Targeting KLF Pathways with Metadichol: Anticancer Strategy
Synergistic Targeting of Krüppel-like factor and Related Signaling Pathways by Metadichol: A Multidimensional Anticancer Strategy
P. R. Raghavan1
- Nanorx Inc. USA, PO Box 131, Chappaqua NY 10514.
OPEN ACCESS
PUBLISHED: 30 June 2025
CITATION: Raghavan, P.R., 2025. Synergistic Targeting of Krüppel-like factor and R Related Signaling Pathways by Metadichol: A Multidimensional Anticancer Strategy. Medical Research Archives, [online] 13(6).
https://doi.org/10.18103/mra.v13i6.6583
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v13i6.6583
ISSN 2375-1924
ABSTRACT
Metadichol, a nanoemulsion of long-chain lipid alcohols, demonstrates a unique capacity to modulate the expression of the entire (KLF) transcription factor family (KLF1–KLF18) in human peripheral blood mononuclear cells (PBMCs) in a concentration-dependent manner. Using qRT-PCR and Western blot techniques, at 1 ng/ml, Metadichol downregulated 14 of 18 KLFs while selectively upregulating KLF4, KLF15, KLF17, and KLF18—factors often implicated in tumor suppression or context-dependent cancer regulation. Lower and higher concentrations produced distinct, biphasic expression patterns, indicating complex dose-dependent regulatory mechanisms. Beyond the KLF family, previous work has shown that metadichol influences interconnected signaling networks involving nuclear receptors, sirtuins, Toll-like receptors, circadian genes, and key tumor suppressors such as TP53 and Klotho. Network analysis suggests that this broad-spectrum modulation may overcome compensatory mechanisms within cancer cells, offering synergistic antitumor effects. The scope of this paper is to comprehensively analyze Metadichol and its dose-dependent regulation of KLFs and related pathways, while the purpose is to establish Metadichol as a multitarget anticancer agent capable of overcoming compensatory mechanisms in cancer cells, with potential applications in other diseases driven by transcriptional dysregulation. By simultaneously targeting multiple pathways, Metadichol offers a novel integrative approach to enhance therapeutic efficacy compared to single-pathway interventions.
Keywords
Metadichol KLF, Toll-like receptors, nuclear receptors, sirtuins, circadian genes, Klotho, TP53, FOXO1, PPARGC1A, telomerase, long-chain lipid alcohols.
List of Abbreviations
- Oct 4 POU class 5 homeobox 1 pseudogene 3
- APOE apolipoprotein E
- CREBBP CREB binding protein
- CRY1 cryptochrome circadian regulator 1
- FOXO1 Forkhead box O1
- GR glucocorticoid receptor or known as NR3C1 (Nuclear receptor subfamily 3, group C, member 1)
- HDAC1 histone deacetylase 1
- KL klotho
- KLF
- MMP9 matrix metallopeptidase 9
- MR Mineralocorticoid receptor also known as NR3C2 (nuclear receptor subfamily 3 group C member 2)
- NFkb Nuclear factor Kappa subunit
- p300 Histone acetyltransferase p300
- p50 Sub-units of NFkb
- p65 Sub-units of NFkb
- PER1 period circadian regulator 1
- PPARa peroxisome proliferator activated receptor alpha
- PPARd peroxisome proliferator activated receptor beta
- PPARg peroxisome proliferator activated receptor gamma
- PPARGC1A PPARG coactivator 1 alpha
- PRP4 (pre-mRNA splicing tri-snRNP complex factor PRPF4
- SOX2 SRY-box transcription factor 2
- SP1 Sp1 transcription factor
- Tert telomerase reverse transcriptase
- TLR Toll Like receptor
- TP53 tumor protein p53
Introduction
Krüppel-like factors (KLFs), a family of 18 mammalian zinc-finger transcription factors, are critical regulators of cellular processes, including differentiation, proliferation, and responses to environmental stress, positioning them as key players in maintaining physiological homeostasis. Dysregulation of KLF expression is implicated in a spectrum of diseases, such as obesity, cardiovascular disease, cancer, and inflammatory disorders. Structurally, KLFs feature a conserved C-terminal DNA-binding domain with three C2H2 zinc fingers that bind GC-rich DNA sequences, while their variable N-terminal regulatory domains confer diverse functional roles. This structural diversity enables KLFs to interact with Sp1 sites, including GC boxes and CACCC boxes, influencing gene expression in a context-dependent manner.
In cancer, KLFs exhibit dual roles as oncogenes or tumor suppressors, modulating cell growth, apoptosis, and metastasis. For instance, KLF4 is essential for inflammatory monocyte differentiation, while KLF8 overexpression in gastric cancer is linked to poor prognosis due to its regulation of glycolysis via GLUT4. KLF4-/- chimerism is related to a decreased number of resident and circulating inflammatory monocytes. KLFs also play significant roles in immune function, with KLF2 controlling T-cell trafficking, and KLF10 regulating colonic macrophage activity. KLF10-deficient animals produce less IL-10 and fewer colonic macrophages. They also produce greater amounts of IL-12p70 after exposure to lipopolysaccharide (LPS). In renal physiology, KLFs are involved in fibrosis progression and glomerular filtration barrier maintenance. Given their involvement in these diverse processes, KLFs are attractive therapeutic targets for cardiovascular disease, cancer, and inflammatory conditions. Obesity, cardiovascular disease, cancer, and inflammatory disorders are linked to dysregulation of KLF expression. KLFs are crucial for optimal physiological function because of their roles in these different disease processes.
Krüppel-like factor in cancer
Cell growth and cancer progression are strongly regulated by KLFs. KLFs participate in many growth signal transduction pathways. The overexpression of these genes can promote or suppress cell proliferation. KLFs can regulate the expression of oncogenes and tumor suppressors. They can even cause cancer. Multiple cancers have altered KLF expression. High KLF8 expression in gastric cancer is associated with a larger tumor size, advanced T and N stages, and shorter patient survival. Different forms of cancer are linked to KLF8. Silencing of KLF8 slows cancer cell glycolysis. The context-dependent activities of KLFs in cancer highlight their intricate roles in carcinogenesis.
Krüppel-like factor in the immune system
Krüppel-like factors are essential for immune cell growth and function. Inflammatory monocyte differentiation requires KLF4. KLF2 modulates chemokine receptor patterns to control T-cell trafficking and recirculation. KLF10 controls intestinal macrophages and causes innate immune colitis. Aortic lesions are more extensive in KLF2+/− ApoE−/− animals than in controls. These findings indicate that KLF2 protects against atherosclerosis. KLF2 expression is downregulated in monocytes from severe atherosclerosis patients. In endothelial and monocyte cells, KLF2 and KLF4 regulate the expression of shear stress-related genes. The various effects of KLFs on immune cells reveal their roles in innate and adaptive immunity.
Therapeutic targeting of KLFs
Krüppel-like factors are attractive therapeutic targets because of their involvement in several illnesses. Modulating KLF expression or activity may be a strategy for the treatment of cardiovascular disease, cancer, and inflammatory illnesses. For example, statins increase endothelial KLF expression, which protects against atherosclerosis. Overexpression of KLF4 prevents atherothrombosis, pulmonary hypertension, and restenosis. KLF13 deficiency in uterine endometrial cells impairs steroid hormone receptor signaling in a mouse model of endometriosis. In the same model, KLF9 deletion in the endometrium increased ectopic lesion formation. Manipulating KLF activity represents a promising treatment strategy for several disorders.
