Breast Cancer Decision Support: Expert vs. Machine Learning
Breast Cancer Decision Support: Expert Systems vs. Machine Learning – have we thrown the baby out with the bathwater?
Mustafa Khanbhai¹, Vivek Patkar³, Danny Ruta², Ashutosh Kothari², Hartmut Kristeleit², Majid Kazmi²*
- AI Centre for Value Based Healthcare, Becket House, 1 Lambeth Place Road, London SE1 7EU
- Guy’s & St Thomas’ NHS Foundation Trust, Guy’s Cancer Center, London SE1 9RT
- Deontics Ltd, Orion House 5 Upper St Martin’s Lane, London, WC2H 9EA, United Kingdom
[email protected]
OPEN ACCESS
PUBLISHED:31 December 2024
CITATION: Khanbhai, M., Patkar, V., et al., 2024. Breast Cancer Decision Support: Expert Systems vs. Machine Learning – have we thrown the baby out with the bathwater? Medical Research Archives, [online] 12(12). https://doi.org/10.18103/mra.v12i12.5998
COPYRIGHT: © 2024 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
DOI https://doi.org/10.18103/mra.v12i12.5998
ISSN 2375-1924
ABSTRACT
Background and Aim: Machine learning enabled clinical decision support systems offer the potential to enhance the efficiency of decision-making processes in breast cancer multidisciplinary team meetings. We examine the circumstances where a traditional rule-based expert system may have advantages over machine learning enabled clinical decision support systems.
Methods: We compared the concordance of an expert system (Deontics) and a machine learning enabled clinical decision support system (Watson for Oncology) with the treatment recommendations of a gold standard consensus panel of breast surgeons and medical oncologists for 208 non-metastatic breast cancer patients, and for 165 patients deemed eligible for triage to an agreed standard of care, and ‘not for discussion at multidisciplinary team meeting’.
Results: The overall concordance between the Deontics clinical decision support system treatment plan recommendations and the gold standard consensus panel was 98% compared to 92% for the machine learning enabled clinical decision support system. Using a clinical decision tree, 79% of patients were eligible for triage to a standard of care and ‘not for discussion at multidisciplinary team meetings’; for these patients the concordance between the Deontics clinical decision support system and gold standard consensus panel was 98.8% (95% CI: 95.6–99.8%), whilst for the machine learning enabled clinical decision support system concordance was 78.8% (95% CI: 71.7–84.8%).
Conclusion: The high level of agreement between the Deontics clinical decision support system and clinical consensus suggests it may be acceptable for use in the breast multidisciplinary team pathway, whereas the level of disagreement observed for the machine learning enabled clinical decision support system would result in a clinically unsafe error rate if used to triage patients away from the multidisciplinary team meetings. These findings, if replicated in prospective studies in a routine clinical setting, could improve the efficiency of UK breast cancer diagnostic and treatment pathways.
Introduction
Breast cancer remains a significant global health challenge, affecting millions of individuals and their families each year. The complexity of this disease necessitates a multidisciplinary approach, wherein healthcare professionals from diverse specialties collaborate to devise optimal treatment strategies. The multidisciplinary team meetings (MDTM), serve as a pivotal forum for deliberation and decision-making in breast cancer care¹². The quality of decisions made during these MDTMs profoundly impacts the course of patient care and outcomes³⁴. MDTMs have become burdened by increasing workloads with an unmatched, limited increase in resources to support such work¹. Some clinicians have raised concerns about the way cancer MDTMs are conducted in UK as frantic business meetings⁵.
The Association of Breast Surgeons published a toolkit with audit tools to assess the performance on Breast MDTM to identify areas for improvement and streamlining⁶. The guidance together with the audit tools have been put forward to standardise the way in which Breast MDTMs run in the UK. Moreover, in January 2020 NHS England and NHS Improvement issued guidance for Cancer Alliances for Streamlining MDTMs⁷. It proposed a process of “Introducing Standards of Care as a routine part of the MDT process to stratify patient cases into those which require full multidisciplinary discussion in the MDTM, and those cases which can be listed but not discussed in the MDT, as patient need is met by a Standard of Care (SoC)”. A SoC is defined as “a point in the pathway of patient management where there is a recognised international, national, regional or local guideline on the intervention(s) that should be made available to a patient”. The guidance states that for a patient to be assigned for ‘no discussion at the MDT’, the SoC must have been reviewed by an appropriate person or triage group.
