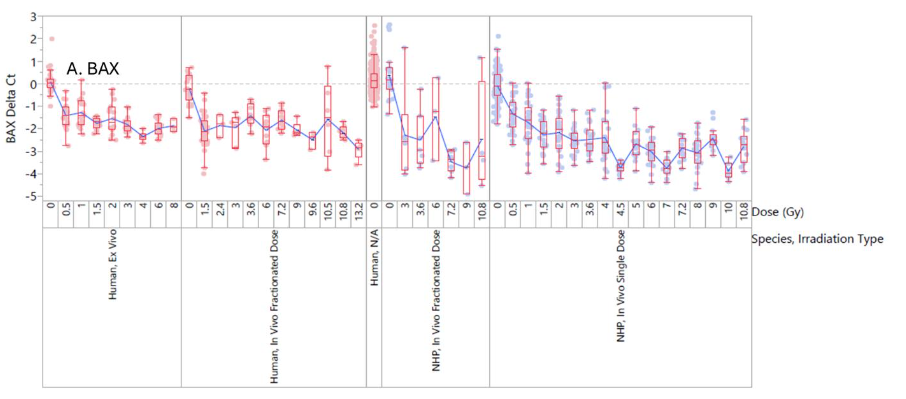
There is clear separation between irradiated and non-irradiated samples for NHP and human samples in all model systems. The limit of detection for the test is approximately 0.5 Gy. This is a very reasonable limit of detection since there are no reported cases of acute radiation syndrome (ARS) from individuals irradiated with less than 0.5 Gy.
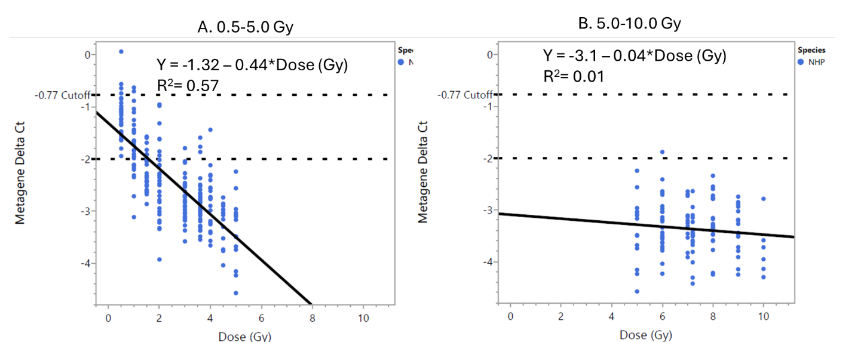
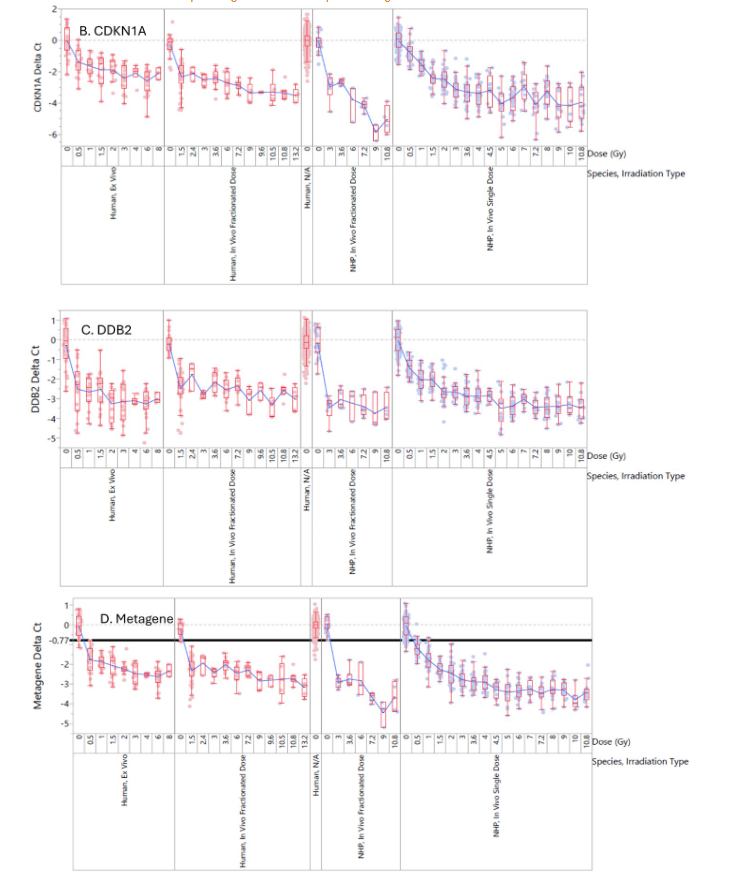
The fold change in the metagene peaks around 72 hours post-exposure and it is still upregulated at 168 hours post-exposure (7 Days). The 0.5 and 1.0 Gy dose samples approach the 1.7-fold cut-off at day 7, while the higher dose samples remain significantly elevated.
