Urinary Biomarkers for Predicting CKD in Diabetes & Hypertension
Cellular Biomarker in the Urine in Predicting Prognosis of Chronic Kidney Disease, Diabetes and Hypertension
Chie Yokoyama 1,2, Kenichi Shukuya 3, Tatsuo Shimosawa 2
- Chie Yokoyama Department of Clinical Laboratory University of Tsukuba Hospital, Ibaragi, Japan; Department of Clinical Laboratory, Faculty of Medicine, International University of Health and Welfare, Chiba, Japan
- Kenichi Shukuya Department of Clinical Medical Technology, Faculty of Medical Science, Juntendo University, Chiba. Japan
- Tatsuo Shimosawa Department of Clinical Laboratory, Faculty of Medicine, International University of Health and Welfare, Chiba, Japan
OPEN ACCESSS
PUBLISHED: 28 Febuary 2025
CITATION: YOKOYAMA, Chie; SHUKUYA, Kenichi; SHIMOSAWA, Tatsuo. Cellular Biomarker in the Urine in Predicting Prognosis of Chronic Kidney Disease, Diabetes and Hypertension. Medical Research Archives Available at: <https://esmed.org/MRA/mra/article/view/6096>.
COPYRIGHT: © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI: https://doi.org/10.18103/mra.v13i2.6096
ISSN 2375-1924
Abstract
Chronic kidney disease (CKD) is a life-threatening condition originated from renal dysfunction. So far we do not have perfect remedies and therefore prevention of progression draw high interests from both researchers and clinicians. There are a lot of risk factors for its development and its prognosis, among them, hypertension and diabetes are the most common risks. We have been using biomarkers for diabetes and hypertension such as HbA1c, diurnal changes of blood pressure, morphological changes in blood vessels and cardiac functions. Recent advances in chemical analysis proposed several biomarkers from blood and urine samples, such as microalbuminuria, L-FABP and others. Animal and cell experiments also suggest potential biomarkers, however, they are not necessarily applicable in human. Besides those chemical markers, studies on genetic or epigenetic factors such as microRNA have been reported. In addition to those upcoming techniques, we learned a lot from classical cellular morphological examination of urinary sediments. In the current review, we focus on podocyte and round cell in the urinary sediment and epigenetic markers in the urine as novel biomarkers for CKD and its risk factors those are hypertension and diabetes.
Keywords
Chronic Kidney Disease, Diabetes, Hypertension, Biomarkers, Podocytes, Urinary Sediment
Introduction
Concept of chronic kidney disease (CKD) is widely accepted, since it is a risk factor for end-stage renal disease (ESRD) and cardiovascular disease. Diabetes mellitus and hypertension are the leading cause of CKD. In Japan, approximately 13% of the Japanese adult population is estimated to have CKD. In the current situation there is no curative treatment for CKD, there is an urgent need to prevent its onset and progression. However, renal lesion assessment for diagnosis and prognosis currently can only be evaluated by renal biopsy. Renal biopsy is an invasive test that requires hospitalization and is difficult to repeat. Since patients with renal disease have few subjective symptoms, they tend to hesitate when it comes to renal biopsy, which leads to interruption of hospital visits without scrutiny. Urinalysis has very long history back to 2000B.C. in India. Urine, as an easily accessible biological fluid, offers a unique opportunity for monitoring disease progression and prognosis. Advances in technologies for analyzing urinary protein and various substances in urine have revealed the existence of substances related to the diagnosis and prognosis of kidney diseases. In acute renal injury (AKI), the expression of molecules such as Kim-1 and NGAL have been confirmed in the renal proximal tubular epithelial cell, and the importance of L-FABP as a biomarker for the progression of renal injury are also attracting attention. However, most of them reflect renal tubular cells injuries, targeting the glomerular injury, nothing exceeds the conventional total protein or albumin excretion, but these are not specific to glomerular injury as they are also resorbed at tubules. Therefore, it is necessary to establish a urinary biomarker that can diagnose glomerular disease and estimate renal prognosis. In addition to these chemicals, cellular components in the urinary sediment has been studied for more than 100 years. Red blood cells, white blood cells and casts were reported from the beginning and recently we and other researchers reported that podocyte, inflammatory cells such as neutrophils and macrophages, round cells in the urine can be biomarkers for glomerular and tubular damages. And by applying molecular biology, investigation of urinary exosomes, analysis of cellular epigenome markers, it become possible to determine epithelial cell origin in the kidney as well as novel biomarkers such as microRNA, cell-derived proteins. and those are reported to be related to CKD prognosis. By multimodal approaches in help of machine learning, it is expected that combining urinary cellular markers with clinical parameters (e.g., blood pressure, glycemic control) improves prognostic accuracy. And algorithms help integrate large biomarker datasets for personalized risk stratification. In the current review, we focus on recent advances in cell components in the urine that is expected to be good markers for CKD, especially in diabetic and/or hypertensive patients.
