ReNeg-AID Peptide’s Role in Triple-Negative Breast Cancer
Effect of ReNeg-AID peptide overexpression in MDA-MB-231 triple-negative breast cancer cells: the noncanonical Notch pathway
Germán Saucedo-Corona1, Rosa E. Nuñez-Añita1, Ulises Maciel-Ponce1, Humberto Contreras-Corona1, Alejandro Bravo-Patrón1
- Centro Multidisciplinario de Estudios en Biotecnología de la Facultad de Medicina Veterinaria y Zootecnia, Universidad Michoacana de San Nicolás de Hidalgo. Posta Veterinaria, Km. 9.5 Carretera Morelia-Zinapécuaro, Col. La Palma, C. P. 58893, Tarímbaro, Michoacán, México.
Address to correspondence: Alejandro Bravo-Patiño, Phone: (+52) 443 186 0263, E-mail: [email protected]
OPEN ACCESS
PUBLISHED: 31 October 2024
CITATION: Saucedo-Correa, G., Nuñez-Anita, RE., et al., 2024. Effect of ReNeg-AID peptide overexpression in MDA-MB-231 triple-negative breast cancer cells: the noncanonical Notch pathway. Medical Research Archives, [online] 12(10). https://doi.org/10.18103/mra.v12i10.5814
COPYRIGHT: © 2024 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v12i10.5814
ISSN 2375-1924
Abstract
The Notch pathway has two general regulatory mechanisms: its canonical pathway and the non-canonical pathway. The non-canonical Notch pathway is known that acts independently of its transcriptional factor to activate its target genes which do not belong to the HER, HES and HERP gene family, and has been linked to oncogenic processes and immune cell activation. Here we report the behavior of triple-negative breast cancer cells that maintain high activity of the non-canonical Notch pathway at the presence of ReNeg-AID a peptide which seems to be provoking both a change in the phenotype of the triple-negative breast cancer cells from epithelial cell to non-functional mesenchymal cell, as well as a cell’s attempt to regain canonical Notch pathway activity. Finally, overexpression of the ReNeg-AID peptide in the triple-negative breast cancer cell line promoted a negative regulation of its transcriptional factor, intensifying the non-canonical Notch pathway activity and causing an oscillation or combination between the activities of the canonical and non-canonical mechanisms of Notch pathway due to the activation of Notch-1 and 3 receptors and the repression of Notch-2 and 4. Added to this must be the interactions that exist between Notch pathway with other signaling pathways that may also be in a state of deregulation in the cancer microenvironment.
Introduction
The non-canonical Notch pathway (NP²) was described by Aster and Pear in 1990. They discovered a chromosomal translocation of the intracellular domain of the Notch-1 gene receptor in T-cell acute lymphoblastic leukemia causing constitutive canonical Notch pathway (NP) activation, which causes T cells to be unable to differentiate to T helper (Th) cells and activate into Th1, Th2 and Th17 cells forms, as T cell activation requires the mechanism of NP² to activate specific interleukins for each of the T helper cells activation forms. From this discovery of a new mechanism of the Notch pathway, its study was extended to more cell types such as lung, ovaries, blood system, bone, and epithelium; and this coincides to organs and systems where either the NP² or NP are deregulated or have changed their activation rhythms caused by a carcinogenic event¹².
Two mechanisms have been postulated that can activate NP² and they are:
-
Ligand-dependent or ligand-independent activation.
The Notch receptor physically interacts in the membrane with certain proteins of other signaling pathways such as Hedgehog (Hh), Wingless (Wnt), Nuclear factor kappa B (NF-kappa B) or Catenin beta-1 (beta-catenin), or, once the Notch receptor is proteolytically processed by the gamma secretase (gamma-secretase) complex and released as Notch intracellular domain (NICD); in the course from the membrane to the nucleus NICD interacts with the same proteins mentioned above, this makes NP² also dependent and independent of gamma-secretases³⁵. -
The Notch pathway transcriptional factor (CSL)-independent activation.
This is based on the capacity of the Notch receptor to bind with certain transcriptional factors and increase their half-life in the cytoplasm. These transcriptional factors such as hypoxia-inducible factor 1 alpha subunit (HIF-1A), the transcription repressor YY1 (YY1) or beta-catenin; these interactions promotes the activation of the target genes downstream of these signaling pathways to have a stricter but effective level of homeostatic regulation⁶⁷.
The role of NP is strictly necessary in the embryonic stage. NP² seems to be exclusive in fully differentiated adult organism processes and acts in regulatory processes of the same NP and other pathways such as NF-kappa B, PI3K/Akt/mTOR pathway, Hh and Wnt, it’s also involved in regulatory processes of apoptosis through the breast cancer type 1 susceptibility protein (BRCA1) and with physical interactions with mitochondria, related to the regulation of cancer cell metabolism under the interaction with beta-catenin⁸¹⁴.