Krüppel-like factor and cellular processes
Krüppel-like factors control proliferation, apoptosis, differentiation, and stem cell maintenance. KLF4 and KLF5 regulate esophageal cancer cell growth, apoptosis, and invasion. KLF5 promotes invasion, whereas KLF4 suppresses proliferation and induces apoptosis. Colorectal cancer tumor suppression by KLF6. In colon cancer cells, insulin promotes FASN expression and proliferation. These essential cellular mechanisms demonstrate the role of KLFs in tissue homeostasis and the regulation of cell destiny.
Genetic regulation by KLFs
Krüppel-like factors regulate gene expression by binding to DNA, and KLF1 is needed for proper globin synthesis. KLF4 affects HBG gene expression in primary erythroid cells. KLF4 preferentially binds to the HBG promoter CACCC region and interacts with CREBBP. In mice, KLF13 controls axonal growth, development, and regeneration. Psychiatric symptoms and ADHD were linked to a de novo heterozygous KLF13 gene variation. Figure 1 provides a summary of the roles of the KLF family of genes in several cellular and physiological processes; these genes exert these effects by regulating the expression of various target genes.
Cross-regulation and Krüppel-like factor family interactions
Cross-regulation between KLF family members complicates their regulatory networks. After arterial damage, KLF4 inhibits neointimal development by inhibiting NF-κB. High KLF7 expression indicates aggressive gastric cancer with a poor prognosis. KLF family members interact with one another, which makes their regulatory effects more complex.
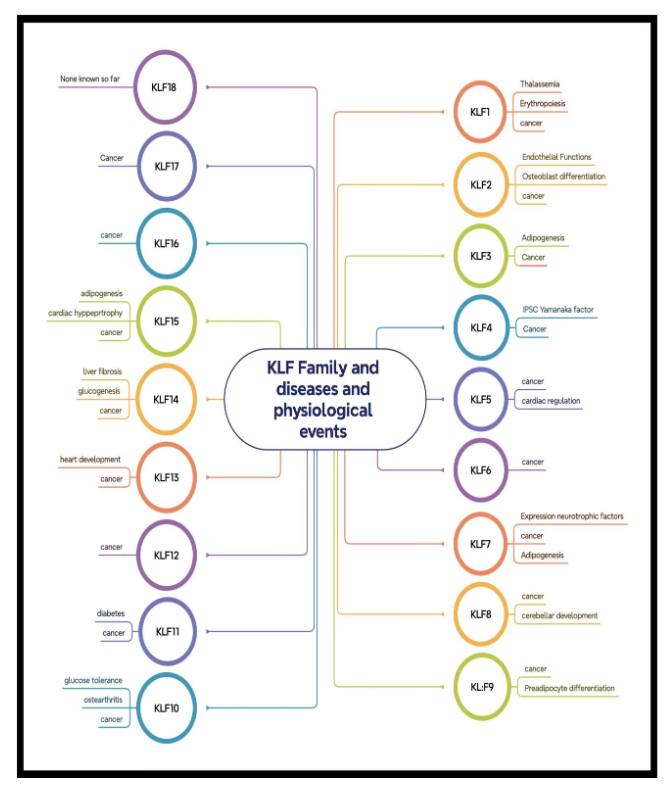
This study investigates the effects of Metadichol, a nanoemulsion of long-chain lipid alcohols, on the expression of the entire KLF family (KLF1–KLF18) in human peripheral blood mononuclear cells (PBMCs), alongside its modulation of interconnected signaling networks, including nuclear receptors, sirtuins, Toll-like receptors, circadian genes, and tumor suppressors such as TP53 and Klotho. The scope of this research is to comprehensively analyze Metadichol’s dose-dependent regulation of KLFs and related pathways using qRT-PCR, Western blot, and network analyses, while the purpose is to establish Metadichol as a multitarget anticancer agent capable of overcoming compensatory mechanisms in cancer cells, with potential applications in other diseases driven by transcriptional dysregulation. By simultaneously targeting multiple pathways, Metadichol offers a novel integrative approach to enhance therapeutic efficacy compared to single-pathway interventions.
Experimental methods
All work was outsourced to Skanda Labs, Bangalore, India. Eurofins Bangalore, India, supplied all the primers. Antibodies were from E-Lab Sciences, Texas, USA. All other reagents and chemicals were from Aldrich chemicals Bangalore, India.
Isolation of human WBCs by cell line and condition.
Blood sample preparation: Fresh human blood from healthy volunteers was collected in EDTA tubes, diluted 1:1 with PBS, and mixed by inverting. Isolation of mononuclear cells: In a 15 ml centrifuge tube, 5 ml of Histopaque-1077 was added, and 5 ml of prepared blood was gently added on top. The tubes were then centrifuged at 400 × g for 30 min at room temperature with the brake off. The upper layer was removed via a Pasteur pipette without disrupting the interphase layer after centrifugation. The interphase layer was carefully placed in a clean centrifuge tube. After being washed with 1× PBS, the cells were centrifuged at 250 × g for 10 min (2×). After centrifugation, the supernatant was discarded, and the pellet was recovered in 10% FBS-RPMI medium. Hemocytometers were used to count the cells and verify their viability. For cell maintenance and seeding, 1 × 106 cells/ml medium were seeded into 6-well plates and incubated for 24 h at 37°C with 5% CO2. After 24 hours of seeding, the medium was carefully removed, and the cells were treated with the selected concentrations of MTT and incubated at 37°C in a CO2 incubator for 24 hours.
Treatment concentrations (Table 1)
| Cell type | Sample name | Treatment details |
|---|---|---|
| Human PBMCs | Metadichol Control | 1 pg/ml, 100 pg/ml, 1 ng/ml, 100 ng/ml |
Sample preparation and RNA isolation
We detached and washed the treated cells with sterile 1× PBS and centrifuged them. The supernatant was decanted, 0.1 ml of TRIzol was added, and the sample was mixed gently by inversion for 1 min. The sample was incubated at room temperature for 10 minutes. To 0.1 ml of TRIzol, 0.75 ml of chloroform was added. The contents were vortexed for 15 seconds. The tube was incubated at room temperature for 5 minutes. The mixture was subsequently centrifuged at 12,000 rpm for 15 minutes at 4°C. The upper aqueous phase was placed in a sterile microcentrifuge tube, 0.25 ml of isopropanol was added, and the mixture was incubated at -20°C for 20 minutes. The contents were subsequently centrifuged at 12,000 rpm for 10 minutes at 4°C. The RNA pellet was rinsed with 0.25 ml of 70% ethanol after the supernatant was discarded. The RNA mixture was subsequently centrifuged at 12,000 rpm at 4°C. The supernatant was carefully removed, and the pellet was air-dried. The RNA pellet was resuspended in 20 µl of DEPC-treated water. The total RNA yield was measured via a Spectra drop (Spectramax i3x, Molecular Devices, USA).