| Time (Hours) | 0.5 Gy | 1 Gy | 2 Gy | 4 Gy | 6 Gy | 8 Gy | 10 Gy |
|---|---|---|---|---|---|---|---|
| 24 | 2.6 | 5.2 | 5.3 | 5.5 | 9.5 | 5.8 | 5.0 |
| 72 | 2.2 | 3.4 | 6.7 | 8.1 | 13.5 | 12.0 | 10.7 |
| 120 | 2.0 | 3.0 | 6.4 | 7.7 | 11.5 | 12.7 | 15.8 |
| 168 | 1.8 | 1.8 | 3.3 | 6.5 | 8.5 | 7.9 | 9.2 |
Table 7. Fold change calculations as a function of dose and time post irradiation for in vivo single dose NHP samples. Fold change equals 2-Ct.
The impact of potentially confounding chronic conditions and commonly taken therapeutics as well as trauma, burn, flu, infection and G-CSF treatment were investigated.
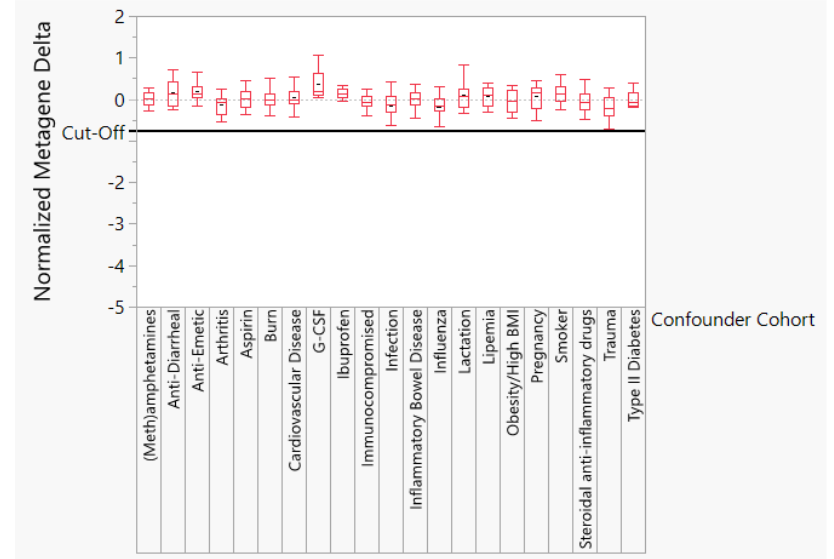
The test sensitivity is 99.0% and specificity is 96.5% based on 1486 samples consisting of 530 NHP TBI, 360 potentially confounding condition, 142 human TBI, 150 human ex vivo, and 304 healthy human samples.
| Threshold | Sensitivity | Specificity | Pos Predictive Value | Negative Predictive Value | Accuracy |
|---|---|---|---|---|---|
| 99.0% | 96.5% | 95.3% | 96.7% | 97.1% |
Table 8. Performance summary for 4-mRNA RT-PCR Biodosimetry Test.
Conclusions
A 4-mRNA biodosimetry test shows high sensitivity and specificity for identifying individuals that have absorbed more than 0.5 Gy of absorbed radiation. Samples must be collected into a blood RNA stabilizing device within 7 days post-exposure. The test is compatible with the large installed base of real time PCR testing equipment that was used to respond to the COVID pandemic, and this test has the potential of being a valuable tool in responding to a mass scale nuclear incident. This test can be particularly useful for calming the public following a nuclear event and avoiding misallocation of scarce resources like hospital beds and Granulocyte Colony Stimulating Factor (G-CSF).
References
- Buddemeier BR, Valentine JE, Millage KK and Brandt LD. Nation Capital Region – Key Response Planning Factors for the Aftermath of Nuclear Terrorism, 2011; Nov. LLNL-TR-512111.
- Coleman CN et al. Triage and treatment tools for use in a scarce resources-crisis standards of care setting after a nuclear detonation, Disaster Med Public Health Prep., 2011; Mar:5 Suppl 1: S111-21.