Effectively, only complex patients, requiring true multidisciplinary input, would be discussed at MDTMs. Streamlining has not been widely adopted in the UK, most likely due to the lack of clinical time required to take on the task of reviewing all patients prior to the MDTM and assigning those eligible to a pre-agreed SoC².
To support MDTMs in reaching the challenging goal of evidence based informed decision-making, information technology and data science can be helpful to manage, register and re-use all relevant data and generate treatment recommendations. Many studies have shown that clinical decision support systems (CDSS) can be effective tools to increase physician concordance⁸, with clinical practice guidelines¹⁰. Clinical decision support systems can be classified into two broad categories (i) knowledge-driven CDSS (previously known as Expert systems), and (ii) machine learning (ML) or learning algorithm-based data driven CDSS¹³. Recently ML based CDSS have emerged as a transformative tool, offering the potential to enhance the efficiency of decision-making processes within MDTMs¹⁴. However, while ML enabled CDSS excel at handling vast amounts of data and complex pattern recognition, expert systems bring to the table unique advantages, such as transparency, interpretability, predictability and reproducibility. Unlike ML where the underlying logic remains a black box to the user, the knowledge-based CDSS recommendations can be traced back to the underlying evidence source and the logic remains human understandable¹⁵. Therefore, it becomes essential to question whether we have perhaps overlooked the capabilities of traditional expert systems in the rush to embrace ML-based solutions. This study aims to explore the circumstances where expert systems can still prove to be invaluable, challenging the notion that we have entirely discarded their potential in favour of ML, and thus, whether we may have, in some instances, “thrown the baby out with the bathwater.”
Methods
DATA COLLECTION
In this retrospective study, we aimed to evaluate the concordance of expert systems and ML
enabled CDSS with gold standard decisions in breast cancer MDTMs. The local best practice MDT breast cancer treatment decisions, or “consensus panel decisions” were derived from the consensus decisions of two consultant medical oncologists and two consultant breast surgeons with knowledge of the historical MDTM outcome and the expert system and ML enabled CDSS therapeutic options, and these decisions were considered the gold standard. The historical MDTM included a consultant breast medical oncologist, consultant breast surgeon, consultant breast radiologist and pathologist, and clinical nurse specialist.
The study encompassed 213 retrospective cases from a previously published study¹⁴ by the co-authors designed to evaluate a ML enabled CDSS. Cases had been discussed at the Guy’s Cancer Centre MDTM between 2017 and 2018, with patients diagnosed with Stage 1-3 invasive breast cancer. Exclusions were made for patients who could not be analysed due to unavailable treatment options, including those with recurrent breast cancer, bilateral breast cancer, male patients, and specific histological types. Metastatic patients as a part of the first-line therapy were also excluded from this study. The same clinicopathological data was analysed as per the previously published study¹⁴. This included demographic details, comorbidities, functional status, endocrine status, tumour characteristics such as biology (grade/stage) including receptor status and nodal status.
KNOWLEDGE-BASED CLINICAL DECISION SUPPORT (EXPERT SYSTEM)
We utilised Deontics, a clinical decision support and workflow management system grounded in cognitive models that simulate human decision-making processes. Deontics technology excels in evaluating and synthesizing multiple, potentially conflicting arguments, a key requirement in the medical field where uncertainty, incomplete data, and conflicting evidence are common challenges. This approach aligns closely with the principles of epidemiology and evidence-based medicine, as it systematically handles and integrates diverse evidence sources, such as randomized controlled trials (RCTs) and Clinical Practice Guidelines, into transparent recommendations. These recommendations, accompanied by clinical justifications, are presented to clinicians in a clear and comprehensive manner. The system operates in two distinct modes. In human-guided mode, it functions as a decision-support tool, offering suggestions for clinicians to consider, such as in MDTMs. In autonomous mode, the system not only supports complex knowledge representation and nuanced decision-making but also facilitates subsequent workflow management. This dual capability makes Deontics an ideal tool for streamlining clinical pathways, integrating evidence-based decision-making with efficient management of clinical workflows.