Table 2: RNA yields
| Test concentrations | RNA yield (ng/µl) |
|---|---|
| 0 | 623.120 |
| 1 pg/ml | 343.123 |
| 100 pg/ml | 792.123 |
| 1 ng/ml | 673.111 |
| 100 ng/ml | 611.123 |
cDNA production
In accordance with the manufacturer’s instructions, 500 ng of RNA was used to synthesize cDNA via the Prime Script RT Reagent Kit (TAKARA) and an oligo dT primer. A 20 μl reaction mixture was used for cDNA synthesis at 50°C for 30 min, followed by RT inactivation at 85°C for 5 min utilizing a Veritii Biosystems instrument. cDNA was used to perform real-time PCR.
RT‒qPCR primers and analysis
The PCR mixture (final volume of 20 µl) included 1.4 µl of cDNA, 10 µl of SYBR Green Master Mix, and 1 µM complementary forward and reverse primers (Table 3) for the target genes. Enzyme activation at 95°C for 2 minutes was followed by a 2-step reaction with initial denaturation and an annealing and extension step at 95°C for 5 seconds, annealing for 30 seconds at the appropriate temperature for 39 cycles, secondary denaturation at 95°C for 5 seconds, and 1 cycle with a melt curve capture step from 65°C to 95°C for 5 seconds each. The results were assessed, and the fold change in expression was estimated via CFX Maestro software. The comparative CT approach was used to compare target gene expression to GAPDH expression as follows: delta Ct = target gene – reference gene. The delta CT was determined to compare gene expression between treated cells and control cells under each condition via the following formula: delta delta Ct = treatment group – control group. The gene expression fold change with each treatment was estimated via the following formula: Fold change = 2^(−delta delta Ct).
SDS‒PAGE and Western blot procedure
PROCEDURE: Isolation of Mononuclear Cells In the 15 ml centrifuge tube, 5 ml of Histopaque-1077 was added, and 5 ml of prepared blood was layered on the Histopaque slowly from the edge of the tube without disturbing the histopaque layer. Then tubes were centrifuged at 400 X g for exactly 30 mins at room temperature with brake-off settings. After centrifugation, the upper layer was discarded with a Pasteur pipette without disturbing the interphase layer. The interphase layer was carefully transferred to clean centrifuge tube. Cells will be washed with 1X PBS and again centrifuged at 250 X g for 10 mins. (2X). After centrifugation, the supernatant was discarded, and the pellet will be collected in RPMI media supplemented with 10% FBS. Cells were counted, and viability was checked with a hemocytometer. The cell count was adjusted to 10×106 cells/2 ml. 2 ml of cell suspension is added to each dish in P35 dishes. Cells were then treated with various concentrations of test sample. Post incubation, the cells were harvested for isolation of protein using RIPA buffer. The cells, post harvesting, were washed twice using 1X PBS. The cell pellets were gently suspended in 300 µl of RIPA buffer with 1X Protease Inhibitor. The cells were incubated for 30 mins by gentle mixing every 5 minutes at 4°C.
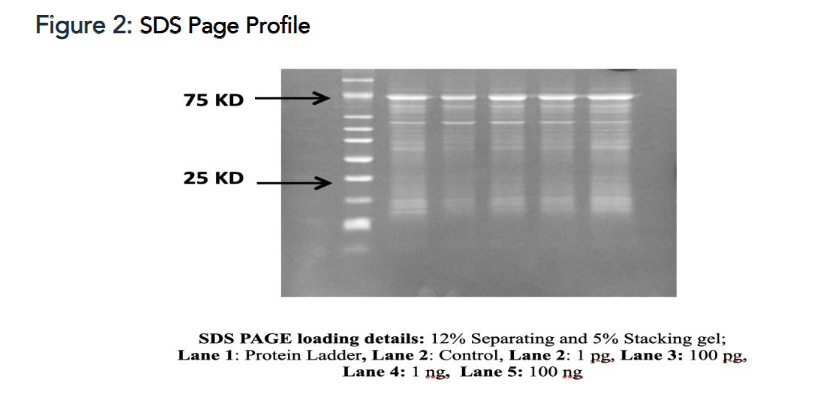
Post incubation, the cells were centrifuged at 10,000 rpm for 12-15 minutes. 25 ug protein sample from each cell lysate was mixed with 5X loading dye and heated for 2 min at 95°C. Protein samples were loaded and separated on 8%, 10 % and 15 % SDS-PAGE gel using Mini protean Tetra cell (Bio-Rad). Nitrocellulose membrane (0.2 µM) was equilibrated in transfer buffer for 10 mins at RT. Protein transfer was done for 15 mins in Turbo Transblot (Bio-Rad) apparatus at 2.5 A and 25 V. Blot was blocked in 3% BSA in TBST for 1 hr at RT followed by incubation with respective 1° Ab at appropriate dilutions O/N at 4°C. Blot was washed thrice with TBST for 5 mins at RT Blot was incubated with 2° Ab (anti-Rabbit or anti-Mouse IgG- HRP) at dilution 1:10000 for 1 hr at RT. Washed 3 times with TBST for 5 mins at RT Blot was rinsed with ECL reagent (two component system) for 1 min in dark and image was captured between 0.5 sec to 15 secs exposure in Chemidoc XRS+ imaging system (Bio-Rad). The protein lysates in the supernatant were transferred to fresh sterile tubes and stored in -20˚C until further use.