Podocyte Injury and Biomarkers
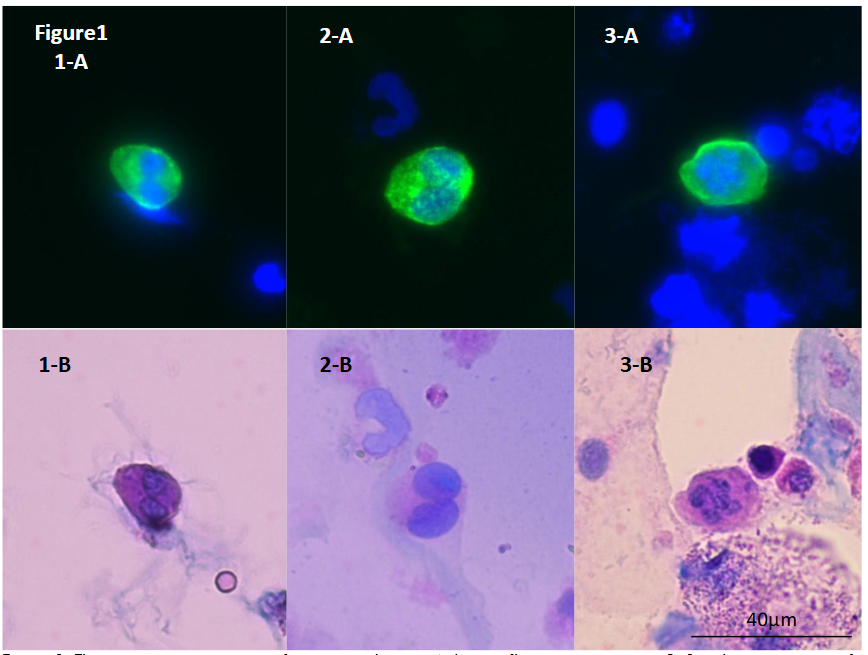
It has been indicated that podocyte injury leads to clinical development of proteinuria and the pathological progression of glomerulosclerosis. Research indicates that podocyte detachment, leading to their presence in urine, is associated with glomerular damage in CKD. Elevated urinary mRNA levels of podocyte-associated genes, such as NPHS2 (podocin) and PODXL (podocalyxin), have been observed in patients with progressive diabetic nephropathy, suggesting their potential as biomarkers for disease progression. However, these markers are mRNA level or podocyte-derived microparticles and difficult to use them in routine clinical work. On the other hand, the detection of podocytes in urinary sediment by an immunofluorescence staining using an antibody to podocalyxin and the number of podocyte relates with prognosis of CKD. In comparison between immunostaining and Sternheimer staining, we characterized podocyte morphology. The size varies from 10 to 40 μm and round-shaped. The bi- or multi-nuclei cells are observed and nucleus localize unevenly. When cells are smaller, the cytoplasm is thicker and vice versa. The surface of cytoplasm is smooth and fine structure. The margin of cytoplasm is clear but in some part is obscure. Cytoplasm is stained deep-purple to light pink and when it is thick, stained in purple. It was considered that the greatest feature of podocytes was to show a very smooth and fine cytoplasmic surface structure regardless of cell size / thickness / shape, nucleus size / shape / number, and stainability.