Locally accepted breast cancer guidelines, National Comprehensive Cancer Network (NCCN) and National Institute for Health and Care Excellence (NICE) guidelines were used in the Deontics software to create a CDSS tool. The outcome from Deontics was labelled as treatment plan using the following categories: Radiotherapy, Surgery, Systemic therapy, Targeted therapy, Endocrine therapy alone or in combination. These outcome labels were matched by two independent surgical oncologists to the gold standard consensus panel outcome labels. Based on the evidence-based guidance described above, one of the breast surgical oncologists then devised a triage clinical ‘decision tree’ that assigned patients either to an appropriate SoC and ‘not for discussion at MDTM’, or to ‘refer to the MDTM for discussion.’ This decision tree was reviewed by the second senior surgical oncologist and a medical oncologist, and a final version was agreed for use as a triage tool.
MACHINE LEARNING-BASED CLINICAL DECISION SUPPORT
In a prior study¹⁴, the authors evaluated a CDSS, Watson for Oncology, developed by IBM, that used natural language processing and machine learning to generate ranked, evidence based therapeutic options. It was developed in collaboration with experts at Memorial Sloan Kettering Cancer Centre. To validate the CDSS as a streamlining tool the authors developed a decision tree utilising fast-and-frugal trees (FFTs), created with the R package FFTrees, for the purpose of determining the appropriate triage pathway for breast cancer patients – whether to triage them to ‘not for discussion at MDTM’ or ‘send to MDTM for discussion’. FFTs represent supervised learning algorithms geared towards binary classification tasks. However, to permit a direct comparison between the two CDSS approaches, in this study the clinical decision tree described above was used to identify patients eligible for triage to a SoC or to the MDTM for discussion, for both the expert system and ML-based CDSS.
CONCORDANCE ASSESSMENT
Concordance, which measures the agreement between different decision-making approaches and a consensus panel’s decisions, was a primary focus of this study. To calculate concordance, each approach’s treatment decision (ML-based and expert system CDSS) was compared to the consensus panel’s decision for each patient case. The concordance rate was determined, representing the proportion of cases with matching decisions. Concordance Rate = (Number of Cases with Matching Decisions)/(Total Number of Cases). Concordance is reported with 95% confidence intervals (CIs) approximated using the Wilson interval, both for overall concordance, and for concordance only for those cases triaged by the CDSS to ‘not for discussion at MDTM’ using the clinical decision tree.