Figure 3: Western blot experimental conditions
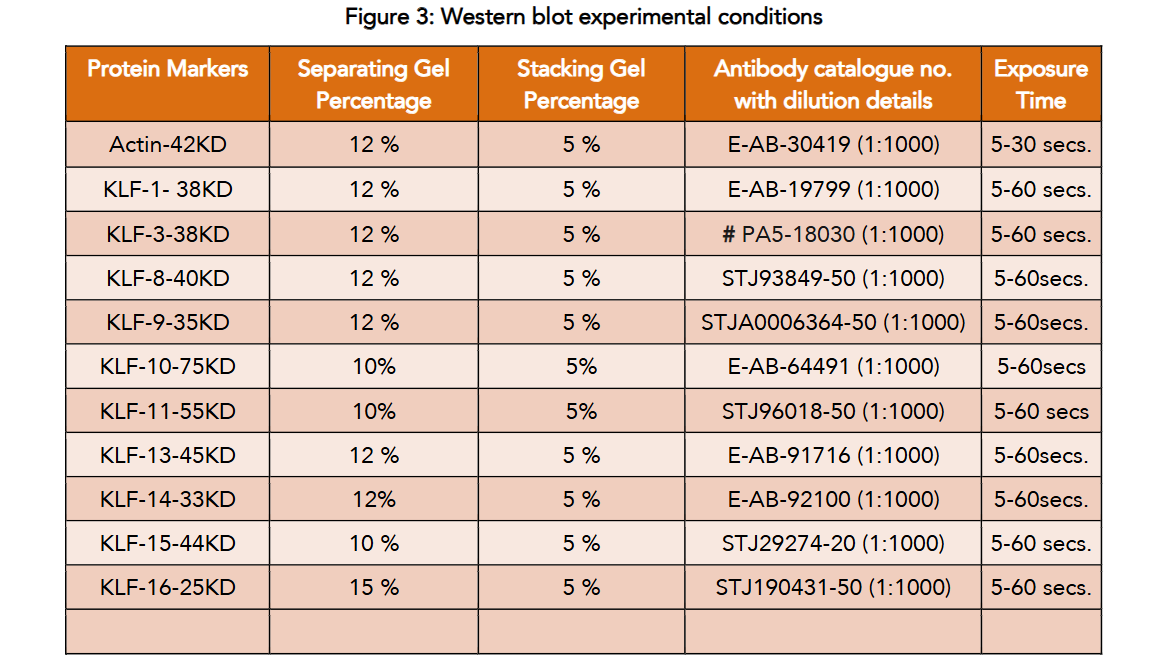
Table 3: Primer details
| Primer | Sequence | Amplicon size | Annealing temperature |
|---|---|---|---|
| GAPDH | GTCTCCTCTGACTTCAACAGCG | 186 | 60 |
| KLF1 | CAGGTGTGATAGCCGAGACC | 111 | 65 |
| KLF2 | CCAAGAGTTCGCATCTGAAGGC | 131 | 65 |
| KLF3 | CTCATGGTCTCCTTATCGGAGG | 131 | 65 |
| KLF4 | CCTTCCTGCCCGATCAGATG | 132 | 62 |
| KLF5 | GGAGAAACGACGCATCCACTAC | 140 | 65 |
| KLF6 | GACAGCTCCGAGGAACTTTCT | 156 | 65 |
| KLF7 | CTCACGAGGCACTACAGGAAAC | 135 | 67 |
| KLF8 | CCTGAAAGCTCACCGCAGAATC | 113 | 61 |
| KLF9 | GGGAAACACGCCTCCGAAAA | 110 | 65 |
| KLF10 | AGGAGTCACATCTGTAGCCACC | 139 | 67 |
| KLF11 | ATGGATGCAGCCACACCTGAAC | 115 | 65 |
| KLF12 | CCTTTCCATAGCCAGAGCAGTAC | 130 | 65 |
| KLF13 | CAGAGGAAGCACAAGTGCCACT | 137 | 65 |
| KLF14 | CATCCAGATATGATCGAGTACCG | 163 | 65 |
| KLF15 | GTGAGAAGCCCTTCGCCTGCA | 114 | 67 |
| KLF16 | GACTGCGCCAAAGCCTACTACA | 171 | 65 |
| KLF17 | GCTGCCCAGGATAACGAGAAC | 128 | 67 |
| KLF18 | TCCATGGGCCAGAAAGTGAC | 197 | 67 |
Results
The data indicate that metadichol exerts a complex regulatory effect, with both upregulation and downregulation of KLF genes depending on the specific KLF member and the treatment concentration. Figures 5 and 6 show the Protein expression of select KLF family members. KLF2 and KLF 4 see reference. At the 1 ng/ml concentration, Metadichol downregulated the expression of 14 out of 18 KLF transcription factors (KLF1, KLF2, KLF3, KLF5, KLF6, KLF7, KLF8, KLF9, KLF10, KLF11, KLF12, KLF13, KLF14, and KLF16), with fold changes ranging from 0.12 (KLF7) to 0.89 (KLF15). In contrast, KLF4, KLF15, KLF17, and KLF18 were upregulated at this concentration, with fold changes of 1.56, 0.89, 0.66, and 1.75, respectively. This selective upregulation suggests that Metadichol may preferentially enhance the activity of specific KLFs that play tumor-suppressive or context-dependent roles in cancer biology while suppressing those associated with oncogenic functions.
At lower concentrations (1 pg/ml and 100 pg/ml), the effects were more variable. For instance, at 1 pg/ml, KLF8 and KLF9 exhibited significant upregulation (fold changes of 4.99 and 3.02, respectively), while KLF17 was markedly downregulated (fold change of 0.39). At 100 pg/ml, most KLFs showed modest changes, with KLF4 notably downregulated (fold change of 0.24). At the highest concentration tested (100 ng/ml), the expression levels of many KLFs returned closer to baseline, with KLF17 and KLF18 showing significant upregulation (fold changes of 2.47 and 1.91, respectively). This biphasic response suggests a dose-dependent mechanism of action, where low concentrations may activate certain KLFs, while higher concentrations predominantly suppress expression, potentially reflecting a saturation effect or feedback regulation within the signaling networks modulated by Metadichol.
qRT‒PCR revealed that at 1 ng/ml metadichol, all KLFs except KLF4, KLF15, KLF17, and KLF18 were downregulated. The experimental results and literature-curated results were well correlated. In esophageal cancer, KLF4 expression is decreased, and KLF5 expression is increased. Metadichol increased KLF4 expression and decreased KLF5 expression at 1 ng/ml. In bladder cancer, 100 ng/ml metadichol increased KLF4 expression and decreased KLF5 and KLF6 expression. In ovarian cancer, KLF8 expression is elevated, and KLF4 expression is decreased. These findings are important features for identifying strategies for combating ovarian cancer. We can compare our experimental results to those in the literature. The findings presented in Tables 4 and 5 indicate that targeting all Krüppel-like factor (KLF) family members in cancer therapy may have several benefits:
Table 4: Fold regulation of KLF genes (1-18)
| Control | 1 pg/ml | 100 pg/ml | 1 ng/ml | 100 ng/ml | |
|---|---|---|---|---|---|
| KLF1 | 1 | 1.42 | 0.94 | 0.22 | 0.7 |
| KLF2 | 1 | 1.03 | 0.67 | 0.27 | 0.77 |
| KLF3 | 1 | 1.63 | 1.35 | 0.38 | 1.35 |
| KLF4 | 1 | 1.1 | 0.24 | 1.56 | 0.8 |
| KLF5 | 1 | 0.98 | 0.7 | 0.3 | 0.96 |
| KLF6 | 1 | 0.57 | 0.54 | 0.46 | 0.47 |
| KLF7 | 1 | 0.87 | 0.5 | 0.12 | 1.02 |
| KLF8 | 1 | 4.99 | 0.85 | 0.34 | 1.02 |
| KLF9 | 1 | 3.02 | 1.3 | 0.41 | 1.08 |
| KLF10 | 1 | 1.4 | 1.38 | 0.26 | 1.15 |
| KLF11 | 1 | 1.45 | 1.5 | 0.28 | 0.91 |
| KLF12 | 1 | 0.59 | 0.76 | 0.22 | 0.88 |
| KLF13 | 1 | 1.84 | 0.92 | 0.27 | 1.12 |
| KLF14 | 1 | 1.45 | 1.05 | 0.32 | 1.51 |
| KLF15 | 1 | 2.9 | 1.49 | 0.89 | 2.14 |
| KLF16 | 1 | 2.31 | 1.47 | 0.36 | 0.93 |
| KLF17 | 1 | 0.39 | 0.41 | 0.66 | 2.47 |
| KLF18 | 1 | 1.23 | 0.87 | 1.75 | 1.91 |
The Western blot results confirm that Metadichol modulates KLF protein expression in PBMCs in a dose-dependent manner, with upregulation at low concentrations (1 pg/ml), downregulation at 1 ng/ml, and stabilization at 100 ng/ml. The data align well with qRT-PCR findings for most KLFs, supporting Metadichol’s potential to target oncogenic KLFs (e.g., KLF-8, KLF-3) while preserving or enhancing tumor-suppressive KLFs (e.g., KLF-15, KLF-9). Discrepancies for some KLFs (e.g., KLF-9, KLF-14).