The mechanisms of NP² are confounded by its ambiguous nature of interacting with many proteins from other signaling pathways. This intricate nature has provided new ways of regulation for NP itself and for the cellular events involved such as differentiation, apoptosis, cell cycle and metabolism. The instigation of a new perspective to understand a new mechanism of gene regulation, not exclusive to NP but in the complex network of interactions that occur in the background between many signaling pathways to maintain and regulate vital processes, if this delicate mechanism is altered, as with any regulatory process will cause complications in the functioning and homeostasis of the organism.
Methods
For cell culture:
Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco, Cat. No. 12100046) for MDA-MB-231 cells, DMEM-F12 Ham (Gibco, Cat. No. 12500096) for MCF12-F cells, fetal bovine serum (FBS, Gibco, Cat. No. 10437028), penicillin, streptomycin, and trypsin-ethylenediaminetetraacetic acid (trypsin-EDTA, supplied by GIBCO-USA). For DMEM-F12 Ham complete growth medium, 20 ng/ml of epidermal growth factor (Gibco, Cat. No. PHG0315), 0.01 mg/ml of human insulin, and 500 ng/ml of hydrocortisone were added. The plasmid used for cell transfection (pFN21A HaloTag® CMV Flexi® Vector) was acquired by Promega™.
ReNeg-AID peptide.
The ReNeg-AID peptide (patent 415236) was previously described at Saucedo-Correa et al. (2021)¹⁵.
Cell culture.
The non-cancer cell lines of human epithelial breast MCF-12F (ATCC CRL-3599) and tumorous triple-negative MDA-MB-231 (ATCC CRM-HTB-26) cells were purchased from ATCC (Manassas, VA, USA).
Transfection & electroporating.
Cells were incubated at 5% of CO₂ at 37°C until reaching 80% confluence. The culture was then tripanized and 1 × 10⁶ cells were harvested and centrifuged until the pill was formed. Then, a 100 μl nucleofection solution (Cell Line Nucleofector® Kit V, protocol T/C-28a2, AMAXA®) was prepared with 2 ng of the plasmid pReNeg-AID and 400 μl of OptiMEM medium (GIBCO-USA, Cat. No. 31985062). The cells were resuspended in nucleofection solution and then were placed in a 4 mm electroporating cuvette in an ECM 630 electroporator (BTX, Harvard Apparatus system, Item #45-2031). Electroporating conditions: 140 V, 70 ms with one pulse; then the cuvette with the cell solution was incubated for five minutes at room temperature between 18°C and 25°C, and the cells were cultured in six-well plates with 1.5 ml of supplemented DMEM in a humidified 37°C/5% CO₂ incubator for 48 h.
Quantitative real time PCR (RT-qPCR).
RT² Profiler PCR (Qiagen, Cat. PAHS-059Z, No. 330231) Notch related gene arrays: Total cell RNA was isolated from MCF12-F and MDA-MB-321, subsequently treated with DNase I, and purified using the RNeasy Mini Kit (Qiagen, Cat. No. 74034). 25 μg of high-quality total RNA was then reverse transcribed using the First Strand Synthesis Kit (Qiagen Cat. No. 330401) and subsequently loaded onto the Human Notch RT2 profiler array (PAHS-059Z). The real time PCR was performed by using SYBR Green as a marker for DNA amplification on a thermocycler StepOnePlus™ System (Applied Biosystems, Thermo Fisher Scientific).
Flow cytometry.
Both the control cells and ReNeg-AID cells were harvested 48 hours post-transfection. A total of 1 × 10⁴ cells in PBS were dyed with 400 μl of IP, followed by adding 50 μl of RNAse and incubated for 30 minutes to 1 hour. Data were collected on the Attune™ NXT Flow Cytometer (Thermo Fischer Scientific) using the BL2-H (lin)/Histogram channel to obtain the cell cycler phases graphics with the following parameters: FCS-260 V, SSC-280 V and BL2-360 V. Analysis was performed with FlowJo software V. 10 (Tree Star, Inc.).
Statistical analysis.
MTT proliferation, luciferase assay and cytometry analysis were analyzed by student’s t-test, and values with P < 0.05 were considered statistically significant. All experiments were performed at least three times, with n = 4 for each of them.
For gene expression, Qiagen’s online web analysis program was used to produce comparative scatter plots, and fold change was calculated by determining the ratio of mRNA levels to control values normalized to an average of two of these housekeeping genes, ACTB and GAPDH. All experiments were performed at least three times, with n = 4 for each of them.
Results
Notch pathway inhibitor-related genes and cell differentiation.
The NP inhibitor-related genes analyzed were amino-terminal enhancer of split (AES), catenin beta 1 (CTNNB1), deitex E3 ubiquitin ligase 1 (DTX1), GLI family zinc finger 1 (GLI1), nuclear receptor corepressor 2 (NCOR2), O-fucosyltransferase 1 (POFUT1) and smoothened, frizzled class receptor (SMO). Figure 1A shows the effect of ReNeg-AID peptide overexpression in MDA-MB-231 control versus ReNeg-AID cells 48 hours post-transfection (hpt). The AES, DTX1 and GLI1 genes presented a positive up-regulation with 8-, 3- and 0.68-fold change with a p-value of 0.002337, 0.00004 and 0.0008 respectively. The NCOR2 gene presented a negative regulation with 3-fold change with a p-value of 0.0005. CTNNB1, POFUT1 and SMO genes did not show significant changes.