Service evaluation by the Guy’s Cancer Information Governance team was granted obviating the need for ethical approval.
Results
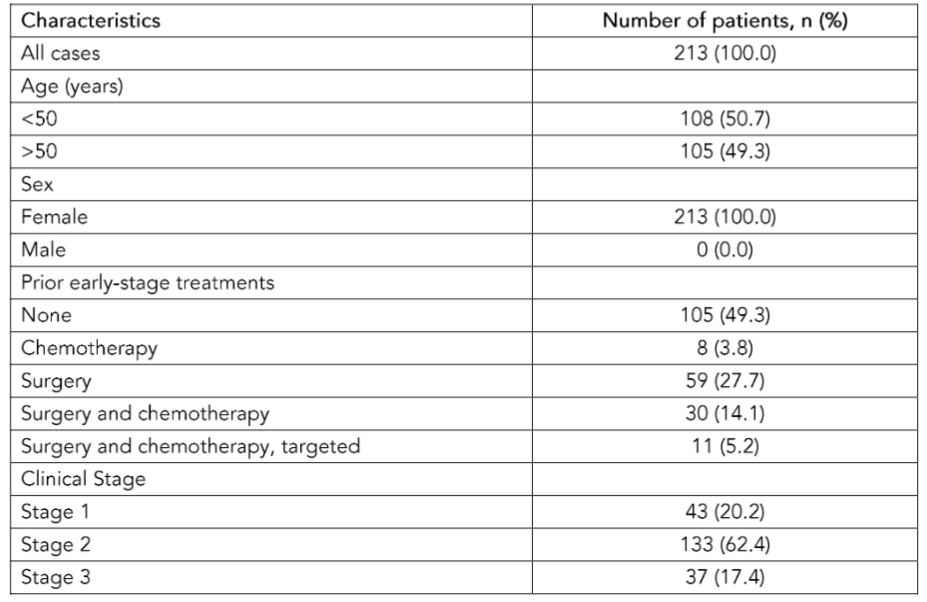
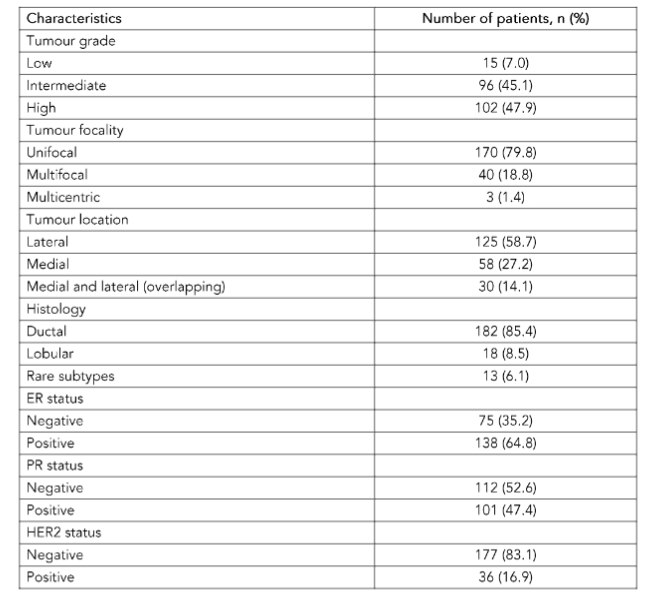
A total of 213 patients were included in the final analysis (Table 1). Case attributes were missing in five patients and therefore concordance was evaluated on 208 patients.
Table 1. The characteristics of each case of breast cancer include in the study, n = 213.
The overall concordance between the ML CDSS treatment plan recommendations and the gold standard consensus panel reported in the previous study¹⁴ was 92% (95% CI: 88–95%, 197/213), following adaptation of the ML CDSS to conform to local best practice¹⁴. Concordance between the Deontics CDSS treatment plan and the gold standard consensus panel was 98% (95% CI: 97–99%, 204/208) (Table 2). The reasons why the Deontics tool provided incorrect recommendations in 5 cases were: 3 cases of recurrence, and the guidance did not cover those; 1 case was a special histological type (adenosquamous); 1 case had a low grade G1 but T3 (large) cancer, where NCCN recommended chemotherapy while local guidelines did not. For reference, the concordance between Deontics and historic MDT was 93% (194/208), and concordance between the gold standard panel and historic MDT was 88% (183/208).
A clinical decision tree was developed based on local recommendations. Appendix 1 shows the clinical decision tree, which was used to triage patients to a SoC, and off to ‘not for discussion at MDTM’. According to the clinical decision tree, a total of 165/208 patients (79%) were eligible to be triaged to CDSS for ‘not for discussion at MDTM’; the main reasons for referral to the MDTM rather than for ‘not for discussion’ were due to multifocality and upgrade to T4 disease. Of the patients sent to ‘not for discussion at MDTM’ the concordance between the Deontics CDSS treatment plan recommendations and gold standard consensus panel was 98.8% (95% CI: 95.6–99.8%, 163/165). The concordance between
the ML CDSS treatment plan recommendations and gold standard consensus panel was 78.8% (95% CI: 71.7–84.8%, 130/165). This gives an observed difference in concordance for the Deontics and ML CDSS when compared with the gold standard panel of 20% (95% CI: 13.6–26.9%).
Table 2. Concordance between gold standard of care consensus panel and machine learning clinical decision support system (CDSS) and Deontics CDSS.