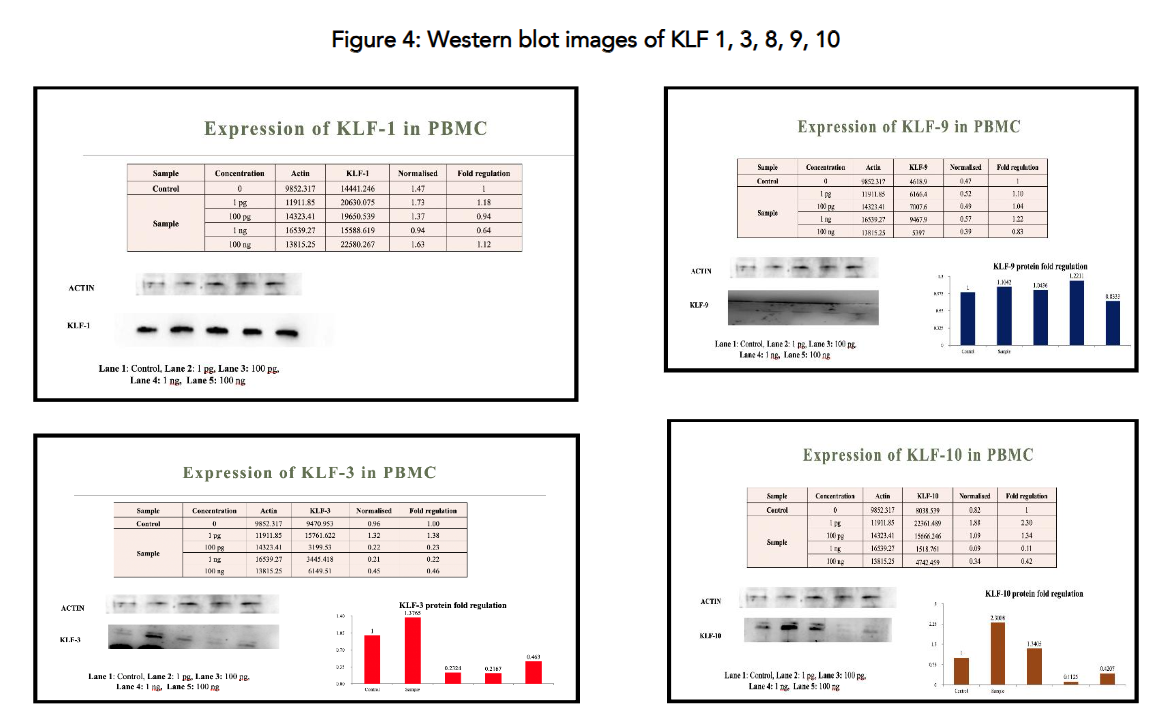
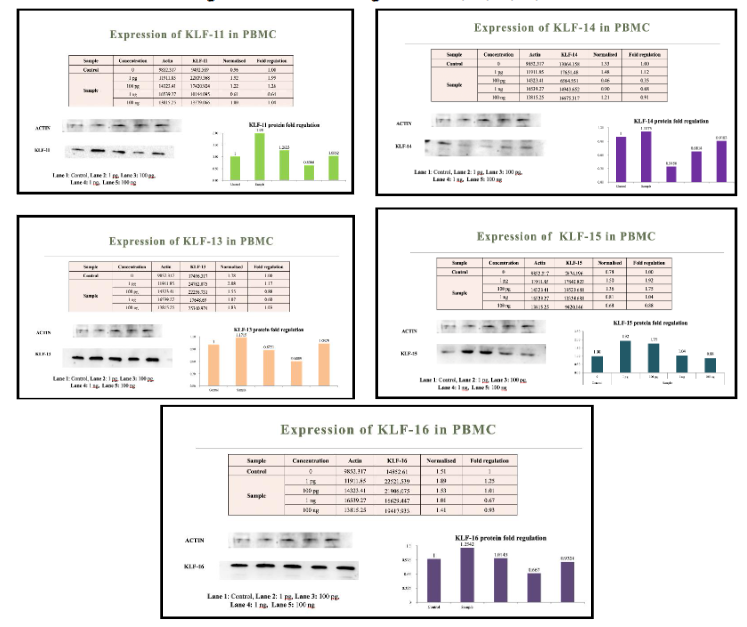
Owing to their different cancer progression functions, several KLFs can have synergistic antitumor effects. Different KLFs influence cancer growth, apoptosis, metastasis, and stemness. Targeting one KLF may result in compensation by another family member because KLF functions commonly overlap. Inhibiting numerous KLFs simultaneously can overcome compensatory processes and improve treatment efficacy.
Table 5: KLFs and their expression levels in cancer malignancies
| Type of cancer | Decreased expression | Increased expression |
|---|---|---|
| Bladder cancer | KLF4 | KLF5, KLF16 |
| Breast cancer | KLF2, KLF4, KLF6, KLF 8-11, KLF14, KLF15, KLF17 | KLF5, KLF16 |
| Cervical cancer | KLF4, KLF14 | KLF1, KLF4 |
| Clear cell carcinoma | KLF6 | |
| Colorectal cancer | KLF2-6, KLF8-10, KLF12-15, KLF17 | KLF1, KLF4, KLF5, KLF7, KLF16 |
| Cutaneous squamous cell carcinoma | KLF4 | |
| Endometrial cancer | KLF9 | |
| Esophagus cancer | KLF4 | KLF5 |
| Gastric cancer | KLF4, KLF6, KLF17 | KLF1, KLF5, KLF8, KLF16 |
| Glioma | KLF6, KLF9 | |
| Head and neck cancer | KLF6 | KLF5 |
| Hemangioma | KLF7 | |
| Hepatocellular carcinoma | KLF2, KLF4, KLF6, KLF17 | KLF4, KLF5, KLF8 |
| Leukemia | KLF3, KLF5 | |
| Lung cancer | KLF4-6, KLF17 | KLF7, KLF16 |
| Lymphoma | KLF4 | |
| Melanoma | KLF10 | KLF5 |
| Myeloma | KLF9, KLF10 | |
| Oral squamous cell carcinoma | KLF6 | KLF16 |
| Osteosarcoma | KLF3, KLF5, KLF8 | |
| Ovarian cancer | KLF4 | KLF8 |
| Pancreatic cancer | KLF9, KLF10 | KLF5 |
| Prostate cancer | KLF3-13, KLF17 | KLF1, KLF15, KLF16 |
| Retinoblastoma | KLF2 | KLF16 |
| Thyroid cancer | KLF17 | KLF5 |
Discussion
It has been observed that metadichol can affect a variety of signaling pathways, including those that involve NRs, for example, peroxisome proliferator-activated receptors (PPARs), sirtuins (SIRT1, SIRT6), TLRs, and circadian genes (for example, CLOCK, CRY1, and PPARGC1A). The expression of genes, including those belonging to the KLF family, are regulated by the convergence of several pathways. In this section, we will investigate the ways in which each route contributes to the activation and expression of KLF, which is supported by evidence that has been mentioned.