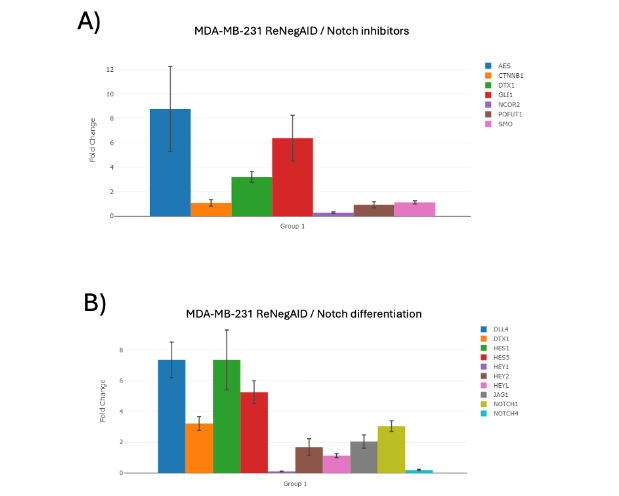
Figure 1. Times of change of NP-dependent genes in MDA-MB-231 ReNeg-AID versus MDA-MB-231 control cells. A) NP inhibitory genes. B) NP-related differentiation genes. ACTB and GAPDH genes were used to normalize mRNA quantifications. The normalized threshold in times of change is ± 2-fold. Genes with statistically significant results are marked with a [*]; (P < 0.05), with n=4.The differentiation-related genes analyzed were: delta like canonical Notch ligand 4 (DLL4), DTX1, hes family bHLH transcription factor 1 (HES1), hes family bHLH transcription factor 5 (HES5), hes related family bHLH transcription factor with YRPW motif 1 (HEY1), hes related family bHLH transcription factor with YRPW motif 2 (HEY2), hes related family bHLH transcription factor with YRPW motif like (HEYL), jagged canonical Notch ligand 1 (JAG1), notch receptor 1 (NOTCH1) and notch receptor 4 (NOTCH4). Figure 1B shows the effect of ReNeg-AID peptide overexpression in MDA-MB-231 control versus ReNeg-AID cells 48 hpt. The DLL4 and JAG1 gene presented a 7.35 and 2.54-fold positive up-regulation with a p-value of 0.000022 and 0.0021 respectively; the NOTCH1 gene presented a 3.05-fold change up-regulation with a p-value of 0.00001, while the NOTCH4 gene presented a -5.09-fold negative down-regulation with a p-value of 0.00005. As well as a positive expression of the HES1 and HES5 genes presented a 7.35- and 5.25-fold change up-regulation with a p-value of 0.00001 and 0.00003 respectively.The Notch and Hedgehog pathways communication.
The NP genes related to the Hh pathway analyzed were: GLI1, glycogen synthase kinase 3 beta (GSK3B), HES5, NOTCH4, sonic hedgehog signaling molecule (SHH), SMO and SUFU negative regulator of hedgehog signaling (SUFU). Figure 2A shows the effect of ReNeg-AID peptide overexpression in MDA-MB-231 control versus ReNeg-AID cells 48 hpt. It was observed that the GLI1 and HES5 genes showed a positive regulation in their times of change, together with the negative regulation of the NOTCH4. Specially the SUFU gene presented a positive regulation of 14.58-fold change up-regulation and a p value of 0.000617.
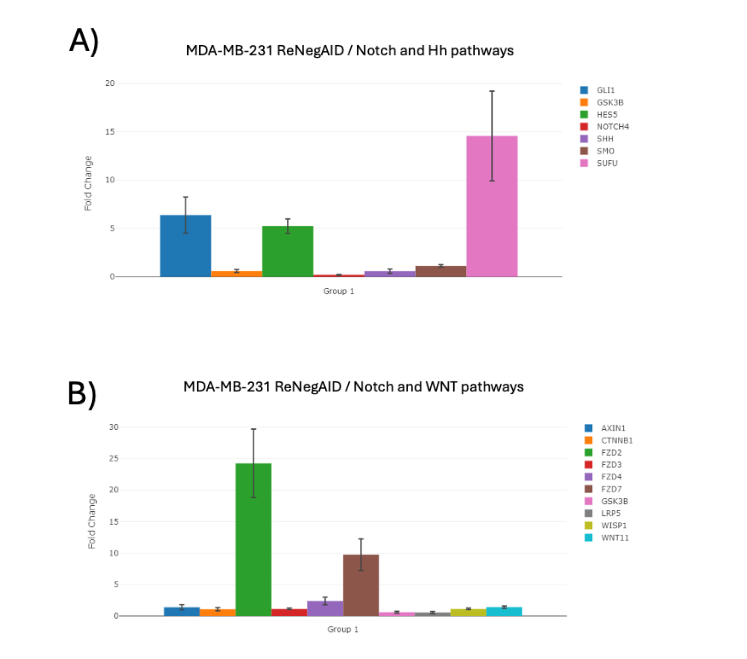
Figure 2. Change times of NP-dependent genes related to the Hh and Wnt pathway in MDA-MB-231 ReNeg-AID versus MDA-MB-231 control cells. A) Hh pathway-related genes. B) Wnt pathway-related genes. ACTB and GAPDH genes were used to normalize mRNA quantifications. The normalized threshold in times of change is ± 2-fold. Genes with statistically significant results are marked with a [*]; (P < 0.05), with n = 4.
