Clinical Tool for Predicting Contralateral Breast Cancer Risk
A Clinical Decision Support Tool to Predict Contralateral Breast Cancer Risk for Women with Primary Breast Cancer
Danielle Braun
Department of Biostatistics, Harvard T.H. Chan School of Public Health, Boston, MA; Department of Data Science, Dana-Farber Cancer Institute, Boston, MA.
Theodore Huang
Department of Biostatistics, Harvard T.H. Chan School of Public Health, Boston, MA; Department of Data Science, Dana-Farber Cancer Institute, Boston, MA; Department of Biostatistics, Vertex Pharmaceuticals, Boston, MA.
Jill Stopfer
Division of Cancer Genetics and Prevention, Department of Medical Oncology, Dana-Farber Cancer Institute, Boston, MA, USA.
Judy Garber
Division of Cancer Genetics and Prevention, Department of Medical Oncology, Dana-Farber Cancer Institute, Boston, MA, USA; Breast Oncology Program, Dana-Farber Brigham Cancer Center, Boston, MA; Harvard Medical School, Boston, MA; Medical Oncology, Dana-Farber Cancer Institute, Boston, MA.
Anna Weiss
Harvard Medical School, Boston, MA; Division of Breast Surgery, Department of Surgery, Brigham and Women’s Hospital, Boston, MA; Breast Oncology Program, Dana-Farber Brigham Cancer Center, Boston, MA; Division of Surgical Oncology, Department of Surgery, University of Rochester, Rochester, NY.
https://doi.org/10.18103/mra.v13i7.6814
COPYRIGHT © 2025 European Society of Medicine. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
ISSN 2375-1924
Abstract
Background/Objectives: The performance of contralateral prophylactic mastectomy significantly reduces the risk of contralateral breast cancer but has also been linked to higher rates of unplanned future operations and psychological stress. The risks of undergoing contralateral prophylactic mastectomy after diagnosis of unilateral primary breast cancer should be weighed against the benefit of risk reduction, which varies widely based on the presence of pathogenic or likely pathogenic variants in high and moderate penetrance genes. In the absence of these mutations, contralateral breast cancer risk varies based on characteristics such as family history, primary tumor characteristics, precursor lesions and other risk factors. Physicians and patients can benefit from risk prediction tools that provide patients personalized risk estimates. Our goal was to develop such a decision support tool to help facilitate informed discussions with breast cancer patients and their clinicians, to make optimal surgical decisions regarding contralateral prophylactic mastectomy.
Methods: For carriers of BRCA1, BRCA2, CHEK2, ATM, PALB2, TP53 data was abstracted from published studies to estimate conditional age-specific contralateral breast cancer risk, which was then used to estimate future risk. For non-carriers, a previously developed and validated contralateral breast cancer risk model was adapted.
Results: We developed an easy-to-use web application that can estimate lifetime and age-conditional risks for both carriers of relatively common breast cancer-related high and moderate penetrance genes as well as for non-carriers.
Conclusions: Our developed tool immediately provides individualized contralateral breast cancer risk estimates and could be extremely helpful during surgical discussions, especially regarding whether or not to perform contralateral prophylactic mastectomy. The tool will be evaluated in a forthcoming randomized controlled trial.
Keywords: Contralateral Breast Cancer; Contralateral Prophylactic Mastectomy; Genetic Counseling; Risk Prediction; Clinical Decision Support Tool
Introduction
Patients with a new diagnosis of primary breast cancer are typically offered more than one option for surgical management. These options include breast conservation, unilateral mastectomy, or bilateral mastectomy which includes removing both the ipsilateral affected breast and the contralateral unaffected breast referred to as contralateral prophylactic mastectomy (CPM). The chance of developing contralateral breast cancer (CBC) can play a significant role in these surgical decisions. CPM is known to reduce the risk of contralateral breast cancer by 90- 95% and in BRCA1/2 carriers who develop breast cancer at a young age CPM is known to reduce the risk of death. However, there is no survival benefit gleaned from the receipt of CPM for non-BRCA patients, and there are also risks associated with CPM including but not limited to frequent need for future revision surgeries and psychological distress, particularly in younger patients.
Contralateral prophylactic mastectomy rates have been on the rise since the early 2000s, sparking debate amongst hereditary professionals, surgeons and the community regarding whether the performance of CPM is medically necessary and if so, for whom. Studies have shown that patients tend to overestimate their own risk of developing contralateral breast cancer, which can lead to higher rates of CPM. It has also been shown that patient education can reduce a patient’s own estimate of developing CBC to be more in line with evidence-based risk predictions and may steer patients away from CPM. Whether performance of CPM is right or wrong, physicians have an obligation to present patients with clear comprehensible data, as available, to help them make their surgical decision.
Contralateral breast cancer risk estimates and the decision to undergo CPM may be relatively straightforward for individuals with pathogenic or likely pathogenic (P/LP) variants in highly penetrant genes such as BRCA1, since lifetime CBC risks range between 50 and 80% and, as mentioned, CPM has been associated with significant reduction in morbidity and mortality. However, a more nuanced decision is required when a patient is a carrier of a P/LP variant in a moderate penetrant gene such as ATM or CHEK2. These genes may still confer considerable CBC risk, possibly 10-40% over a lifetime, which is substantially lower than the CBC risk observed with high penetrance genes such as BRCA1 but also higher than the CBC risk observed among patients who do not carry a P/LP variant. Ambiguity exists because the CBC risks associated with moderate penetrance genes are less well calibrated due to more limited studies with smaller sample sizes. Even high penetrance genes such as TP53 are difficult to study due to low prevalence, and larger sample sizes are needed to be able to accurately measure CBC risk. The development of a CBC risk model, with the ability to estimate CBC risks for individuals with P/LP variants in these understudied moderate penetrance, and less common high penetrance, genes, would have great potential to impact patient care. If more precise CBC risks for these genes can be shared, patients and providers will be better equipped to discuss surgical decisions related to CPM.
Our group has previously developed the All Syndromes Known to Man Evaluator (ASK2ME) clinical decision support tool (https://ask2me.org/). This tool contains CBC risk estimates for BRCA1, BRCA2, and CHEK2 mutation carriers, but does not contain risk estimates for ATM, PALB2, or TP53 carriers, and is not designed specifically for affected breast cancer patients. Giardiello et al. developed a multivariable Fine and Gray prediction model, PredictCBC, to calculate CBC risk with and without BRCA1/2 mutation status, and Sun et al. used a multivariate Cox regression to develop BRCA-CRisk for BRCA1/2 carriers. Lastly, for non-carriers, Chowdhury et al. previously developed and validated the CBCRisk tool (https://cbc-predictor-utd.shinyapps.io/CBCRisk/). Weaknesses of these existing models include the lack of a clinical user interface, and lack of applicability to a range of both carriers and non-carriers for CBC risk estimation.