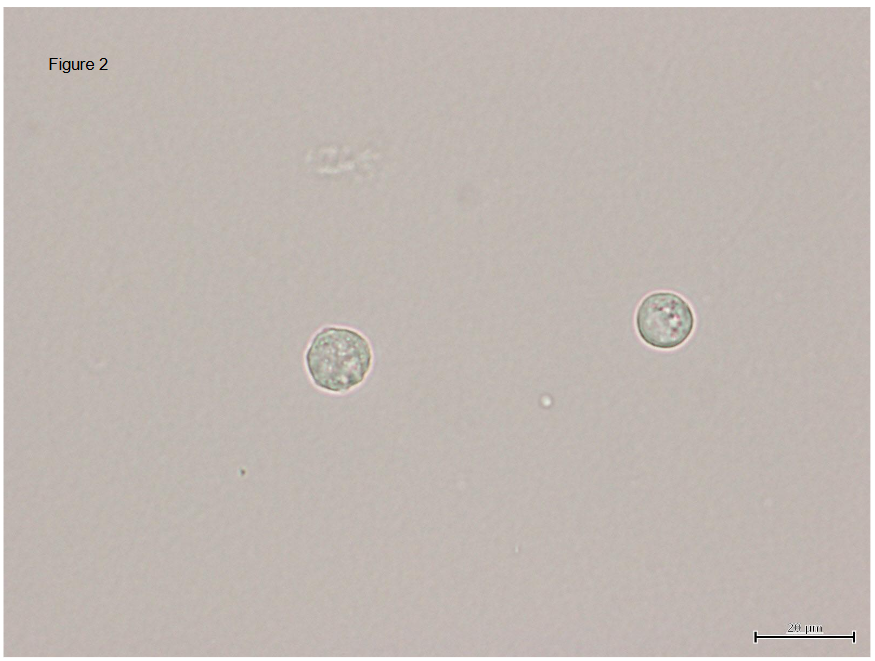
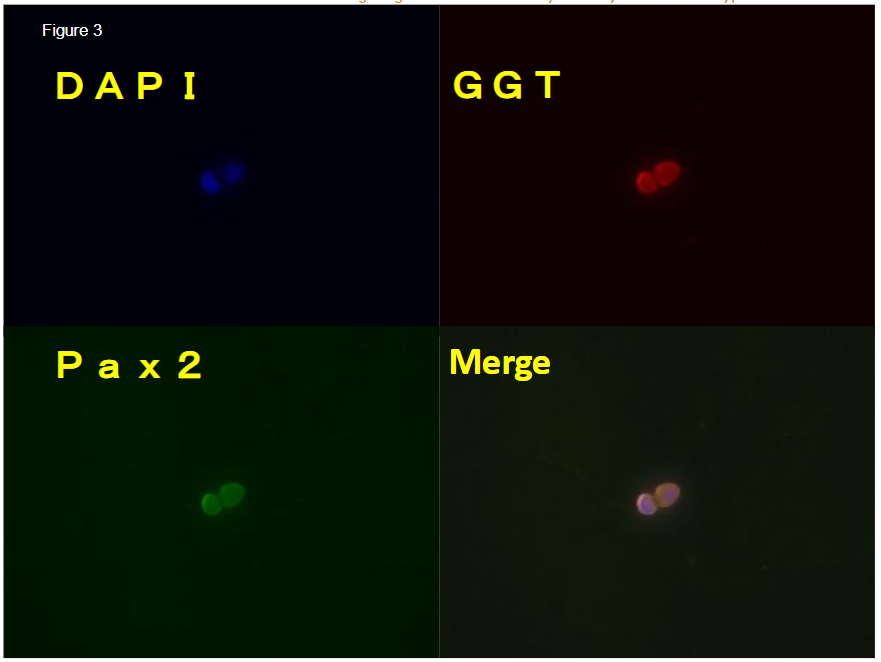
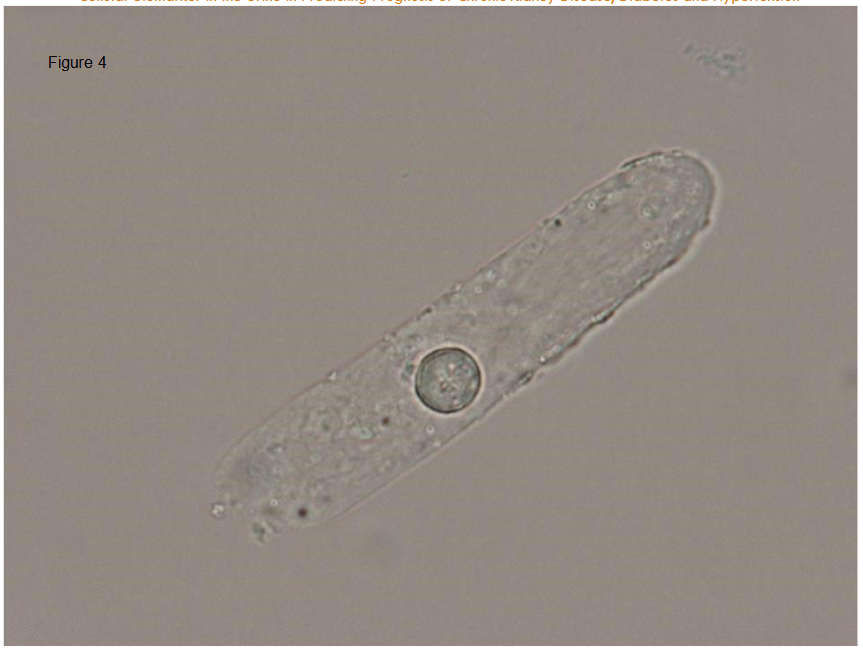
Round cell in the urine sediment was first reported by us. It is characterized as a cell about 10μm in diameter and nuclear/cytoplasm ration is large. Its molecular markers are PAX2, HOX11, OSR1, WT1 and SIX2 which suggests the round cell is undifferentiated. and negative for podocalyxin. and when it is cultured, it differentiates into epithelial cell and positive for GGT-1. Round cell number correlate with the prognosis of renal function and predict risks for hemodialysis in CKD patients. Prospective study shows in severe CKD patients, area under the curve of receiver-operated characteristic (ROC) curve in round cell number is better than urinary protein. The round cell is also found in hyaline casts, that suggests the cell are originated from tubules but not from urinary tract. In contrast to podocyte, which can not be divided and regenerate in the glomerulus and podocyturia suggests podocyte injury, renal tubules are regenerated rapidly and actively and in urine sediment we observe a lot of differentiated tubular epithelial cells. In order to regenerate tubules, there must be progenitor cells. We assume that this round cell is a progenitor cells and healthy kidney can recruit the round cell to the injured tubular lesion and it can differentiate into epithelial cells but under disease condition, the cell fail to home to the lesion and drop into the urine. Therefore, the number of round cell reflects tubular condition and/or progenitor cell condition and can be a biomarker to predict renal dysfunction. Recent preliminary study shows that in hypertensive nephropathy, round cell number increases and it is independent from proteinuria and can be used as tubular damage marker.