- Ricks RC, Berger ME, and O’Hara FM. The Medical Basis for Radiation Accident Preparedness – The Clinical Care of Victims, The Parthenon Publishing Group 2002.
- Satyamitra M, Turcu F, Pantoja-Galicia N, Wathen L. Challenges and Strategies in the Development of Radiation Biodosimetry Tests for Patient Management, Radiat Res. 2021; 196(5):455-467. Doi:10.1667/RADE-21-00072.1.
- Wathen LK, EDER PS, Horwith G, Wallace RL, Using biodosimetry to enhance the public health response to a nuclear incident, International Journal of Radiation Biology, 2021; DOI:10.1080/09553002.2020.1820605.
- Jacobs AR et al. Role of a high throughput biodosimetry test in treatment prioritization after a nuclear incident, International Journal of Radiation Biology, 2018; DOI: 10.1080/09553002.2018.1532615.
- Badie C et al, NATO Biodosimetry Study – Laboratory Intercomparison of Gene Expression Assays, Radiation Research, 2013; 180, 138-148.
- Rothkamm K et al, 2013 “NATO Biodosimetry Study: Comparison of Established and Emerging Biodosimetry Assays”, Radiation Research, 2013; 180, 111-119.
- Bhagi S, Alok A, Chaudhury NK. Radiation-Induced Gene Expression Signatures for Triage Biodosimetry, Defence Life Science Journal, 2021; 6(1):84-92, DOI:10.14429/dlsj.6.15540.
- Schreyer S, Karkanitsa L, Albanese J, Ostapenko V, Shevchuk V, et al., Analysis of radiation-associated changes in gene expression using microarray technology. Brit J Radiol, 2002; Supple 26: 129–139.
- Meadows S, Dressman H, Muramoto G, Himburg H, Salter A, et al. Gene expression signatures of radiation response are specific, durable and accurate in mice and humans, PLoS ONE 2008; 3: e1912.
- Amundson SA, Do KT, Shahab S, Bittner M, Meltzer P, Trent J, Fornace AJJ. Identification of potential mRNA biomarkers in peripheral blood lymphocytes for human exposure to ionizing radiation. Radiation Res. 2000; 154:342–346.
- Ostheim et al. Gene expression for biodosimetry and effect prediction purposes: promises, pitfalls and future directions – key session ConRad, International Journal of Radiation Biology, 2021; 98(5):843-854. DOI: 10.1080/09553002.2021.1987571.
- Iversen ES, McCarthy J, Lipton G, Phillips G, Dressman H, Ross J, Owzar K, and Chao N. [2018]. Bridging the Gaps: Using an NHP Model to Predict Single Dose Radiation Absorption in Humans, International Journal of Radiation Biology, 2018; DOI:10.1080/09553002.2018.1532614.
- Farese, AM, Cohen, MV, Katz, BP, Smith, CP, Gibbs, AM, Cohen, DM, MacVittie, TJ. Filgrastim Improves Survival in Lethally Irradiated Nonhuman Primates. Rad Research, 2013; 179(1):89-100.
- Ossetrova NI et al, Non-human Primate Total-body irradiation Model with Limited and Full Medical Supportive Care Including Filgrastim for Biodosimetry and Injury Assessment, Radiation Protection Dosimetry, 2016; 172(1-3):174-101 DOI:10.1093/rpd/ncw176.
- Ghandhi SA, Smilenov L, Shuryak I, Pujol-Canadell M, Amundson SA, Discordant gene responses to radiation in humans and mice and the role of hematopoietically humanized mice in the search for radiation biomarkers, Sci Rep 2019, 9:19434 DOI:10.1038/s41598-019-55982-2.
- Park JG, Paul S, Brionea N et al, Developing Human Radiation Biodosimetry Models: Testing Cross-Species Conversion approaches Using an Ev Vivo Model System, Radiation Res. 2017; 187(6): 708-721 DOI:10.1667/RR14655.1.
- Lucas J, Dressman HK, Suchindran S et al, A Translatable Predictor of Human Radiation Exposure, PLoS One 2014; 9(9): e107897. DOI:10.1371/journal.pone.0107897.
- Abend M, Blakely WF, Ostheim P, Schuele S and Port M. Early molecular markers for retrospective biodosimetry and prediction of acute health effects, J. Radiol. Prot. 2022; Jan 25;42(1).
- O’Brien G et al, FDXR is a biomarker of radiation exposure in vivo, Sci Rep. 2018; 8, 684 https://doi.org/10.1038/s41598-017-19043-w.