Our goal was to develop one decision support tool for both carriers and non-carriers that can provide high-quality CBC risk estimates to medical providers and their patients, based on estimated risks from the literature and using statistical techniques similar to those in Braun et al. (ASK2ME) for carriers, and the previously developed and validated CBCRisk for non-carriers. The goal of the work presented here is not to evaluate model performance or estimate model accuracy but rather provide surgeons and patients with a patient-friendly reference with reasonable CBC risk estimations to aid surgical discussions, while acknowledging inherent uncertainties in these estimates. A tool like this could anchor a patient’s perceived risks with scientific, evidence-based estimates and assist them in making better informed surgical decisions by 1) providing personalized risk estimates based on multiple models for germline mutation carriers and non-carriers that considers a wide range of patient characteristics and previously published data as inputs, 2) presenting the output in a format easily understood by patients, and 3) producing results instantaneously. The tool is intended for genetic counselors and surgeons to review contralateral breast cancer risk with patients in the intervention arm of the randomized clinical trial, called GEnetic Testing For All breast Cancer patienTS (GET FACTS; NCT04245176, PI: Weiss), which is examining patient’s self-reported CBC risk estimates and propensity to undergo CPM.
Here we describe the development of this tool for carriers of six different breast cancer genes based on published studies, the adaptation of the CBCRisk model selected for non-carriers, and the software development process. We also describe the final decision support tool (https://hereditarycancer.dfci.harvard.edu/CBCApp/) developed, the resulting risk estimates from the model, and the limitations that were encountered.
Materials and Methods
The steps required to build our tool were divided accordingly:
- For the carrier model, use risk estimates from previously published studies to develop a CBC risk model for carriers of P/LP gene variants for high and moderate penetrance breast cancer genes.
- For the non-carrier model, identify and adapt an existing validated CBC risk model for non-carriers to incorporate into the tool so that it can be utilized by non-carrier patients.
- Iteratively develop, with continuous feedback from genetic counselors, a single web application framework to allow clinical staff to interact with the two models interchangeably to provide a detailed risk report for patients.
EXTERNAL ADVISORY BOARD
Given the uncertainty of most existing risk estimates, including those calculated by our previous and below methods, we convened an external advisory board (EAB) comprised of national experts in medical genetics, surgery, medical oncology, epidemiology, and statistics. The EAB supported development of the decision support tool in several ways, including but not limited to the following: 1) ensured carrier estimates were derived based on highest quality published studies and reviewed modeling assumptions, 2) deliberated and critically evaluated the CBC risk estimate statistical modeling for P/LP variants with less evidence, 3) discussed and adapted the existing CBC risk model for non-carriers, and 4) provided suggested language for providers to use when conveying risk estimation uncertainty to patients.
CARRIER MODEL
Defining the model objectives. Our first objective was to determine CBC risk estimates for as many high and moderate penetrant breast cancer genes as feasible based on available literature. To make our model useful for as many breast cancer patients as possible, we prioritized the following genes for inclusion in our model based on their penetrance and prevalences: BRCA1 (high penetrance/high prevalence), BRCA2 (high penetrance/high prevalence), CHEK2 (moderate penetrance/high prevalence), ATM (moderate penetrance/high prevalence), PALB2 (moderate penetrance/mid prevalence), and TP53 (high penetrance/low prevalence). For CHEK2, we were specifically interested in the high prevalence c.1100delC variant which has been linked to breast cancer. For each gene, our goal was to estimate age-specific CBC risk based on published studies, which can then be used to estimate future CBC.
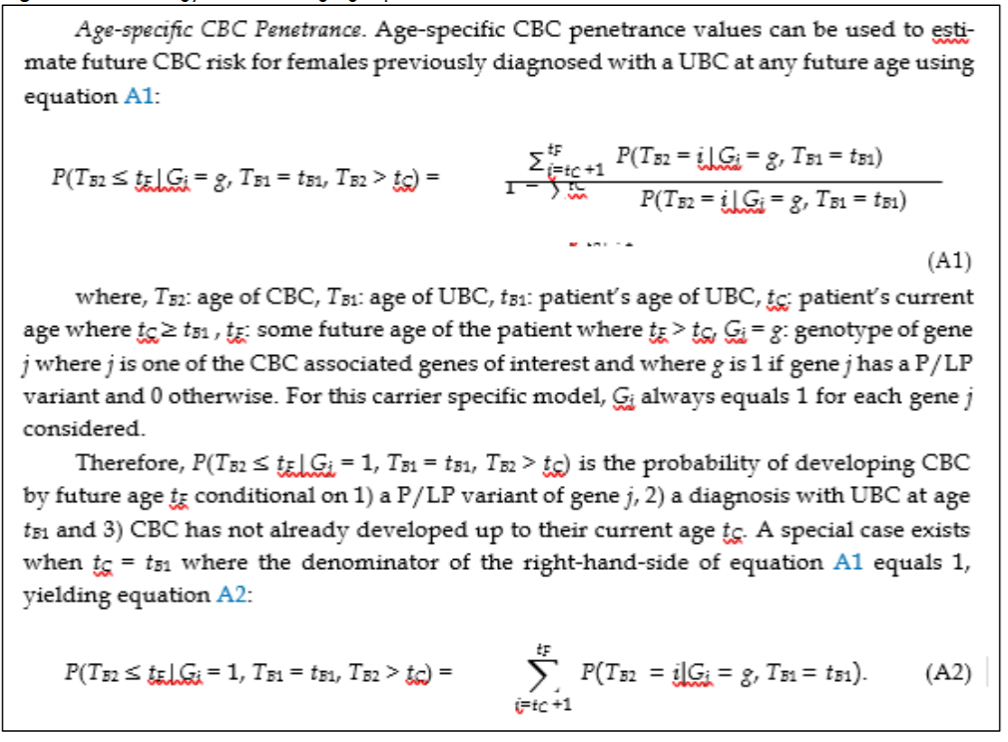

Further details on the estimation of the conditional future risk are available.
Data acquisition and literature review.
For each gene, we did not train new risk prediction models; rather, we conducted a literature review of articles with CBC risk estimations and then derived age-specific risks, as described previously. For genes for which age-specific CBC estimates were previously estimated, including BRCA1, BRCA2, CHEK2, our literature review focused on studies published thereafter. Our inclusion criteria were large studies that reported contralateral breast cancer risk stratified by gene, sex, age at the time of diagnosis, and by either current age or years since diagnosis as described in more detail previously. The identified studies by gene and the quantity extracted from each study are presented in Table 1.
| Gene | Published Study | Extracted Quantity |
|---|---|---|
| BRCA1/BRCA2 | Kuchenbaecker et al. | CBC incidence rates |
| CHEK2/c.1100delC | Kriege et al. | CBC incidence rates |
| ATM/PALB2 | N/A | Estimated from unilateral breast cancer penetrance for these genes |
| TP53 | Guo et al. | Cumulative risk of CBC |
Penetrance estimation for BRCA1 and BRCA2.