| The cellular findings in Sternheimer stain | Shape | Nucleus | Size | Thickness of cytoplasm | Structure of cytoplasmic surface | Structure of cytoplasmic edges | Stainability |
|---|---|---|---|---|---|---|---|
| Various shapes, round-shape | The bi- or multi- nuclei cells are observed and nucleus localize unevenly. | 10μm~40μm | Regarding the thickness of the cytoplasm, the identified cells tended to be thicker as the cell size was smaller and thinner as the cell was increased, but this could not be said unconditionally | A very smooth and fine cytoplasmic surface structure | Clarity | A red purple of a light pink. |
Conclusion
CKD affects millions of people worldwide, making it a widespread health concern. It often goes undiagnosed in its early stages because symptoms can be subtle or absent. With early detection through screening, CKD can often be managed or even prevented from progressing to more severe stages. Lifestyle changes, medications, and regular monitoring can help maintain kidney function and reduce complications. Current biomarkers are not high enough in both sensitivity and specificity. By multimodal approaches, including chemical, genetical, and epigenetic markers as well as cell population in the urine sediment will provide us new set of biomarkers. By using the biomarker set, we can detect early phase of CKD and monitor the progression of disease status. Understanding CKD’s importance emphasizes the need for awareness, regular check-ups, and proactive measures to reduce the risk of kidney damage.