The Notch and Wingless pathways communication
The NP genes related to the Wnt pathway analyzed were:
axin 1 (AXIN1), CTNNB1, frizzled class receptor 2 (FZD2), frizzled class receptor 3 (FZD3), frizzled class receptor 4 (FZD4), frizzled class receptor 7 (FZD7), GSK3B, LDL receptor related protein 5 (LRP5), WNT1 inducible signaling pathway protein 1 (WISP1) and Wnt family member 11 (WNT11).
Figure 2B shows the effect of ReNeg-AID peptide overexpression in MDA-MB-231 control versus ReNeg-AID cells 48 hpt.
The genes with significant changes were FZD2, FZD4 and FZD7, which presented a positive regulation with 24.24-, 2.39- and 9.74-fold change with a p value of 0.0001, 0.0002 and 0.0003 respectively.
AXIN1, CTNNB1, FZD4, GSK3B, LRP5, WISP1 and WNT11 genes showed no significant changes.
Apoptosis-related genes
The apoptosis-related NP genes analyzed were:
AXIN1, CASP8 and FADD like apoptosis regulator (CFLAR), CTNNB1, Fos proto-oncogene, AP-1 transcription factor subunit (FOS), FOS like 1, AP-1 transcription factor subunit (FOSL1), interleukin 2 receptor subunit alpha (IL2RA), neuralized E3 ubiquitin protein ligase 1 (NEURL1), nuclear receptor subfamily 4 group A member 2 (NR4A2) and pre T cell antigen receptor alpha (PTCRA). Figure 3A shows the effect of ReNeg-AID peptide overexpression in MDA-MB-231 control versus ReNeg-AID cells 48 hpt. The only genes that showed significant changes were FOS and NEURL1, both with a positive regulation in their change times with 9.19- and 6.40-fold change and a p-value of 0.00019 and 0.0002 respectively.
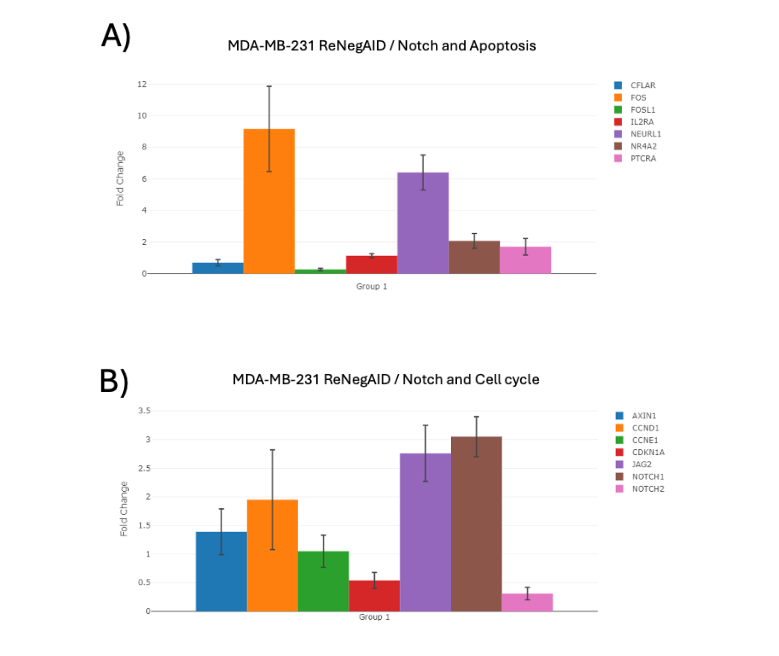
Figure 3. Change times of NP-dependent genes related apoptosis and cell cycle in MDA-MB-231 ReNeg-AID versus MDA-MB-231 control cells.
A) Apoptosis-related genes.
B) Cell cycle-related genes.
ACTB and GAPDH genes were used to normalize mRNA quantifications. The normalized threshold in times of change is ± 2-fold. Genes with statistically significant results are marked with a [*]; (P < 0.05), with n=4.
Cell cycle-related genes.
The cell cycle-related NP genes analyzed were:
AXIN1, cyclin D1 (CCND1), cyclin E1 (CCNE1), cyclin-dependent kinase 1 S homeolog (CDK1A), jagged canonical Notch ligand 2 (JAG2) and notch receptor 2 (NOTCH2).