Our literature review identified a large, high quality 2017 study previously not incorporated into our previous work for BRCA1 and BRCA2 carriers. The study by Kuchenbaecker et al. provided CBC incidence rates per 1,000 person-years for women stratified by BRCA1/BRCA2, age at primary diagnosis, and years since diagnosis. For each stratum, years since diagnosis were grouped into five year increments out to 20 years post-diagnosis. An additional age category was provided for greater than 20 years post-diagnosis. These reported incidence rates enabled us to estimate age-specific CBC penetrance through linear interpolation. For each of the two genes, age at cancer diagnosis, and future age, the five year incidence rate was divided over the five years in the interval to obtain a probability density value assumed to be equivalent to age-specific CBC penetrance.
Penetrance estimation for CHEK2.
Our literature review did not identify any additional studies on CHEK2 CBC risk since, therefore, the final risks were estimated based on age-conditional cancer penetrance derived from previously published primary breast cancer penetrance.
Penetrance estimation for ATM and PALB2.
Braun et al. did not include age-specific CBC penetrance for ATM and PALB2 nor could we identify additional literature that would allow us to estimate age-specific risks accurately. Instead, we utilized primary breast cancer penetrance for ATM and PALB2 and the BRCA1 gene to estimate CBC penetrance. This required two assumptions: 1) BRCA1 CBC penetrance is larger than that of ATM and PALB2 at every age and 2) the ratios between BRCA1 primary breast cancer penetrance and ATM and PALB2 primary breast cancer penetrances were equivalent to the ratios of CBC penetrances.
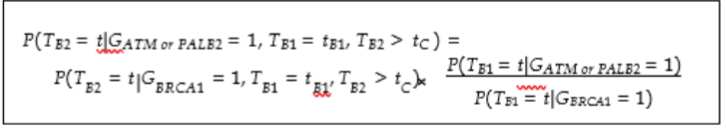
For example, using ASK2ME, we knew the primary penetrance at each age from 1 to 85 for BRCA1 and ATM; however, we only had the CBC penetrance at each age from 1 to 85 for BRCA1. We applied the ratio between the BRCA1 and ATM primary penetrance estimates at every age to obtain the ATM CBC age-conditional penetrance.
Penetrance estimation for TP53.
Braun et al. did not include age-specific CBC penetrance for TP53. Our literature review identified work by Guo et al. (2022) which contained cumulative CBC risks for carriers of TP53 since the first primary breast cancer diagnosis. We obtained the cumulative risk point values from the time to event plot in Guo et al. using the website https://apps.automeris.io/wpd/ which extracts data points from images of plots. Next, the R package functional data analysis was used to fit a monotonically increasing smoothed B-spline to the extracted data points. Using the fitted spline, risk estimates at integer year values from 1 to 15 years after a first breast cancer diagnosis were determined. Next, probability densities at each year post-diagnosis were determined from cumulative risk values and we assumed these approximated to the CBC penetrance values for years 1 to 15 after any age at cancer diagnosis. In the absence of data beyond 15 years post-diagnosis, we assumed the risk from years 16 and on had age-specific CBC penetrance equivalent to non-carrier CBC penetrance obtained from the Surveillance, Epidemiology, and End Results Program (SEER).
NON-CARRIER MODEL
The next step was to incorporate a model for patients without any P/LP gene associated with elevated risk of CBC. For this we selected the CBCRisk model which is contained in the publicly available R package “CBCrisk”. With permission from the package owner, we incorporated the code from the CBCrisk package into our application so that the same tables and graphs displayed for the carriers could be displayed for the non-carriers. Based on feedback from the EAB, patient characteristics with extremely wide confidence intervals were adapted.
ITERATIVE WEB APPLICATION DEVELOPMENT
Genetic counselors from the Cancer Genetics and Prevention program at Dana-Farber Cancer Institute guided the design of the tool from the outset. From selection of the non-carrier risk model through the development of the gene-specific carrier risk models, they provided continual feedback on model assumptions, which genes should be included, and whether the estimated risks were in line with their training and clinical experience. Regular meetings were held to provide the genetic counselors with updates and receive feedback over the course of two years. They also helped select the risk visualizations that patients would be presented in the report and the layout of the web application. Additionally, a medical oncologist and expert medical geneticist, and a breast surgeon were included in the bi-weekly meetings, providing continual feedback so that the decision support tool could also be used during a patient’s surgery consultation.
The tool was coded in the open source R programming language using the R Shiny framework. R Shiny enables R developers to build powerful but light-weight, highly customized web applications capable of handling complex statistical models and custom data visualizations. Its lightweight design gives it an advantage over similar platforms because it allows for rapid iteration of the user interface and back-end calculations. R Shiny allows users to enter model inputs and can produce Portable Document Format (PDF) reports for download. It was crucial that the data visualizations in the report would clearly and simply communicate the results of each of the complex risk models to patients without a medical or other quantitative background. The tool was deployed to the web via R Studio console and was hosted on a R Connect server owned and operated by Dana-Farber Cancer Institute. For privacy and data security, no data was stored beyond a single user session and the connection.
Results
CONTRALATERAL BREAST CANCER RISK TOOL
The resulting app, https://hereditarycancer.dfci.harvard.edu/CBCApp/, has two tabs, one for each risk model depending on the patient’s carrier status. The carrier model has three inputs: patient current age, age at first primary breast cancer diagnosis, and P/LP gene. The non-carrier model has nine inputs: current age, age at first breast cancer diagnosis, if anti-estrogen therapy is being used to treat the first breast cancer, if there are any first degree relatives with BC, history of high risk preneoplasia, breast density, estrogen receptor status, type of first breast cancer (pure DCIS, pure invasive, or mixed), and age at first child birth. Other than the current age, age at cancer diagnosis, and first breast cancer type, all other model inputs have an option to select “Unknown”. For both models, the app collects a patient identifier to assist clinical staff with reporting and storing results in a patients’ electronic medical record.
Once the model inputs are entered, the risk estimates in both table and graph formats appear. A download button generates a PDF report with three data visuals that communicate the patient’s CBC risk.