References
- Imai E, Horio M, Watanabe T, et al. Prevalence of chronic kidney disease in the Japanese general population. Clin Exp Nephrol. Dec 2009;13(6):621-30. Doi:10.1007/s10157-009-0199-x
- Bonventre JV, Vaidya VS, Schmouder R, Feig P, Dieterle F. Next-generation biomarkers for detecting kidney toxicity. Nat Biotechnol. May 2010;28(5):436-40. Doi:10.1038/nbt0510-436
- Inoue K. Urinary Podocyte Biomarkers and Glomerular Histologic Change. Kidney360. Mar 31 2022;3(3):407-409. Doi:10.34067/KID.0008212021
- Ding F, Gao Q, Tian X, Mo J, Zheng J. Increasing urinary podocyte mRNA excretion and progressive podocyte loss in kidney contribute to the high risk of long-term renal disease caused by preterm birth. Sci Rep. Oct 19 2021;11(1):20650. Doi:10.1038/s41598-021-00130-y
- Fukuda A, Sato Y, Shibata H, Fujimoto S, Wiggins RC. Urinary podocyte markers of disease activity, therapeutic efficacy, and long-term outcomes in acute and chronic kidney diseases. Clin Exp Nephrol. Jun 2024;28(6):496-504. Doi:10.1007/s10157-024-02465-y
- Lu J, Hu ZB, Chen PP, et al. Urinary levels of podocyte-derived microparticles are associated with the progression of chronic kidney disease. Ann Transl Med. Sep 2019;7(18):445. Doi:10.21037/atm.2019.08.78
- Hara M, Yamamoto T, Yanagihara T, et al. Urinary excretion of podocalyxin indicates glomerular epithelial cell injuries in glomerulonephritis. Nephron. 1995;69(4):397-403. Doi:10.1159/000188509
- Vogelmann SU, Nelson WJ, Myers BD, Lemley KV. Urinary excretion of viable podocytes in health and renal disease. Am J Physiol Renal Physiol. Jul 2003;285(1):F40-8. Doi:10.1152/ajprenal.00404.2002
- Kriz W, Lemley KV. A potential role for mechanical forces in the detachment of podocytes and the progression of CKD. J Am Soc Nephrol. Feb 2015;26(2):258-69. Doi:10.1681/ASN.2014030278
- Yokoyama C, Usui J, Kobayashi M, et al. Sustained Appearance of Urinary Podocytes Suggests Poor Renal Prognosis in Kidney Transplant Patients with Focal Segmental Glomerulosclerosis: Case Reports and Review of Literature. Clin Lab. 2015;61(12):1961-6. Doi:10.7754/clin.lab.2015.150515
- Sternheimer R. A supravital cytodiagnostic stain for urinary sediments. JAMA. Feb 24 1975;231(8):826-32.
- Shukuya K, Ogura S, Tokuhara Y, et al. Novel round cells in urine sediment and their clinical implications. Clin Chim Acta. Jun 01 2016;457:142-9. Doi:10.1016/j.cca.2016.04.017
- Thongboonkerd V, Kanlaya R. The divergent roles of exosomes in kidney diseases: Pathogenesis, diagnostics, prognostics and therapeutics. Int J Biochem Cell Biol. Aug 2022;149:106262. Doi:10.1016/j.biocel.2022.106262
- Sinha N, Kumar V, Puri V, et al. Urinary exosomes: Potential biomarkers for diabetic nephropathy. Nephrology (Carlton). Dec 2020;25(12):881-887. Doi:10.1111/nep.13720
- Gámez-Valero A, Lozano-Ramos SI, Bancu I, Lauzurica-Valdemoros R, Borràs FE. Urinary extracellular vesicles as source of biomarkers in kidney diseases. Front Immunol. 2015;6:6. Doi:10.3389/fimmu.2015.00006
- Oba S, Ayuzawa N, Nishimoto M, et al. Aberrant DNA methylation of Tgfb1 in diabetic kidney mesangial cells. Sci Rep. Nov 05 2018;8(1):16338. Doi:10.1038/s41598-018-34612-3
- Li S, Li W, Wu R, et al. Epigenome and transcriptome study of moringa isothiocyanate in mouse kidney mesangial cells induced by high glucose, a potential model for diabetic-induced nephropathy. AAPS J. Dec 05 2019;22(1):8. Doi:10.1208/s12248-019-0393-z
- Sun G, Reddy MA, Yuan H, Lanting L, Kato M, Natarajan R. Epigenetic histone methylation modulates fibrotic gene expression. J Am Soc Nephrol. Dec 2010;21(12):2069-80. Doi:10.1681/ASN.2010060633
- Bell JT, Tsai PC, Yang TP, et al. Epigenome-wide scans identify differentially methylated regions for age and age-related phenotypes in a healthy ageing population. PLoS Genet. 2012;8(4):e1002629. Doi:10.1371/journal.pgen.1002629
- Chen Z, Satake E, Pezzolesi MG, et al. Integrated analysis of blood DNA methylation, genetic variants, circulating proteins, microRNAs, and kidney failure in type 1 diabetes. Sci Transl Med. May 22 2024;16(748):eadj3385. Doi:10.1126/scitranslmed.adj3385