Figure 3B shows the effect of ReNeg-AID peptide overexpression in MDA-MB-231 control versus ReNeg-AID cells 48 hpt.
The only genes that showed significant changes were JAG2 and NOTCH1 with a positive regulation in their change times of 2.76- and 3.05-fold change with a p-value of 0.0001 and 0.00001 respectively, while the NOTCH2 gene underwent a negative regulation in its change times of -3.25-fold change with a p-value of 0.0016.
Discussion
Notch pathway inhibitor-related genes and cell differentiation.
The positive regulation of DTX1, AES and the negative regulation of NCOR2 indicates that the canonical NP is being negatively regulated at the level of protein complex formation by the action of AES and at the cytoplasmic level by the action of DTX1¹⁶ ¹⁷ (Fig. 1A), however, theGLI1 gene could be promoting the expression of the Hypoxia-inducible factor 1-alpha (HIF-1α) factor of the Hh pathway and thus contributing to the positive regulation of NP²; although the effect of the interaction of the Notch receptor with HIF-1a assists the expression of genes towards the oxidative stress response such as BCL2 binding component 3 (PUMA), which encodes to a protein that cooperates with direct activator proteins to induce mitochondrial outer membrane permeabilization and apoptosis, or BCRA; it has also been reported that the HIF-1a factor in turn regulates the intensity of the expression of NF-kappa B dependent genes involved in the recognition response of the immune system such as IL-6 and 17 as well as the metabolism of cancer cells.
The fact that CTNNB1, POFUT1 and SMO genes did not show a change in expression after transfection with pReNeg-AID could indicate that the aberrant state of the Wnt pathway has not undergone changes in this cell type; the case of the POFUT1 gene suggests a correlation between the ligand-dependent pathway that NP² can exert⁴⁹,¹⁷.
On the other hand, the discrepancy between the positive expression of Delta4, Jagged1 ligands and Notch-1 receptor but a negative expression of Notch-4 receptor would suggest that NP² is being promoted in its ligand-dependent form at a higher frequency (Fig. 1B), however, the positive expression of the HES1 and HES5 genes suggests that NP is being favored with ReNeg-AID peptide expression and this suggests a regulation and attempt to regain cellular identity in this type of mammary glandular tissue, however further evidence is needed to fully elucidate the NP² machinery and its ultimate implications with the other signaling pathways related to cell differentiation¹⁶–¹⁸.
The expression of HEY1, HEY2 and HEYL genes did not present significant changes, this coincides with what has been reported for the differential phenotype of this cell type⁷,¹⁸–²¹.
The Notch and Hedgehog pathways communication.
The genes with a positive regulation (Fig. 2A) in their times of change such as GLI1 and HES5, together with the negative regulation of the NOTCH4 gene suggests that the regulatory effect that occurs between NP and the Hh pathway through the Notch-4/HIF-1a receptor is being negatively regulated, implying that cell migration or metastatic behavior reported by this interaction is being promoted, however, a positive regulation of the SUFU gene means that the NP² is compromised, since the SUFU gene codes for the protein with the same name and is a negative regulator of Hh pathway activation. This could indicate an approach to understand that, although the NP² via HIF-1a factor and GLI1 protein is being activated by MDA-MB-231 cells, the overexpression of the SUFU gene promoted by the ReNeg-AID peptide prevents the metastatic phenotype in this cell line from being mechanistically non-functional²²,²³.
The Notch and Wingless pathways communication.
In this context between the intercommunication between NP and NP² suggests that NP regulates its activity by being activated by positive expression of its FZD ligand gene (Fig. 2B), however, WISP1 and WNT11 ligands genes did not undergo significant changes causing the Wnt pathway to be unaffected by NP, but the effects of NP² are uncertain in this context and require further analysis in understanding the molecular mechanisms that may be affected²⁴,²⁵.
Apoptosis-related genes.
The positive expression of FOS and NEURL1 genes (Fig. 3A) suggest an up regulation of NP-dependent apoptotic processes by the intrinsic pathway. Nevertheless, the MDA-MB-231 cells cannot terminate their process of apoptosis signaling path. These should be possible because the FOS gene is dependent on the joint function with FOSL1 gene who showed a diminished expression; and the NEURL1 gene is not functional since this gene and NR4A gene, who showed a diminished expression, are known to be both the last mediators in the apoptosis signaling steps under the control of NP¹²⁶. All this data together suggests that the attempt to recover this apoptosis signaling path in the NP context fails because some of the proteins involved are not present.
Cell cycle-related genes.