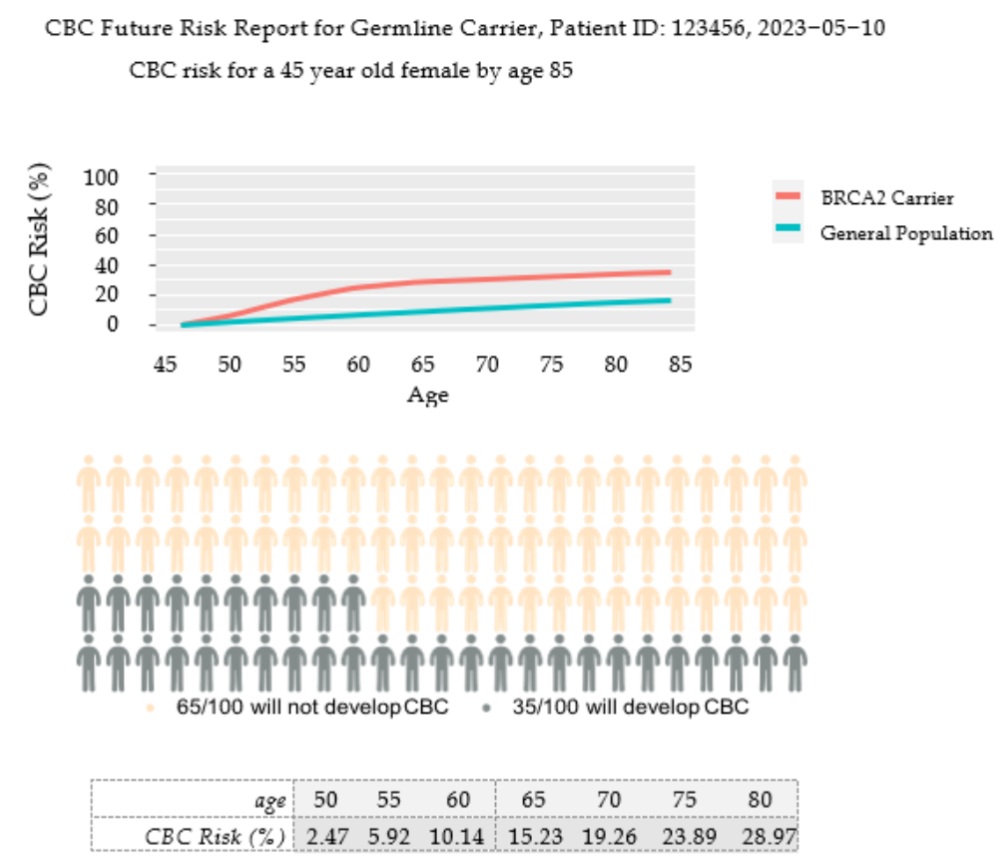
CONTRALATERAL BREAST CANCER RISK ESTIMATES
Carrier model
Table 2 shows the mean lifetime relative CBC risk values stratified by P/LP gene for newly diagnosed breast cancer patients from the carrier model. The mean was taken for all possible ages from 1 to 85. The highly penetrant genes BRCA1, BRCA2, and TP53 had relative risks greater than 3 while the moderately penetrant genes CHEK2, PALB2, and ATM had values between 2 and 3.
| Gene | Mean Relative Risk |
|---|---|
| BRCA1 | 4.32 |
| TP53 | 4.11 |
| BRCA2 | 3.44 |
| CHEK2 | 2.89 |
| PALB2 | 2.80 |
| ATM | 2.49 |
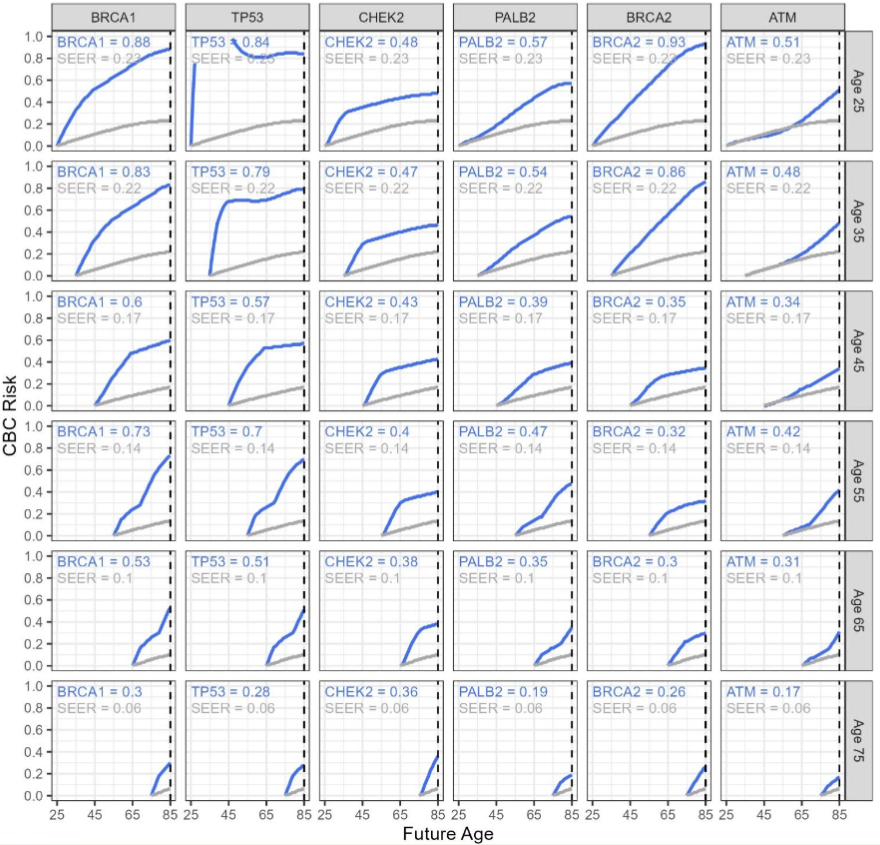
Because this figure is for newly diagnosed patients, the patients’ current ages are equivalent to their ages at the time of their breast cancer diagnosis. Within each column, lifetime risks decrease as patient age increases. BRCA1, BRCA2, and TP53 are generally considered high risk genes; however, our estimates indicate large changes in BRCA2 risks depending on age at diagnosis. Specifically, BRCA2 carriers have the highest risk, as compared to the other genes, for a patient newly diagnosed before age 40 with lifetime risks above 85%. However, if the diagnosis is after age 40, BRCA2 ranks as either the lowest or second lowest risk conferring gene with risks ranging between 26% and 35%. BRCA1 estimates range between greater than 80% at younger ages to as low as 30% at age 75. At each age, the BRCA1 risk estimates are roughly 2-3 percentage points higher than TP53. TP53 is the second highest risk gene when the patient is diagnosed with breast cancer over 40 years of age. CHEK2 risk estimates are consistently moderately penetrant and range from 48% at younger ages down to 36% at older ages. PALB2 and ATM show mid-level risk values ranging from above 50% risk at younger ages to below 20% risk at older ages. Examining the figure down each row reveals that for younger patients (age less than 40), risks from the different gene estimates range from around 50% to above 90%. For middle aged patients between 40 and 60 years old, risk estimates range between around 30% for moderate penetrant genes and above 70% for high penetrant genes. For patients above age 60, risks range from below 20% to above 50%. It is also observed that for P/LP ATM carriers between 25 and 55 years of age, the CBC risk estimates closely follow the SEER estimates 10 to 40 years after diagnosis before increasing thereafter.