- Overall concordance ML CDSS — 92% (88–95%)
- Overall concordance Deontics CDSS — 98% (97–99%)
- Concordance of patients triaged to SoC Deontics CDSS — 98.8% (95.6–99.8%)
- Concordance of patients triaged to SoC ML CDSS — 78.8% (71.7–84.8%)
SoC; standard of care, reported as 95% CI
Discussion
A CDSS is defined as a system intended to improve healthcare delivery by enhancing medical decisions with targeted clinical knowledge, patient information, and other health information¹⁶. Many studies have shown that CDSS can be effective tools to increase physician concordance with clinical practice guidelines¹⁰¹¹. A systematic review focusing on CDSSs impact on process outcomes (e.g. percentage change MDT treatment decision after using CDSS), guideline adherence and clinical outcomes found that CDSS implementation did not significantly improve process outcomes and guideline adherence but no improvement in clinical outcomes¹⁷. Machine learning is already proving of enormous value in areas such as diagnosing, early detection, recognising patterns, or predicting outcomes from many unselected variables¹⁸, leading to a vast and increasing range of clinical and drug discovery AI applications¹⁹. However, its relative advantages over traditional rules based expert systems may be less convincing when used in the context of clinical decision support, where predictable outcomes are needed, based on the representation of scientifically valid clinical guidelines derived from RCTs.
Ideally, a CDSS should import relevant (standardized) data from the electronic health record automatically and uses these error-free copied source data for decision-support. Machine learning enabled CDSSs are expensive and resource hungry to produce and to update as new research becomes available, both in terms of clinical supervision and training, and computational requirements. They also lack transparency (the so called ‘black box’ problem) making it difficult to build clinical confidence in their decisions, or to identify potential bias or errors in their clinical recommendations. Expert system enabled CDSS on the other hand can be produced relatively cheaply and their algorithms require no data on which to train; only knowledge in the form of written clinical guidelines or protocols, which can be inputted quickly, with no programming skills required². This knowledge can be easily and instantly updated, for example as new research evidence becomes available. Perhaps most importantly for clinical practice, the outputs from an expert system are clinically ‘explainable’ since they can be traced back to the specific guideline recommendation and evidence from which they derive¹. Expert systems like Deontics also make uncertainties and conflicts in the source evidence transparent. In this context, rule-based CDSSs are more intuitive for clinicians to understand compared to systems using machine learning techniques²².
Our study showed a surprising and unexpectedly large advantage in performance of the Deontics CDSS over the ML enabled CDSS when applied to streamlining non-metastatic breast cancer patients in an MDTM pathway. The Deontics CDSS had 98% overall concordance with the gold standard consensus panel, compared to 92% with the ML enabled CDSS. Of greater clinical relevance when considering the use of AI to streamline the breast cancer MDT pathway, when a clinical decision tree was used to identify those patients eligible for triage to an AI CDSS and ‘not for discussion at MDTM’, the Deontics CDSS had 98.8% concordance with the treatment recommendations of a gold standard consensus panel. This compared to only 78.8% concordance for the ML enabled CDSS, which would result in an unacceptable clinically unsafe error rate in SoC decisions for those patients triaged to ‘not for discussion at MDTM’ if the tool were deployed in an MDT pathway. On the other hand, a 98.8% agreement between the expert system CDSS and the gold standard consensus panel suggests it may be acceptable for use in the MDT pathway, where the treating clinician acts as the ‘human in the loop’. If this level of performance was replicated in prospective studies in a routine clinical setting, the potential efficiencies for UK breast cancer diagnostic and treatment pathways from automated streamlining with tools like Deontics would be transformational.
There are several possible reasons for the low performance observed from the ML enabled CDSS when compared with the Deontics CDSS. It may reflect differences between US and UK oncology practice that was not fully adjusted for (although the original authors did attempt to ‘localise’ the ML enabled CDSS to UK practices). The ML enabled CDSS treatment recommendations were generated in 2020 based on the version of IBM Watson for Oncology available at the time. Whilst the gold standard consensus panel treatment recommendations were initially created in 2020, they were reviewed and updated in 2023, and the Deontics CDSS was presented with NCCN²³ and NICE guidelines²⁴, in addition to local Guy’s Cancer Centre protocols current in 2023. IBM discontinued Watson for Oncology in 2021, and Watson Health was subsequently sold to Merative, which until now has not replaced the product. The studies²⁵²⁶ using WFO did not prospectively evaluate these tools, and there remains a significant opportunity to demonstrate the value of AI-based CDSS systems in oncology²⁹.