The fact that CCND1 and CCNE1 genes showed no significant changes under the influence of the ReNeg-AID peptide can be directly related to a non-functional canonical NP (Fig. 3B), because both genes are direct targets of canonical NP. On the other hand, the ReNeg-AID peptide expression promotes a positive regulation for JAG2 and NOTCH1 which in the MDA-MB-231 cell type context are strictly required for differentiation (Fig. 3B), however, the negative regulation of the Notch-2 receptor indicates that the cell cycle is not being adequately regulated by NP, or, alternatively, it is promoting a transcriptional factor CSL-dependent NP² by presenting a regulation of the Notch-1 receptor but not of the Notch-2 receptor; recalling that the Notch-2 receptor in mammary gland epithelial cells is mainly responsible for activating the cell cycle and carrying out the final progression of differentiation in this cellular context²⁷–²⁹.
Previously, Saucedo-Correa et al. (2021), had been reported that in a MTT proliferation assay the MDA-MB-231 cell line presented an irregular proliferation, due to its genetic background with respect to the NP that is activated in its non-canonical way, which is more evident when the ReNeg-AID peptide has no effect on non-cancerous MCF-12F mammary gland cells at any of the tested time, and cell proliferation remains unaltered.
Conservation of NP cellular communication
Conservation of NP cellular communication mechanism at the molecular level over time and under the inherent effects of evolution, has allowed the emergence of specific clades of metazoans from simple animals such as sponges to complex animals such as mammals. NP is part of the 7 major pathways that give rise to the intricate cellular and systemic morphological patterns of metazoans and at the same time with the interaction of other signaling-pathways promotes cell-cycle regulation, apoptosis, differentiation, and cell migration. These cellular events are the key points for maintaining the state of homeostasis that is characteristic of a healthy state of the individual. If the NP mechanism ceases to function properly it promotes a drastic change in the dynamics of information flow and cell-cell communication. These changes manifest themselves in a loss of cellular identity, meaning that the cell has stopped receiving or has increased the number of commands to other cells; stimulating the cell cycle to lose its self-regulation, apoptosis to fail to activate, cell migration to be activated in cell types that no longer need it, causing the cell to adopt a cancerous and metastatic behavior.With the above described and integrating the results obtained in this research and based on the principle of conservation of the domains that make up the family of proteins belonging to and related to NP, we were able to design a molecular switch from a domain of the H protein of D. melanogaster¹⁵ that was used to bind the transcriptional factor CSL, which negatively regulates the activation of the target genes of NP in human mammary epithelial cancer cell lines that express Notch constitutively. Although mammalian mechanisms of negative regulation of NP exist; the mammalian genome has apparently lost the hairless protein gene that has been found only in insects and mollusks, however, the mammalian CSL transcriptional factor is still able to recognize the Hairless protein domain¹⁵,¹⁶. This recognition may indicate that the loss of a gene does not necessarily mean that the entire system has lost the ability to use or recognize it, and that the H protein functions as a synapomorphy for insects and mollusks.The understanding of this mechanism will benefit and facilitate new and future ways to take advantage of this quality in scenarios where constitutive activation of NP is present and is the origin of diseases such as cancer in different tissues. This also implies that the same method can be applied to design strategies in scenarios where NP is constitutively repressed and even to design peptides with the same principle that function in other signaling pathways with malfunctioning mechanisms.Notch-1 receptor but maintains normal Notch-4 receptor levels in MCF-7 cells¹⁵; however, in MDA-MB-231 cells it promotes an opposite effect, negatively regulating the Notch-4 receptor and positively regulating the Notch-1 receptor. While the effect of ReNeg-AID peptide under the context of the canonical Notch pathway works in a predicted manner in MCF-7 cells; but the effect in MDA-MB-231 cells is unclear, however, it retains certain patterns suggesting that NP² is promoted by its two described forms; ligand-dependent and ligand-independent and CSL-independent.The main effect of the ReNeg-AID peptide in MCF-7 cells was the interruption of the cell cycle in the transition from G1 to S phase¹⁵; this has a direct impact on the arrest of uncontrolled cell growth characteristic of cancer. In contrast, the main effect in MDA-MB-231 cells was not on the cell cycle but on the cross talk that exists between the NP and the Hh pathway, promoting the positive regulation of the SUFU protein and consequently negatively regulating the expression of Hh pathway target genes that in the context of cancer are directly related to cell metastasis and cancer metabolism.
It is advisable to do more detailed studies, such as step-by-step analysis of the possible interactions that can occur with Notch receptors with undescribed proteins belonging to the Hh and Wnt pathways; since cross-communication between these three pathways govern and determine much of the nature of cancer as well as promote its occurrence and maintain its homeostatic state of cancer itself, and thus understand whether the non-canonical Notch pathway serves as a regulatory, promoter or inhibitory mechanism in cellular processes that regulates canonically way.
Declaration of interest:
None.
Funding sources:
GSC recently received a Ph.D. scholarship from CONAHCYT (registration number 270234). ABP has received SEP-CONAHCYT funding (registration number CB-2011-01 167947) and CIC-UMSNH funding (registration number 521765 CIC 2015), which help to support the development of this manuscript. The remaining authors have no financial interests.
Acknowledgements:
None.