Table 3
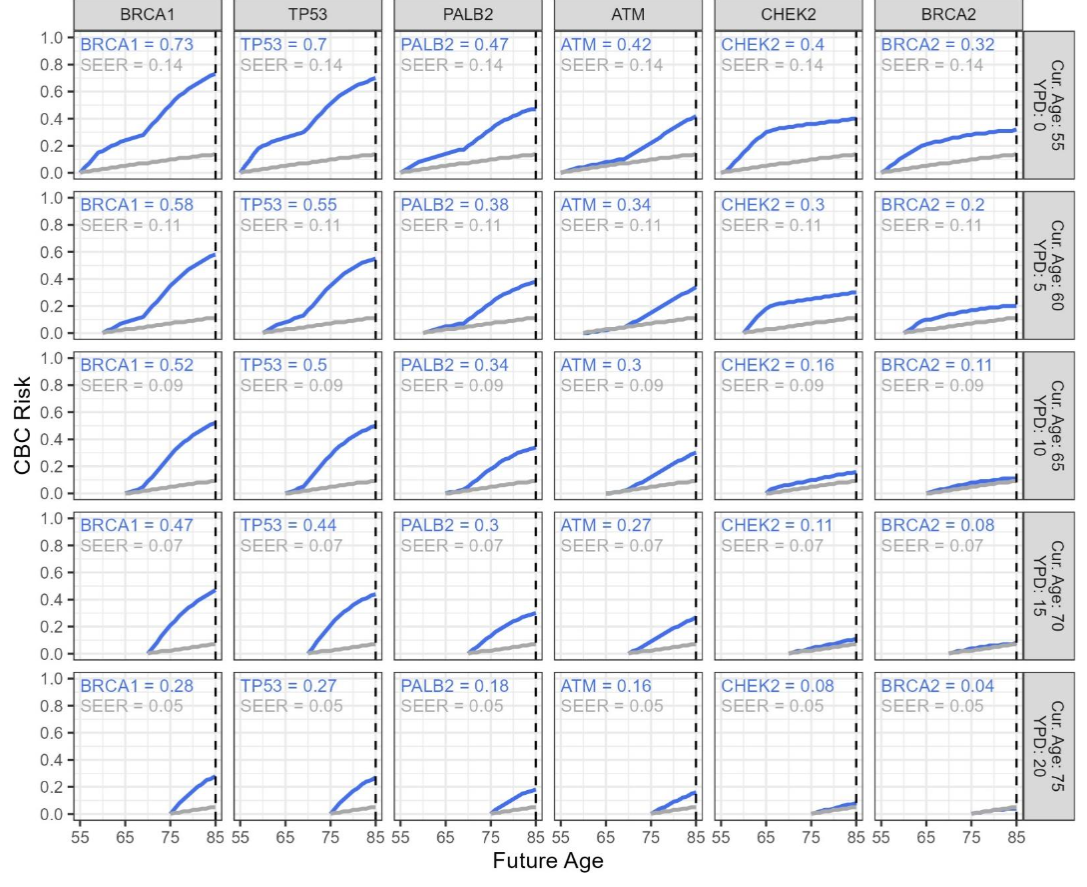
Figure 5 and Table 3 show risk estimates for a patient diagnosed at age 55. Age 55 was chosen as illustration based on the median age of patients enrolled to the American College of Surgeons Oncology Group Z0011 study, a large, representative clinical trial.
| Gene | Current Age/ Age at Breast Cancer Diagnosis | Risk by Age 65 | Risk by Age 80 |
|---|---|---|---|
| BRCA1 | 55 | 23.93% | 64.21% |
| BRCA2 | 55 | 21.01% | 30.23% |
| CHEK2/c.1100delC | 55 | 29.55% | 38.07% |
| ATM | 55 | 7.54% | 32.66% |
| PALB2 | 55 | 14.14% | 41.93% |
| TP53 | 55 | 26.3% | 62.99% |
NON-CARRIER MODEL
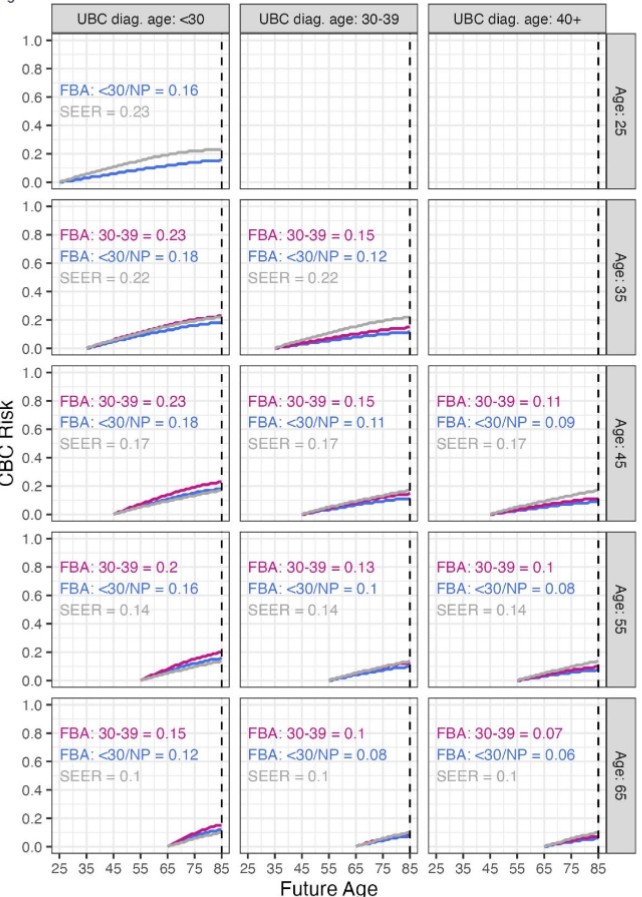
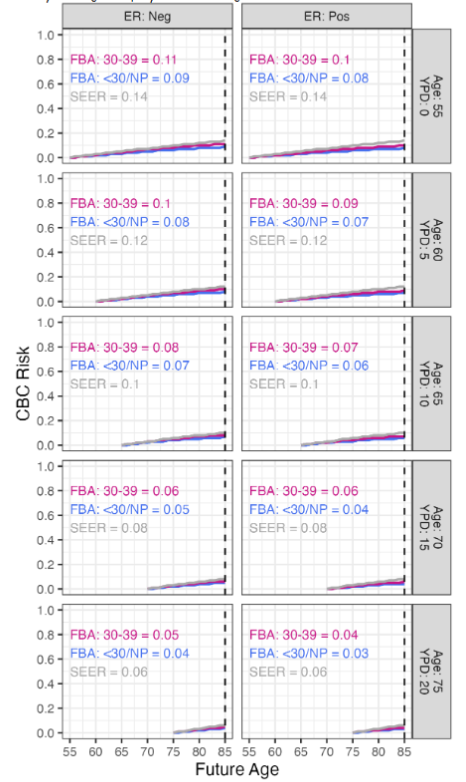
Figure 6 shows CBC risk estimates for non-carriers stratified by diagnosis age, patient age, and age at first live childbirth. The age categories for cancer diagnosis and first live childbirth were dictated by the CBCRisk model and were divided into <30 and 30 − 39. For the first live birth age categories, nulliparous patients were grouped together with the <30 category, and patients with first live birth >40 were grouped together with 30 − 39. First live birth age was included as a stratum in the figure over the other model inputs because it contributes to the greatest variability in CBC risk estimates. As the age of diagnosis increases from less than 30 to between 30 and 39, the risk estimates decrease by 5 to 8 percentage points. The risk estimates decrease an additional 3 to 4 percentage points as the breast cancer diagnosis age range changes from 30 to 39 to age 40 and above. As patient ages increase by each 10 year increment, a two to five percentage point decrease in risk is also observed.