Since 2020, a new generation of AI technologies have emerged using transformer based large language models that represent a leap forward in AI as great as that from rules based expert systems to deep learning. More recently, large multimodal models have been developed that use text, images, video and audio data. In medicine, these foundation models can be ‘fine-tuned’ on clinical data from electronic patient records in addition to images from radiology, histopathology, and even multi-omic and video data. One example is MedGemini³⁰, trained and fine-tuned by Google DeepMind from its Gemini foundation model to perform multiple clinical tasks, even tasks for which it was not specifically trained, including providing treatment recommendations. MedGemini and other ML technologies based on large foundation models are still at an early research stage. Like other ‘generative’ AI, they tend to ‘hallucinate’, providing superficially plausible answers to clinical questions which are in fact incorrect, with potentially lethal consequences. Until this problem is solved, which may be some years off, these technologies are not safe for use in routine clinical practice. Even then there are other problems relating to costs, and to safe deployment and routine monitoring that may limit their use; and the issue of transparency and explainability remains.
The Deontics CDSS has since been validated at Guy’s Cancer in the prostate MDT pathway as part of an NIHR (National Institute for Health and Care Research) funded study, where in both retrospective and prospective studies it was able to automatically identify and correctly assign 33% of all prostate cancer patients to ‘not for discussion at MDTM’, with 96% concordance with the actual MDTM decision (not yet published, data available on request). On the strength of these findings, and the findings reported here, Guy’s Cancer is now planning to deploy the Deontics CDSS across all the prostate and breast MDT pathways, and potentially to all solid tumour MDT pathways in the future.
Conclusion
Our study has demonstrated the feasibility and validity of using a rules based expert system to automate the streamlining of the majority of non-metastatic breast cancer patients in the MDT pathway to ‘not for discussion at MDTM’. This could reduce the referral to treatment time, and free up significant clinician time to discuss more complex patients or treating more patients. If these findings are replicated in prospective studies, and for other tumour types, cancer centres across the UK should consider deploying such expert systems, and indeed in all health care systems that are concerned with delivering cost-effective cancer care.
Conflict of Interest Statement
VP is the Chief Medical Officer for Deontics Ltd.
Funding Statement:
Mustafa Khanbhai was funded by the AI Centre for Value Based Healthcare which is funded by public sector grants from UK Research and Innovation (UKRI) and the Department of Health and Social Care (DHSC), Office of Life Sciences, delivered through Innovate UK.
Acknowledgements:
Martha Martin contributed to the populating of the initial dataset.
References
1. Taylor C, Munro AJ, Glynne-Jones R, et al. Multidisciplinary team working in cancer: what is the evidence? BMJ. 2010;340:c951.
2. Patkar V, Acosta D, Davidson T, et al. Cancer multidisciplinary team meetings: evidence, challenges, and the role of clinical decision support technology. Int J Breast Cancer. 2011;2011:831605.
3. Prades J, Remue E, van Hoof E, Borras JM. Is it worth reorganising cancer services on the basis of multidisciplinary teams (MDTs)? A systematic review of the objectives and organisation of MDTs and their impact on patient outcomes. Health Policy. 2015;119(4):464-474.
4. Eaker S, Dickman PW, Hellstrom V, et al. Regional differences in breast cancer survival despite common guidelines. Cancer Epidemiol Biomarkers Prev. 2005;14(12):2914-2918.
5. De Ieso PB, Coward JI, Letsa I, et al. A study of the decision outcomes and financial costs of multidisciplinary team meetings (MDMs) in oncology. Br J Cancer. 2013;109(9):2295-2300.
6. Sibering M. Improving the Efficiency of Breast Multidisciplinary Team Meetings: A Toolkit for Breast Services 2022.
7. Streamlining multi-disciplinary team meetings: guidance for cancer alliance. NHS England and NHS Improvement.
8. Mazo C, Kearns C, Mooney C, Gallagher WM. Clinical Decision Support Systems in Breast Cancer: A Systematic Review. Cancers (Basel). 2020;12(2).