ORCID ID:
-
Germán Saucedo-Correa, ORCID 0000-0003-1468-1268
-
Rosa E. Nuñez-Anita, ORCID 0000-0002-1117-9136
-
Ulises Maciel-Ponce, ORCID 0009-0003-1524-9474
-
Humberto Contreras-Cornejo, ORCID 0009-0000-1713-6244
-
Alejandro Bravo-Patiño, ORCID 0000-0001-9897-7634
References
[1]. Sade H, Krishna S, Sarin A. The Anti-apoptotic Effect of Notch-1 Requires p56lck-dependent, Akt/PKB-mediated Signaling in T Cells. J. Biol. Chem. 2004;279:2937–2944. https://doi.org/10.1074/jbc.m309924200.
[2]. Allenspach EJ, Maillard I. Notch Signaling in Cancer. Cancer Biol. Ther. 2002;1:466–476. https://doi.org/10.4161/cbt.1.5.159.
[3]. Behrens J, Lustig B. The Wnt connection to tumorigenesis. Int. J. Dev. Biol. 2004;48:477–487. https://doi.org/10.1387/ijdb.041815jb.
[4]. Di Mauro C, Rosa R, D’Amato V, Ciciola P, Servetto A, Marciano R, Orsini RC, Formisano L, De Falco S, Cicatiello V, Di Bonito M, Cantile M, Collina F, Chambery A, Veneziani BM, De Placido S, Bianco R. Hedgehog signalling pathway orchestrates angiogenesis in triple-negative breast cancers. Br. J. Cancer. 2017;116:1425–1435. https://doi.org/10.1038/bjc.2017.116.
[5]. Dongre A, Surampudi L, Lawlor RG, Fauq AH, Miele L, Golde TE, Minter LM, Osborne BA. Non-canonical Notch signaling drives activation and differentiation of peripheral CD4+ T cells. Front. Immunol. 2014;5:1–14. https://doi.org/10.3389/fimmu.2014.00054
[6]. Maier D. The evolution of transcriptional repressors in the Notch signaling pathway: a computational analysis. Hereditas. 2019;156:5. https://doi.org/10.1186/s41065-019-0081-0
[7]. Zeng C, Xing R, Liu J, Xing F. Role of CSL-dependent and independent Notch signaling pathways in cell apoptosis. Apoptosis. 2016;21:1–12. https://doi.org/10.1007/s10495-015-1188-z
[8]. Ayaz F, Osborne BA. Non-canonical Notch signaling in cancer and immunity, Front. Oncol. 2014;4:1–7. https://doi.org/10.3389/fonc.2014.00345
[9]. Brechbiel J, Miller-Moslin K, Adjei AA. Crosstalk between hedgehog and other signaling pathways as a basis for combination therapies in cancer. Cancer. Treat. Rev. 2014;40:750–759. https://doi.org/10.1016/j.ctrv.2014.02.003.
[10]. Salmena L, Carracedo A, Pandolfi PP. Tenets of PTEN Tumor Suppression. Cell. 2008;133:403–414. https://doi.org/10.1016/j.cell.2008.04.013.
[11]. Hossain F, Sorrentino C, Ucar DA, Peng Y, Matossian M, Wyczechowska D, Crabtree J, Zabaleta J, Morello S, Del Valle L, Burow M, Collins-Burow B, Pannuti A, Minter LM, Golde TE, Osborne BA, Miele L. Notch Signaling Regulates Mitochondrial Metabolism and NF-kB Activity in Triple-Negative Breast Cancer Cells via IKKa-Dependent Non-canonical Pathways. Front. Oncol. 2018;8:575. https://doi.org/10.3389/fonc.2018.00575.
[12]. Minter LM, Osborne BA. Canonical and non-canonical Notch signaling in CD4+ T cells. Curr. Top. Microbiol. Immunol. 2012;360:99–114. https://doi.org/10.1007/82_2012_233.
[13]. Kyu-Sun Lee L, Wu Z, Song Y, Mitra SS, Feroze AH, Cheshier SH, Lu B. Roles of PINK1, mTORC2, and mitochondria in preserving brain tumor-forming stem cells in a noncanonical Notch signaling pathway. Genes Dev. 2013;27:2642–2647. https://doi.org/10.1101/gad.225169.113.
[14]. Thorsen J, Micci F, Heim S. Identification of chromosomal breakpoints of cancer-specific translocations by rolling circle amplification and long-distance inverse PCR. Cancer Genet. 2011; 204:458–461. https://doi.org/10.1016/j.cancergen.2011.07.007.
[15]. Saucedo-Correa G, Nuñez-Anita RE, Maciel-Ponce U, Contreras-Cornejo H, Bravo-Patiño A. A peptide derived from D. melanogaster Hairless protein promotes the negative regulation of Notch aberrant constitutive signaling on human breast cancer cells. Int. J. Clin. Exp. Med. 2021;14:62-75. https://e-century.us/files/ijcem/14/1/ijcem0108892.pdf.