Discussion
We created a clinical decision support tool for newly diagnosed breast cancer patients to estimate their risk of developing contralateral breast cancer to aid surgical treatment decisions. The user interface was designed for clinicians to enter the patient characteristics into one of two models, depending on whether the patient is a carrier of one of six P/LP variants associated with increased CBC risk (ATM, BRCA1, BRCA2, CHEK2, PALB2, or TP53). For carriers of one of these six P/LP gene variants, we estimated risk based on published literature linking these genes to CBC diagnoses and statistical techniques. It is important to note that we did not train new risk models for carriers, rather we used risk estimated in previously published studies. For non-carriers, we leveraged the existing CBCRisk model. Both model outputs take the form of a PDF document that contains figures and a table detailing a patient’s estimated risk at various ages, up until age 85. This report was designed to be easily understood by patients and serve as a discussion guide during their genetic counseling and surgical consultative sessions. Several visualizations were chosen for the report based on EAB and patient advocate input to accommodate different patient learning styles. The table provides the simplest way of viewing the risk estimates in 5 year increments up to age 85. The line graph shows the most detailed information and allows the user to see how risk accumulation accelerates or decelerates throughout their lifetime. The personograph was included because these graphs have been shown to be effective at communicating risk to people of all numeracy levels. Overall, the tool received positive feedback from the genetic counselors and surgeons involved in its development who felt that the estimated risks reflected their clinical training and experience.
What sets our risk estimations apart from others is that we provide risk estimates not just as total lifetime estimates but also for each future age out to age 85, in increments of 5. These more detailed risk estimates provide patients and clinicians with important information required to make well-informed treatment decisions. The decision to undergo CPM has serious implications and should be guided by detailed risk estimates grounded in peer reviewed literature and sound statistical techniques. For example, Figure 5 shows that an ATM carrier diagnosed with breast cancer at age 55 has a lifetime CBC risk of 42%. However, her risk is low through age 70 and is only slightly higher than general population risk. Over the next 10 years of her life, her risk is projected to be less than 10%. Because our tool gives the patient visibility of her relatively low CBC risk for the next 10 to 15 years, the patient may decide not to undergo CPM until after this acute period; whereas, if the patient was only presented with the lifetime risk, they may choose to undergo CPM as soon as possible.
This ability to discern CBC risk trajectories is critically important, especially for young patients, among whom breast cancer incidence is on the rise. As of the early 2010s, bilateral mastectomy was the most common surgical treatment for patients <40 years of age, which is reasonable as younger patients have a higher cumulative lifetime risk of contralateral breast cancer. However, on the other hand, bilateral mastectomy can lead to chronic post-surgical pain and younger patients are both at a higher risk of chronic pain syndrome after mastectomy and also have more years to suffer the side effects of breast cancer treatments administered. Additionally, there are unique considerations in the surgical treatment of young patients that are not always well appreciated, such as future childbearing and desires to breast feed that may drive surgery decisions. If CBC risk magnitude over time can be presented to young breast cancer patients, most importantly carriers of P/LP variants who are at an elevated risk of contralateral breast cancer, it may influence if and when they proceed with CPM. They may feel reassured by a lower than expected risk of CBC over the next 5-10 years, decide to bear a child, breastfeed, and undergo CPM later in life, for example. This clinical decision support tool greatly contributes to surgical decision making.
This tool does have several limitations that should be mentioned. First, the non-carrier model was previously validated, but the carrier model cannot be validated for the same reason that the model is necessary in the first place, specifically we do not have reliable CBC rates for most of the gene mutation carriers. In fact, the carrier model is based on previously published CBC risk estimates that have not been validated either. Additionally, not all breast cancer associated genes were included in the carrier model. For example, the Fam3PRO (previously named PanelPRO) multi-gene/multi-cancer risk model contains age-specific breast cancer penetrance for 11 genes: ATM, BARD1, BRCA1, BRCA2, CDH1, CHEK2, NBN, PALB2, PTEN, STK11, and TP53. We included 6 of these 11 in the carrier model (ATM, BRCA1, BRCA2, CHEK2, PALB2, and TP53) based on a holistic view of each P/LP gene’s relative penetrance, prevalence, and feasibility of CBC risk estimation. It is very likely the other five genes are also associated with CBC, however, to what extent is unknown and a review of available literature determined these risks have not been studied in enough detail to present to patients with any level of certainty. In the absence of these estimations, we planned to present patients who carried variants in these 5 genes with CBC risk estimates for a non-carrier, along with the EAB language around risk and uncertainty. We could have estimated the penetrance for these 5 genes using the same ratio method applied to ATM and PALB2, however, the accuracy of this method has not been validated in other studies of P/LP gene carriers, and we did not want to extend this unvalidated method further to these other more rare and lower penetrance gene variants. Consequently, the method used for estimating the ATM and PALB2 is itself a limitation, relying on a strong assumption that the ratio of risk for a primary breast cancer between BRCA1 and the gene of interest is equivalent to the ratio of risk for CBC between BRCA1 and the gene of interest. While the EAB found this assumption to be reasonable, this has not been validated. Estimating penetrance for additional genes, and others not included in the Fam3PRO model, would require future published literature which stratify CBC risk at least in terms of sex, age at diagnosis, gene, and either patient current age or years since their diagnosis. Literature has suggested that other genes, such as NBN, NF1, BRIP1, RAD51C and RAD51D, may be associated with an inherited predisposition to developing breast cancer; however, to-date, age-specific primary breast cancer risks have not been estimated for these genes, let alone for CBC risks.