9. Sutton RT, Pincock D, Baumgart DC, et al. An overview of clinical decision support systems: benefits, risks, and strategies for success. NPJ Digit Med. 2020;3:17.
10. Garg AX, Adhikari NK, McDonald H, et al. Effects of computerized clinical decision support systems on practitioner performance and patient outcomes: a systematic review. JAMA. 2005;293 (10):1223-1238.
11. Roshanov PS, Fernandes N, Wilczynski JM, et al. Features of effective computerised clinical decision support systems: meta-regression of 162 randomised trials. BMJ. 2013;346:f657.
12. Pawloski PA, Brooks GA, Nielsen ME, Olson-Bullis BA. A Systematic Review of Clinical Decision Support Systems for Clinical Oncology Practice. J Natl Compr Canc Netw. 2019;17(4):331-338.
13. Sloane EB JSR. Artificial intelligence in medical devices and clinical decision support systems. Clinical Engineering Handbook 2020:556-568.
14. Martin M, Kristeleit, H., Ruta, D., et al. Augmentation of a multidisciplinary team meeting with a clinical decision support system to triage breast cancer patients in the United Kingdom. Future Medicine AI. 2023;1(1):1-12.
15. Scott IA. Machine Learning and Evidence-Based Medicine. Ann Intern Med. 2018;169(1):44-46.
16. Greenes RA, Bates DW, Kawamoto K, et al. Clinical decision support models and frameworks: Seeking to address research issues underlying implementation successes and failures. J Biomed Inform. 2018;78:134-143.
17. Klarenbeek SE, Weekenstroo HHA, Sedelaar JPM, et al. The Effect of Higher Level Computerized Clinical Decision Support Systems on Oncology Care: A Systematic Review. Cancers (Basel). 2020;12(4).
18. Johnson KB, Wei WQ, Weeraratne D, et al. Precision Medicine, AI, and the Future of Personalized Health Care. Clin Transl Sci. 2021; 14(1):86-93.
19. Singh S, Kumar R, Payra S, Singh SK. Artificial Intelligence and Machine Learning in Pharmacological Research: Bridging the Gap Between Data and Drug Discovery. Cureus. 2023;15(8):e44359.
20. Amann J, Vetter D, Blomberg SN, et al. To explain or not to explain?-Artificial intelligence explainability in clinical decision support systems. PLOS Digit Health. 2022;1(2):e0000016.
21. Tonekaboni S, Joshi, S., McCradden, M., Goldenberg, A. What Clinicians Want: Contextualizing Explainable Machine Learning for Clinical End Use. Proceedings of Machine Learning Research. 2019:1-21.
22. Bradley A, van der Meer R, McKay C. Personalized Pancreatic Cancer Management: A Systematic Review of How Machine Learning Is Supporting Decision-making. Pancreas. 2019;48 (5):598-604.
23. Gradishar WJ, Moran MS, Abraham J, et al. Breast Cancer, Version 3.2024, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw. 2024;22(5):331-357.
24. Early and locally advanced breast cancer: diagnosis and management 2024.
25. Yu SH, Kim MS, Chung HS, et al. Early experience with Watson for Oncology: a clinical decision-support system for prostate cancer treatment recommendations. World J Urol. 2021;39(2):407-413.
26. Pan H, Tao J, Qian M, et al. Concordance assessment of Watson for Oncology in breast cancer chemotherapy: first China experience. Transl Cancer Res. 2019;8(2):389-401.
27. Zhuang YD, Zhou MC, Liu SC, et al. Effectiveness of personalized 3D printed models for patient education in degenerative lumbar disease. Patient Educ Couns. 2019;102(10):1875-1881.
28. Zhou N, Zhang CT, Lv HY, et al. Concordance Study Between IBM Watson for Oncology and Clinical Practice for Patients with Cancer in China. Oncologist. 2019;24(6):812-819.
29. Nafees A, Khan M, Chow R, et al. Evaluation of clinical decision support systems in oncology: An updated systematic review. Crit Rev Oncol Hematol. 2023;192:104143.
30. Saab K, Tu, T., Weng, WH., et al. Capabilities of Gemini Models in Medicine. arXiv. 2024.