[16]. Contreras-Cornejo H, Saucedo-Correa G, Oviedo-Boyso J, Valdez-Alarcón JJ, Baizabal-Aguirre VM, Cajero-Juárez M, Bravo-Patiño A. The CSL proteins, versatile transcription factors and context dependent corepressors of the notch signaling pathway. Cell. Div. 2016;11:12. https://doi.org/10.1186/s13008-016-0025-2.
[17]. Pedrosa AR, Trindade A, Carvalho C, Graça J, Carvalho S, Peleteiro MC, Adams RH, Duarte A. Endothelial Jagged1 promotes solid tumor growth through both pro-angiogenic and angiocrine functions. Oncotarget. 2015;6:24404-24423. https://doi.org/10.18632/oncotarget.4380.
[18]. Weijzen S, Rizzo P, Braid M, Vaishnav R, Jonkheer SM, Zlobin A, Osborne BA, Gottipati S, Aster JC, Hahn WC, Rudolf M, Siziopikou K, Kast WM, Miele L. Activation of Notch-1 signaling maintains the neoplastic phenotype in human Ras-transformed cells. Nat. Med. 2002;8:979–986. https://doi.org/10.1038/nm754.
[19]. Asnaghi L, Lin MH, Lim KS, Lim KJ, Tripathy A, Wendeborn M, Merbs SL, Handa JT, Sodhi A, Bar EE, Eberhart CG. Hypoxia promotes uveal melanoma invasion through enhanced notch and MAPK activation. PLoS One. 2014;9:e105372. https://doi.org/10.1371/journal.pone.0105372.
[20]. Hartman J, Müller P, Foster JS, Wimalasena J, Gustafsson JA, Ström A. HES-1 inhibits 17beta-estradiol and heregulin-beta1-mediated upregulation of E2F-1. Oncogene. 2004;23:8826–8833. https://doi.org/10.1038/sj.onc.1208139.
[21]. Corbin EA, Kong F, Lim CT, King WP, Bashir R. Biophysical properties of human breast cancer cells measured using silicon MEMS resonators and atomic force microscopy. Lab. Chip. 2015;15:839–847. https://doi.org/10.1039/c4lc01179a.
[22]. Karamboulas C, Ailles L. Developmental signaling pathways in cancer stem cells of solid tumors. Biochim. Biophys. Acta. 2013;1830:2481–2495. https://doi.org/10.1016/j.bbagen.2012.11.008.
[23]. Zhang J, Tian XJ, Xing J. Signal Transduction Pathways of EMT Induced by TGF-β, SHH, and WNT and Their Crosstalks. J. Clin. Med. 2016;5:41. https://doi.org/10.3390/jcm5040041.
[24]. Guo S, Liu M, Gonzalez-Perez RR. Role of Notch and its oncogenic signaling crosstalk in breast cancer. Biochim. Biophys. Acta. 2011;1815:197-213. https://doi.org/10.1016/j.bbcan.2010.12.002.
[25]. Sanchez-Vega F, Mina M, Armenia J, Chatila WK, Luna A, La KC, Dimitriadoy S, Liu DL, Kantheti HS, Saghafinia S, Chakravarty D, Daian F, Gao Q, Bailey MH, Liang WW, Foltz SM, Shmulevich I, Ding L, Heins Z, Ochoa A, Gross B, Gao J, Zhang H, Kundra R, Kandoth C, Bahceci I, Dervishi L, Dogrusoz U, Zhou W, Shen H, Laird PW, Way GP, Greene CS, Liang H, Xiao Y, Wang C, Lavarone A, Berger AH, Bivona TG, Lazar AJ, Hammer GD, Giordano T, Kwong LN, McArthur G, Huang C, Tward AD, Frederick MJ, McCormick F, Meyerson M, Van Allen EM, Cherniack AD, Ciriello G, Sander C, Schultz N. Oncogenic Signaling Pathways in The Cancer Genome Atlas. Cell. 2018;173:321-337. https://doi.org/10.1016/j.cell.2018.03.035.
[26]. Perumalsamy LR, Nagala M, Banerjee P, Sarin A. A hierarchical cascade activated by non-canonical Notch signaling and the mTOR-Rictor complex regulates neglect-induced death in mammalian cells. Cell Death & Differentiation. 2009;16:879–889. https://www.nature.com/articles/cdd200920.
[27]. Wang Z, Zhang Y, Li Y, Banerjee S, Liao J, Sarkar FH. Down-regulation of Notch-1 contributes to cell growth inhibition and apoptosis in pancreatic cancer cells. Mol. Cancer Ther. 2006;5:483–493. https://doi.org/10.1158/1535-7163.mct-05-0299.
[28]. Arnold A, Papanikolaou A. Cyclin D1 in breast cancer pathogenesis. J. Clin. Oncol. 2005;23:4215 –4224. https://doi.org/10.1200/jco.2005.05.064.
[29]. Sweeney KJ, Swarbrick A, Sutherland RL, Musgrove EA. Lack of relationship between CDK activity and G1 cyclin expression in breast cancer cells. Oncogene. 1998;16:2865–2878. https://doi.org/10.1038/sj.onc.1201814.