The method used to estimate the TP53 risk also has limitations. Guo et al. only reported risks in terms of years since first breast cancer diagnosis and did not consider patient age at diagnosis. Based on the BRCA1, BRCA2, and CHEK2 risk estimates, it is evident that diagnosis age can significantly alter risks with patients diagnosed at younger ages who often face much higher lifetime risk estimates. Furthermore, our method for extrapolating penetrance values out past the end of the study’s 15 year follow up period has its limitations. We assumed that the risk elevation from 16 years after diagnosis to age 85 matched or paralleled the general population’s increase in risk with age. This is unlikely to be the case but, in absence of other data, we felt the assumption was reasonable. Generally, CBC risk is highest from 2 to (7 or 9) years after diagnosis and given that the Guo study reported risks 15 years beyond diagnosis, we felt that our assumption would have minimal impact on clinical decisions. The CBC risk estimates for BRCA2 carriers with primary breast cancer diagnosis before 40 need to be validated using future literature. Figure 4 shows very high risk estimates for this patient type; the BRCA2 risk estimates rank among the lowest risk genes when the primary breast cancer diagnosis age is above 40, however below age 40 BRCA2 becomes the highest risk gene, while the other genes do not show a drastic increase in risk when diagnosis age changes from below age 40 to above age 40. These inconsistencies were noted by the genetic counselors involved in developing the risk tool and lead us to believe the accuracy of these estimates may be in question. Examining the study upon which the risk estimates were based reveals a very wide confidence interval of 25% to 98% for CBC cumulative risks of BRCA2 carriers with a diagnosis age less than 40 who were observed for more than 20 years. Therefore, the root cause is related to the underlying data and not our statistical methods for estimating penetrance.
Another limitation is that the non-carrier model uses additional inputs, other than age at first diagnosis and genotype, as risk modifying factors, yet these factors are not considered in the carrier model. It is likely these risk factors like age at first birth and history of atypia also increase CBC risk for carriers even though our carrier model does not consider it; however, it is also reasonable to assume that the risk conferred by the inherited P/LP variant may drive a patient’s CBC risk more so than the small impacts made by gynecologic risk factors. Developing a carrier model that includes these other factors such as age at first live birth, breast density, estrogen receptor status, and others would have been prohibitive given the currently available literature.
Future iterations of the decision tool presented herein are being considered. Confidence intervals around the CBC risk estimates would allow clinicians to better assess the accuracy of the estimates. For example, we know that the confidence intervals for BRCA2 carriers diagnosed at younger ages are very wide. If the genetic counselor can view confidence intervals on the report, they could communicate the high level of uncertainty to their patients, which could lead to a more informed treatment decision. In the development of the current decision support tool, confidence intervals would have been available for some genes but not others. For example, confidence intervals were reported in the primary literature for breast cancer incidence rates for BRCA1, BRCA2, and CHEK2 which would enable us to estimate confidence intervals for these genes’ CBC risk estimates. However, the paper that the TP53 risk estimates were based on reported point estimates only. Without access to the underlying data, additional literature would need to be published in order to estimate TP53 risk confidence intervals. Similarly for ATM and PALB2, these risks were estimated using ratios relative to BRCA1 and given that the prevalences and penetrance of these two genes are different from BRCA1, using ratios to determine their confidence intervals would be inaccurate. Ultimately, while developing this tool the EAB deemed the confidence intervals too wide and thus too confusing for most of the genes to present to patients during counseling, but this is something to consider for future support tools, and iterations of our own. As new literature on these genes becomes available, we intend to incorporate confidence intervals into the risk tool, which can be toggled on or off depending on the intended audience. We also intend to add more genes to the tool and improve the existing risk estimates for TP53, ATM and PALB2 so that the methods used to estimate their penetrance are more like those for BRCA1, BRCA2, and TP53.
Conclusions
The currently presented clinical decision support tool for contralateral breast cancer risk, https://hereditarycancer.dfci.harvard.edu/CBCApp/, provides patients diagnosed with breast cancer with their personalized CBC risk estimates to aid them in their surgical treatment decisions. More specifically, the information provided by the tool is helpful in weighing the risks and benefits of undergoing CPM. We provide tailored risk estimates based on the characteristics of each patient for both carriers and non-carriers of P/LP variants at future ages increasing by 5 years to the age of 85. For the carriers, we provide CBC risk estimates for six different associated genes which include the highly penetrant genes BRCA1, BRCA2, and TP53 and the moderately penetrant genes ATM, CHEK2, and PALB2. These risk estimates are based on age-specific contralateral breast cancer risk derived from literature. To estimate non-carrier risk, we used the existing CBCRisk model with adaptations based on EAB input. The risk estimates provided by either model are curated in a PDF report designed to communicate CBC risk directly to patients in an easily understood format that includes a table, a line graph, and a personograph. This customized risk report provides CBC risk estimates throughout the patient’s lifetime for carriers and non-carriers alike and enables patients, in consultation with their clinical team, to make a well-informed surgical decision, the efficacy of which will be presented in the forthcoming results of the GEnetic Testing For All breast Cancer patienTS (GET FACTS) randomized clinical trial in which the tool is evaluated as an intervention.
Author Contributions
Conceptualization, D.B., J.S., J.G., A.W.; methodology, D.B., T.H.; software, D.B., T.H.; formal analysis, D.B., T.H.; resources, J.G, A.W.; data curation, D.B., T.H.; writing—original draft preparation, D.B., A.W.; writing—review and editing, D.B., T.H., J.S., A.W.; supervision, D.B., J.G., A.W.; project administration, A.W.; funding acquisition, A.W. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Dana-Farber/Harvard Cancer Center Breast Specialized Program of Research Excellence (SPORE), a National Cancer Institute (NCI)-funded program (grant 1P50CA168504). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health/NCI. Additional funding includes: DFCI’s Friends of Dana-Farber Award, Susan F. Smith Center for Women’s Cancers Breast Gynecologic Cancer Innovation Award.
Conflicts of Interest
Anna Weiss reports an institutional research agreement with Myriad Laboratories Inc. Danielle Braun co-leads the Bayes-Mendel Laboratory, which develops and maintains the BayesMendel and PanelPRO software package. This includes a variety of risk assessment tools, including BRCAPRO, PancPRO, MelaPRO, MMRpro, and PanelPRO/Fam3PRO, which are licensed for commercial use. Theodore Huang, Jill Stopfer, and Judy E. Garber have no conflicts.
Data Availability Statement
The tool is available openly at https://hereditarycancer.dfci.harvard.edu/CBCApp/.