The activation of the KLF is mediated by nuclear receptors
- Regulation of gene expression is accomplished through the binding of certain DNA sequences by nuclear receptors, which are ligand-activated transcription factors. Metadichol and its ability to affect PPARs, specifically PPARα and PPARδ, play a crucial role in its impact on lipid metabolism and inflammation, as suggested by previous research. In order to facilitate the activation of NRs, KLFs interact with them in a number of different ways.
- Coregulation with PPARs: There is a physical interaction between KLF15 and PPARα, which is responsible for regulating genes involved in lipid metabolism, including those implicated in Gluconeogenesis. The act of silencing KLF15 has the effect of reducing the expression of PPARα target genes, which suggests that the activation of PPARα by metadichol may lead to an increase in KLF15 activity. In a similar manner, KLF5 assists in the regulation of lipid oxidation by working in conjunction with PPARδ, and its activity is influenced by SUMOylation. The regulation of PPARδ by Metadichol has the potential to activate KLF5 and increase its expression in peripheral blood mononuclear cells (PBMCs).
- Direct Nuclear Receptor Targets: Krüppel-like factor 2, KLF9, and KLF15 are direct targets of glucocorticoid receptors (GRs) and mineralocorticoid receptors (MRs). During times of stress or circadian variation, there is an increase in the amount of binding that occurs to glucocorticoid response elements (GREs) in the promoters of these genes. Metadichol NR activation may result in an increase in the binding efficiency of GR or MR, which in turn leads to an upregulation of these KLFs in PBMCs.
- Feedback loops: Krüppel-like factor 14 improves insulin sensitivity by increasing the expression of SIRT1 and PPARγ coactivator-1α (PGC-1α), two proteins that are associated with NR signaling. Through the use of metadichol NR modulation, the expression of KLF14 might be amplified, so generating a feedback loop that drives other KLFs.
Regulation of the KLF by Sirtuin’s Mediation
- Nicotinamide adenine dinucleotide-dependent deacetylases, known as sirtuins, are responsible for regulating metabolic processes, inflammation, and epigenetic changes. It is believed that the elevation of SIRT1 and SIRT6 by metadichol has an effect on the activity of KLF.
- The synergy between Krüppel-like factor 14 and SIRT1: KLF14 overexpression in hepatocytes leads to an increase in SIRT1 mRNA levels, which in turn leads to an increase in insulin sensitivity. There is a possibility that the activation of SIRT1 by metadichol could stabilize KLF14, hence boosting its expression as well as the expression of related KLFs through transcriptional networks.
- Krüppel-like factor 10 and SIRT6 in Inflammation: KLF10 promotes SIRT6 to decrease NF-κB signaling in pancreatic cancer, hence assisting in the reduction of epithelial-mesenchymal transition. Metadichol’s regulation of SIRT6 in PBMCs may lead to an increase in KLF10 expression, which in turn contributes to the anti-inflammatory properties of the compound.
- Transformations of the Epigenome: The accessibility of chromatin is altered as a result of sirtuins’ ability to deacetylate histones and transcription factors. When it comes to histone deacetylases (HDACs) and acetyltransferases (such as CBP/p300), KLF4 and KLF6 are involved in interactions. The sirtuin activity brought about by metadichol.
Krüppel-like factor expression That Is Mediated By TLR
- Innate immune responses are mediated by pattern-recognition receptors known as TLRs. The regulation of TLRs in peripheral blood mononuclear cells (PBMCs) by metadichol has the potential to promote the expression of KLF.
- Krüppel-like factor 4 and NF-κB Suppression: KLF4 inhibits the signaling pathway of NF-κB by interacting with HDAC1 and p50/p65, hence decreasing the expression of proinflammatory genes including MMP9. An anti-inflammatory phenotype can be promoted in peripheral blood mononuclear cells (PBMCs) by the anti-inflammatory actions of metadichol, which are mediated by TLR regulation.
- The role of KLF2 in immunological regulation is demonstrated by the fact that it controls the responses of macrophages to polymicrobial infections and endotoxic shock by means of TLR signaling. To reduce inflammation in peripheral blood mononuclear cells (PBMCs), the regulation of proinflammatory TLR pathways by metadichol could lead to a rise in KLF2 expression.
- Transforming growth factor beta 1 (TGFb1) and its receptors are activated by KLF6 in response to injury, which is associated to TLR-mediated inflammation. KLF6 is also involved in the process of tissue repair. There is a possibility that TLR regulation with metadichol could boost KLF6 expression, hence aiding immune cell repair tasks.
The regulation of Krüppel-like factor by circadian rhythm
- Metadichol has been shown to drastically alter the expression of circadian genes in human fibroblasts, including CLOCK, CRY1, and PPARGC1A, which suggests that it may play a role in the expression of rhythmic genes. There is a strong connection between KLFs and circadian regulation.
- Krüppel-like factor 9 and KLF13 as Clock Outputs: The CLOCK-BMAL1 complex, which binds E-boxes in the promoters of KLF9 and KLF13, is responsible for regulating the expression of these two proteins. It is possible that the overexpression of CLOCK by metadichol could lead to an increase in the expression of KLF9 and KLF13, which could allow for the modulation of immunological rhythms in PBMCs.
- Krüppel-like factor 15 and metabolic rhythms: KLF15 is responsible for regulating the circadian expression of metabolic genes in the heart and liver. It does this by interacting with PPARα and oscillating with the sleep-wake cycle. The activation of PPARGC1A by metadichol may result in an increase in the expression of KLF15, which in turn affects the equilibrium of lipids and nitrogen in PBMCs.
- Krüppel-like factor 2, KLF9, and KLF15 exhibit enhanced GR/MR binding to GREs during circadian variation and Stress. It is a relationship between stress and the integration of circadian rhythms. It is possible that the regulation of CRY1 by metadichol could exacerbate these effects, thereby coordinating the expression of KLF with immunological and metabolic cycles.
Role of Posttranslational Modifications with Regard to Molecular Switches
- The effects of metadichol on NRs, sirtuins, TLRs, and circadian genes are expected to focus on posttranslational modifications (PTMs) that influence the activity of KLF.
- Acetylation: Krüppel-like factor 2, KLF4, and KLF6 are all acetylated by CBP/p300, which has the effect of enhancing their binding to coactivators like EP300. Metadichol acts as a promoter of Sirtuin-mediated deacetylation, which has the potential to stabilize the contacts of KLF with repressors like Sin3A, thereby fine-tuning the activity of these repressors.
- Enhanced transcriptional activity or nuclear localization can be achieved through the process of phosphorylation, which involves the phosphorylation of KLF5, KLF11, and KLF13 by kinases such as ERK/MAPK or PRP4. The ability of metadichol to modify TLR and NR signals, which are responsible for activating kinases, can phosphorylate KLFs, which in turn increases the activity of these kinases.
- The process of SUMOylation is responsible for regulating the repressor activity of KLF5 in the context of lipid metabolism. Metadichol and its regulation of PPARδ have the potential to change the SUMOylation of KLF5, hence causing it to transition into an activator state in peripheral blood mononuclear cells (PBMCs).
- The nanoemulsion of Metadichol increases its bioavailability, which in turn makes it easier for it to interact with membrane-bound receptors (such as TLRs) and nuclear factors (such as NRs and sirtuins). The upregulation of circadian genes suggests a rhythmic component that synchronizes KLF expression in PBMCs, which is critical for immune regulation.