- Chu AY, Tin A, Schlosser P, et al. Epigenome-wide association studies identify DNA methylation associated with kidney function. Nat Commun. Nov 03 2017;8(1):1286. Doi:10.1038/s41467-017-01297-7
- Kim JE, Jo MJ, Cho E, et al. The Effect of DNA Methylation in the Development and Progression of Chronic Kidney Disease in the General Population: An Epigenome-Wide Association Study Using the Korean Genome and Epidemiology Study Database. Genes (Basel). Jul 21 2023;14(7). Doi:10.3390/genes14071489
- Li KY, Tam CHT, Liu H, et al. DNA methylation markers for kidney function and progression of diabetic kidney disease. Nat Commun. May 15 2023;14(1):2543. Doi:10.1038/s41467-023-37837-7
- Schlosser P, Tin A, Matias-Garcia PR, et al. Meta-analyses identify DNA methylation associated with kidney function and damage. Nat Commun. Dec 09 2021;12(1):7174. Doi:10.1038/s41467-021-27234-3
- Smyth LJ, Dahlström EH, Syreeni A, et al. Epigenome-wide meta-analysis identifies DNA methylation biomarkers associated with diabetic kidney disease. Nat Commun. Dec 22 2022;13(1):7891. Doi:10.1038/s41467-022-34963-6
- Hishikawa A, Hayashi K, Yoshimoto N, Nakamichi R, Homma K, Itoh H. DNA damage and expression of DNA methylation modulators in urine-derived cells of patients with hypertension and diabetes. Sci Rep. Feb 25 2020;10(1):3377. Doi:10.1038/s41598-020-60420-9
- Marumo T, Hoshino J, Kawarazaki W, et al. Methylation pattern of urinary DNA as a marker of kidney function decline in diabetes. BMJ Open Diabetes Res Care. Sep 2020;8(1). Doi:10.1136/bmjdrc-2020-001501
- Marumo T, Yoshida N, Inoue N, et al. Aberrant proximal tubule DNA methylation underlies phenotypic changes related to kidney dysfunction in patients with diabetes. Am J Physiol Renal Physiol. Sep 01 2024;327(3):F397-F411. Doi:10.1152/ajprenal.00124.2024
- Elijovich F, Weinberger MH, Anderson CA, et al. Salt Sensitivity of Blood Pressure: A Scientific Statement From the American Heart Association. Hypertension. Sep 2016;68(3):e7-e46. Doi:10.1161/HYP.0000000000000047
- Morimoto A, Uzu T, Fujii T, et al. Sodium sensitivity and cardiovascular events in patients with essential hypertension. Lancet. Dec 13 1997;350(9093):1734-7. Doi:10.1016/S0140-6736(97)05189-1
- Gu D, Zhao Q, Chen J, et al. Reproducibility of blood pressure responses to dietary sodium and potassium interventions: the GenSalt study. Hypertension. Sep 2013;62(3):499-505. Doi:10.1161/HYPERTENSIONAHA.113.01034
- Mu S, Shimosawa T, Ogura S, et al. Epigenetic modulation of the renal beta-adrenergic-WNK4 pathway in salt-sensitive hypertension. Nature Medicine. MAY 2011 2011;17(5):573-U92. Doi:10.1038/nm.2337
- Cao N, Lan C, Chen C, et al. Prenatal Lipopolysaccharides Exposure Induces Transgenerational Inheritance of Hypertension. Circulation. Oct 04 2022;146(14):1082-1095. Doi:10.1161/CIRCULATIONAHA.122.059891
- Kawakami-Mori F, Nishimoto M, Reheman L, et al. Aberrant DNA methylation of hypothalamic angiotensin receptor in prenatal programmed hypertension. JCI Insight. Nov 02 2018;3(21). Doi:10.1172/jci.insight.95625
- Gildea JJ, Lahiff DT, Van Sciver RE, et al. A linear relationship between the ex-vivo sodium mediated expression of two sodium regulatory pathways as a surrogate marker of salt sensitivity of blood pressure in exfoliated human renal proximal tubule cells: the virtual renal biopsy. Clin Chim Acta. Jun 05 2013;421:236-42. Doi:10.1016/j.cca.2013.02.021
- Isobe K, Mori T, Asano T, et al. Development of enzyme-linked immunosorbent assays for urinary thiazide-sensitive Na-Cl cotransporter measurement. Am J Physiol Renal Physiol. Nov 01 2013;305(9):F1374-81. Doi:10.1152/ajprenal.00208.2013
- Xu P, Sudarikova AV, Ilatovskaya DV, et al. Epithelial Sodium Channel Alpha Subunit (αENaC) Is Associated with Inverse Salt Sensitivity of Blood Pressure. Biomedicines. Apr 23 2022;10(5). Doi:10.3390/biomedicines10050981
- Gildea JJ, Carlson JM, Schoeffel CD, Carey RM, Felder RA. Urinary exosome miRNome analysis and its applications to salt sensitivity of blood pressure. Clin Biochem. Aug 2013;46(12):1131-1134. Doi:10.1016/j.clinbiochem.2013.05.052