References
- Rebbeck TR, Friebel T, Lynch HT, et al. Bilateral prophylactic mastectomy reduces breast cancer risk in BRCA1 and BRCA2 mutation carriers: the PROSE Study Group. J Clin Oncol. 2004;22(6):1055-1062. doi:10.1200/JCO.2004.04.188
- Li X, You R, Wang X, et al. Effectiveness of Prophylactic Surgeries in BRCA1 or BRCA2 Mutation Carriers: A Meta-analysis and Systematic Review. Clin Cancer Res. 2016;22(15):3971-3981. doi:10.1158/1078-0432.CCR-15-1465
- Blondeaux E, Sonnenblick A, Agostinetto E, et al. Association between risk-reducing surgeries and survival in young BRCA carriers with breast cancer: an international cohort study. Lancet Oncol. 2025;26(6):759-770. doi:10.1016/S1470-2045(25)00152-4
- Kurian AW, Lichtensztajn DY, Keegan TH, Nelson DO, Clarke CA, Gomez SL. Use of and mortality after bilateral mastectomy compared with other surgical treatments for breast cancer in California, 1998-2011. JAMA. 2014;312(9):902-914. doi:10.1001/jama.2014.10707
- Zion SM, Slezak JM, Sellers TA, et al. Reoperations after prophylactic mastectomy with or without implant reconstruction. Cancer. 2003;98(10):2152-2160. doi:10.1002/cncr.11757
- Frost MH, Slezak JM, Tran NV, et al. Satisfaction after contralateral prophylactic mastectomy: the significance of mastectomy type, reconstructive complications, and body appearance. J Clin Oncol. 2005;23(31):7849-7856. doi:10.1200/JCO.2005.09.233.
- King TA, Sakr R, Patil S, et al. Clinical management factors contribute to the decision for contralateral prophylactic mastectomy. J Clin Oncol. 2011;29(16):2158-2164. doi:10.1200/JCO.2010.29.4041
- Wong SM, Freedman RA, Sagara Y, Aydogan F, Barry WT, Golshan M. Growing Use of Contralateral Prophylactic Mastectomy Despite no Improvement in Long-term Survival for Invasive Breast Cancer. Ann Surg. 2017;265(3):581-589. doi:10.1097/SLA.0000000000001698
- Abbott A, Rueth N, Pappas-Varco S, Kuntz K, Kerr E, Tuttle T. Perceptions of contralateral breast cancer: an overestimation of risk. Ann Surg Oncol. 2011;18(11):3129-3136. doi:10.1245/s10434-011-1914-x.
- Alexander NE, Ross J, Sumner W, Nease RF Jr, Littenberg B. The effect of an educational intervention on the perceived risk of breast cancer. J Gen Intern Med. 1996;11(2):92-97. doi:10.1007/BF02599584.
- Yadav S, Boddicker NJ, Na J, et al. Contralateral Breast Cancer Risk Among Carriers of Germline Pathogenic Variants in ATM, BRCA1, BRCA2, CHEK2, and PALB2. J Clin Oncol. 2023;41(9):1703-1713. doi:10.1200/JCO.22.01239
- Kuchenbaecker KB, Hopper JL, Barnes DR, et al. Risks of Breast, Ovarian, and Contralateral Breast Cancer for BRCA1 and BRCA2 Mutation Carriers. JAMA. 2017;317(23):2402-2416. doi:10.1001/jama.2017.7112.
- Braun D, Yang J, Griffin M, Parmigiani G, Hughes KS. A Clinical Decision Support Tool to Predict Cancer Risk for Commonly Tested Cancer-Related Germline Mutations. J Genet Couns. 2018;27(5):1187-1199. doi:10.1007/s10897-018-0238-4.
- Giardiello D, Steyerberg EW, Hauptmann M, et al. Prediction and clinical utility of a contralateral breast cancer risk model. Breast Cancer Res. 2019;21(1):144. Published 2019 Dec 17. doi:10.1186/s13058-019-1221-1.
- Giardiello D, Hooning MJ, Hauptmann M, et al. PredictCBC-2.0: a contralateral breast cancer risk prediction model developed and validated in ~ 200,000 patients. Breast Cancer Res. 2022;24(1):69. Published 2022 Oct 21. doi:10.1186/s13058-022-01567-3.
- Sun J, Chu F, Pan J, et al. BRCA-CRisk: A Contralateral Breast Cancer Risk Prediction Model for BRCA Carriers. J Clin Oncol. 2023;41(5):991-999. doi:10.1200/JCO.22.00833.
- Chowdhury M, Euhus D, Onega T, Biswas S, Choudhary PK. A model for individualized risk prediction of contralateral breast cancer. Breast Cancer Res Treat. 2017;161(1):153-160. doi:10.1007/s10549-016-4039-x.
- Chowdhury M, Euhus D, Arun B, Umbricht C, Biswas S, Choudhary P. Validation of a personalized risk prediction model for contralateral breast cancer. Breast Cancer Res Treat. 2018;170(2):415-423. doi:10.1007/s10549-018-4763-5.
- Kriege M, Hollestelle A, Jager A, et al. Survival and contralateral breast cancer in CHEK2 1100delC breast cancer patients: impact of adjuvant chemotherapy. Br J Cancer. 2014;111(5):1004-1013. doi:10.1038/bjc.2014.306.
- Guo Y, Wan Q, Ouyang T, et al. Risk of ipsilateral breast tumor recurrence and contralateral breast cancer in patients with and without TP53 variant in a large series of breast cancer patients. Breast. 2022;65:55-60. doi:10.1016/j.breast.2022.07.002.
- Ramsay, J. fda: Functional Data Analysis, 2024. R package version 6.1.8.
- Giuliano AE, Hunt KK, Ballman KV, et al. Axillary dissection vs no axillary dissection in women with invasive breast cancer and sentinel node metastasis: a randomized clinical trial. JAMA. 2011;305(6):569-575. doi:10.1001/jama.2011.90.
- Galesic M, Garcia-Retamero R, Gigerenzer G. Using icon arrays to communicate medical risks: overcoming low numeracy. Health Psychol. 2009;28(2):210-216. doi:10.1037/a0014474.
- Xu S, Murtagh S, Han Y, Wan F, Toriola AT. Breast Cancer Incidence Among US Women Aged 20 to 49 Years by Race, Stage, and Hormone Receptor Status. JAMA Netw Open. 2024;7(1):e2353331. Published 2024 Jan 2. doi:10.1001/jamanetworkopen.2023.53331.
- Donaldson J, Swadi S, Frampton C, Brett C. Acute and persistent post-operative pain following mastectomy: a descriptive study in a tertiary hospital cohort. N Z Med J. 2024;137(1601):36-47. Published 2024 Aug 23. doi:10.26635/6965.6524.
- Tesch ME, Partridge AH. Treatment of Breast Cancer in Young Adults. Am Soc Clin Oncol Educ Book. 2022;42:1-12. doi:10.1200/EDBK_360970.
- Lee G, Liang JW, Zhang Q, et al. Multi-syndrome, multi-gene risk modeling for individuals with a family history of cancer with the novel R package PanelPRO. Elife. 2021;10:e68699. Published 2021 Aug 18. doi:10.7554/eLife.68699.
- Couch FJ, Shimelis H, Hu C, et al. Associations Between Cancer Predisposition Testing Panel Genes and Breast Cancer. JAMA Oncol. 2017;3(9):1190-1196. doi:10.1001/jamaoncol.2017.0424.
- Easton DF, Pharoah PD, Antoniou AC, et al. Gene-panel sequencing and the prediction of breast-cancer risk. N Engl J Med. 2015;372(23):2243-2257. doi:10.1056/NEJMsr1501341.