Multiple Signaling Pathway Modulation
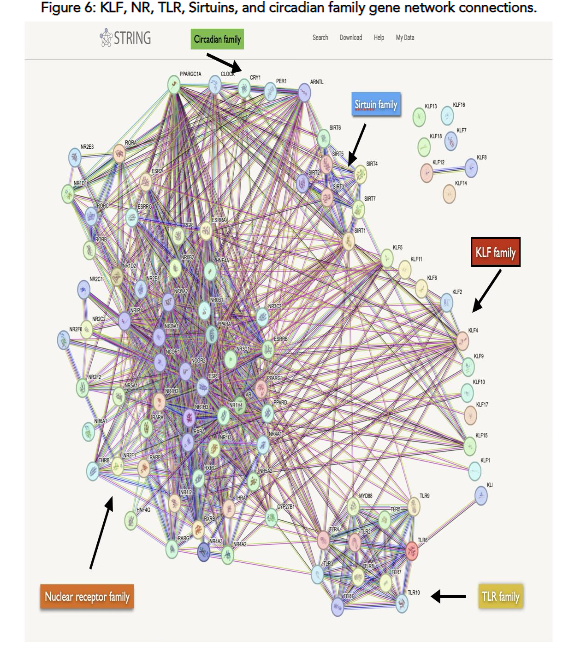
All the genes expressed by Metadichol were input into the STRING curated database and a network analysis was carried out. Network analysis program that shows interactions between and relationships between the gene families that are expressed by Metadichol see Figure 6. Each of the genes that are controlled by Metadichol can be synergistically active in a variety of disease processes, including cancer, as seen in Figure 7, which depicts the complex signaling network that regulates biological processes relevant to cancer where all the transcription factors expressed by Metadichol play synergistically important roles.
Concluding Remarks
Within peripheral blood mononuclear cells (PBMCs), metadichol has a role in facilitating the activation and expression of genes that belong to the KLF family. This is accomplished through a complicated system that includes nuclear receptors, sirtuins, TLRs, circadian genes, and posttranslational modifications. KLFs such as KLF2, KLF4, KLF6, KLF9, KLF10, KLF13, KLF14, and KLF15 are regulated by a transcriptional network that is produced by Metadichol. This network is regulated by PPARs, GRs, SIRT1, SIRT6, TLR signaling, and circadian genes such as CLOCK and CRY1. Due to the fact that they regulate immune responses, inflammation, and metabolism, KLFs have the potential to be utilized as therapeutic agents for the treatment of immunological and metabolic disorders.
The fact that metadichol is capable of exerting an influence on a wide variety of molecular families that are linked to the formation and progression of cancer presents a tremendously promising potential for the study and treatment of cancer and other diseases. In addition to being non-toxic, the fact that Metadichol contains nanoemulsion properties is another factor that contributes to the multitarget medicine potential that it possesses. When compared to treatments that concentrate on a specific pathway, Metadichol is preferable since it has the capacity to change a wide variety of cancer hallmarks.

Declarations
This work has been published as a preprint available at https://www.preprints.org/manuscript/202502.0271/v1.
Author Contributions:
PPR designed and supervised the work and is entirely responsible for its content.
Conflict of Interest Statement:
The author is a major shareholder in Nanorx, Inc., NY, USA, and the author founded the company.
Funding Statement:
Nanorx, Inc.’s R&D budget provided funding.
Supplementary Files
- File. PDF; Short synopsis of our results
- File: PDF; Raw Q-RT-PCR data
- File: PDF; Western blot of selected KLF genes.
Acknowledgements:
None.
References:
- Yuce K, Ozkan AI. The Krüppel-like factor (KLF) family, diseases, and physiological events. Gene. 2024;895:148027. doi:10.1016/j.gene.2023.148027
- Swamynathan SK. Krüppel-like factors: three fingers in control. Hum Genomics. 2010;4(4):263-270. doi:10.1186/1479-7364-4-4-263
- Black AR, Black JD, Azizkhan-Clifford J. Sp1 and Krüppel-like factor family of transcription factors in cell growth regulation and cancer. J Cell Physiol. 2001;188(2):143-160. doi:10.1002/jcp.1111
- Alder JK, Georgantas RW III, Hildreth RL, et al. Krüppel-like factor 4 is essential for inflammatory monocyte differentiation in vivo. J Immunol. 2008;180(8):5645-5652. doi:10.4049/jimmunol.180.8.5645
- Mao A, Zhou X, Liu Y, Ding J, Miao A, Pan G. KLF8 is associated with poor prognosis and regulates glycolysis by targeting GLUT4 in gastric cancer. J Cell Mol Med. 2019;23(8):5087-5097. doi:10.1111/jcmm.14378
- Sebzda E, Zou Z, Lee JS, Wang T, Kahn ML. Transcription factor KLF2 regulates the migration of naive T cells by restricting chemokine receptor expression patterns. Nat Immunol. 2008;9(3):292-300. doi:10.1038/ni1565
- Papadakis KA, Krempski J, Svingen P, et al. Krüppel-like factor KLF10 deficiency predisposes to colitis through colonic macrophage dysregulation. Am J Physiol Gastrointest Liver Physiol. 2015;309(11):G900-G909. doi:10.1152/ajpgi.00309.2015
- Mallipattu SK, Estrada CC, He JC. The critical role of Krüppel-like factors in kidney disease. Am J Physiol Renal Physiol. 2017;312(2):F259-F265. doi:10.1152/ajprenal.00550.2016
- Papadakis K, Krempski J. Krüppel-like factor KLF10 regulates intestinal macrophages and innate immune colitis. Inflamm Bowel Dis. 2014;20(suppl 1):S106. doi:10.1097/01.MIB.0000456963.59724.80
- Dabravolski SA, Sukhorukov VN, Kalmykov VA, Grechko AV, Shakhpazyan NK, Orekhov AN. The role of KLF2 in the regulation of atherosclerosis development and potential use of KLF2-targeted therapy. Biomedicines. 2022;10(2):254. doi:10.3390/biomedicines10020254
- Sathanoori R, Rosi F, Gu BJ, et al. Shear stress modulates endothelial KLF2 through activation of P2X4. Purinergic Signal. 2015;11(1):139-153. doi:10.1007/s11302-014-9442-3
- Liu C, Shen M, Tan WLW, et al. Statins improve endothelial function via suppression of epigenetic-driven EndMT. Nat Cardiovasc Res. 2023;2:467-485. doi:10.1038/s44161-023-00267-1
- Tugal D, Jain MK, Simon DI. Endothelial KLF4: crippling vascular injury? J Am Heart Assoc. 2014;3(1):e000769. doi:10.1161/JAHA.113.000769
- Heard ME, Velarde MC, Giudice LC, Simmen FA, Simmen RCM. Krüppel-like factor 13 deficiency in uterine endometrial cells contributes to defective steroid hormone receptor signaling but not lesion establishment in a mouse model of endometriosis. Biol Reprod. 2015;92(6):140. doi:10.1095/biolreprod.115.130260
- Heard ME, Simmons CD, Simmen FA, Simmen RC. Krüppel-like factor 9 deficiency in uterine endometrial cells promotes ectopic lesion establishment associated with activated notch and hedgehog signaling in a mouse model of endometriosis. Endocrinology. 2014;155(4):1532-1546. doi:10.1210/en.2013-1947
- Homeister AW, Patterson C. Zinc fingers in the pizza pie. Circ Res. 2008;103(7):665-667. doi:10.1161/CIRCRESAHA.108.185763
- Yang Y, Goldstein BG, Chao HH, Katz JP. KLF4 and KLF5 regulate proliferation, apoptosis, and invasion in esophageal cancer cells. Cancer Biol Ther. 2005;4(11):1216-1221. doi:10.4161/cbt.4.11.2090
- Brown AR, Simmen R, Simmen F. Suppression of insulin-induced fatty acid synthase gene expression and colon cancer cell proliferation by members of the Krüppel-like family of transcription factors. Cancer Prev Res. 2011;4(10 Suppl):A44.
- Bieker JJ. Krüppel-like factors: three fingers in many pies. J Biol Chem. 2001;276(37):34355-34358. doi:10.1074/jbc.R100043200
- Pang CJ, Lemsaddek W, Alhashem YN, et al. Krüppel-like factor 1 (KLF1), KLF2, and Myc control a regulatory network essential for embryonic erythropoiesis. Mol Cell Biol. 2012;32(13):2628-2644. doi:10.1128/MCB.00104-12
- Kalra IS, Alam MM, Choudhary PK, Pace BS. Krüppel-like factor 4 activates HBG gene expression in primary erythroid cells. Br J Haematol. 2011;154(2):248-259. doi:10.1111/j.1365-2141.2011.08710.x
- Vinci M, Greco D, Treccarichi S, et al. Bioinformatic evaluation of KLF13 genetic variant: implications for neurodevelopmental and psychiatric symptoms. Genes. 2024;15(8):1056. doi:10.3390/genes15081056
- Shaverdashvili K, Padlo J, Weinblatt D, et al. KLF4 activates NFκB signaling and esophageal epithelial inflammation via the Rho-related GTP-binding protein RHOF. PLoS One. 2019;14(4):e0215746. doi:10.1371/journal.pone.0215746
- Jiang Z, Yu T, Fan Z, Yang H, Lin X. Krüppel-like factor 7 is a marker of aggressive gastric cancer and poor prognosis. Cell Physiol Biochem. 2017;43(3):1090-1099. doi:10.1159/000481748
- Raghavan PR, inventor. US patent 8,722,093. May 13, 2014.
- Raghavan PR, inventor. US patent 9,006,292. April 14, 2015.
- Raghavan PR. Metadichol®-induced expression of sirtuins 1-7 in somatic and cancer cells. Med Res Arch. 2024;12(6). doi:10.18103/mra.v12i6.5371
- Raghavan PR. Metadichol-induced expression of toll receptor family members in peripheral blood mononuclear cells. Med Res Arch. 2024;12(8). doi:10.18103/mra.v12i8.5610
- Raghavan PR. Metadichol®: a nano lipid emulsion that expresses all 49 nuclear receptors in stem and somatic cells. Arch Clin Biomed Res. 2023;7:524-536. doi:10.26502/acbr.50170368
- Raghavan PR. Metadichol, a natural ligand for the expression of Yamanaka reprogramming factors in human cardiac, fibroblast, and cancer cell lines. Med Res Arch. 2024;12(6). doi:10.18103/mra.v12i6
- Raghavan PR. Metadichol®-induced expression of circadian clock transcription factors in human fibroblasts. Med Res Arch. 2024;12(6). doi:10.18103/mra.v12i6.5371
- Raghavan PR. Metadichol-induced KLF expression in PBMC cells links SIRTs, NRs, TLRs, and circadian genes: a systems-wide biology approach. Preprints. 2025:2025020271. doi:10.20944/preprints202502.0271.v1
- Tetreault MP, Wang ML, Yang Y, et al. Klf4 overexpression activates epithelial cytokines and inflammation-mediated esophageal squamous cell cancer in mice. Gastroenterology. 2010;139(6):2124-2134.e9. doi:10.1053/j.gastro.2010.08.048
- Tarapore RS, Yang Y, Katz JP. Restoring KLF5 in esophageal squamous cell cancer cells activates the JNK pathway leading to apoptosis and reduced cell survival. Neoplasia. 2013;15(5):472-480. doi:10.1593/neo.122126
- Lahiri SK, Zhao J. Krüppel-like factor 8 emerges as an important regulator of cancer. Am J Transl Res. 2012;4(3):357-363.
- McConnell BB, Yang VW. Mammalian Krüppel-like factors in health and diseases. Physiol Rev. 2010;90(4):1337-1381. doi:10.1152/physrev.00058.2009
- Yang Y, Katz JP. Krüppel-like factors in the biology of cancer. In: Nagai RF, Friedman SL, Kasuga M, eds. The Biology of Krüppel-like Factors. Springer; 2009:67-82.
- Tetreault MP, Yang Y, Katz J. Krüppel-like factors in cancer. Nat Rev Cancer. 2013;13:701-713. doi:10.1038/nrc3582
- Zhang Y, Yao C, Ju Z, et al. Krüppel-like factors in tumors: key regulators and therapeutic avenues. Front Oncol. 2023;13:1080720. doi:10.3389/fonc.2023.1080720
- Rascio F, Spadaccino F, Rocchetti MT, et al. The pathogenic role of PI3K/AKT pathway in cancer onset and drug resistance. Cancers (Basel). 2021;13(16):3949. doi:10.3390/cancers13163949
- Zhu B, Liu Q, Han Q, Zeng B, Chen J, Xiao Q. Downregulation of Krüppel-like factor 1 inhibits the metastasis and invasion of cervical cancer cells. Mol Med Rep. 2018;18(4):3932-3940. doi:10.3892/mmr.2018.9401
- Xu R, Chen Y, Wei S, Chen J. Comprehensive pan-cancer analysis of the prognostic role of KLF transcription factor 2 (KLF2) in human tumors. Onco Targets Ther. 2024;17:887-904. doi:10.2147/OTT.S476179
- Zhu J, Teng H, Zhu X, Yuan J, Zhang Q, Zou Y. Pancancer analysis of Krüppel-like factor 3 and its carcinogenesis in pancreatic cancer. Front Immunol. 2023;14:1167018. doi:10.3389/fimmu.2023.1167018
- He Z, He J, Xie K. KLF4 transcription factor in tumorigenesis. Cell Death Discov. 2023;9:118. doi:10.1038/s41420-023-01416-y
- Luo Y, Chen C. The roles and regulation of the KLF5 transcription factor in cancers. Cancer Sci. 2021;112(6):2097-2117. doi:10.1111/cas.14910
- DiFeo A, Martignetti JA, Narla G. The role of KLF6 and its splice variants in cancer therapy. Drug Resist Updat. 2009;12(1-2):1-7. doi:10.1016/j.drup.2008.11.001
- Li Z, Liu Q. The oncogenic role of KLF7 in colon adenocarcinoma and therapeutic perspectives. Int J Genomics. 2023;2023:5520926. doi:10.1155/2023/5520926
“`




